Abstract
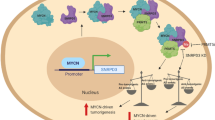
The nuclear factor erythroid 2-like 2 (NFE2L2; NRF2) signaling pathway is frequently deregulated in human cancers. The critical functions of NRF2, other than its transcriptional activation, in cancers remain largely unknown. Here, we uncovered a previously unrecognized role of NRF2 in the regulation of RNA splicing. Global splicing analysis revealed that NRF2 knockdown in non-small cell lung cancer (NSCLC) A549 cells altered 839 alternative splicing (AS) events in 485 genes. Mechanistic studies demonstrated that NRF2 transcriptionally regulated SMN mRNA expression by binding to two antioxidant response elements in the SMN1 promoter. Post-transcriptionally, NRF2 was physically associated with the SMN protein. The Neh2 domain of NRF2, as well as the YG box and the region encoded by exon 7 of SMN, were required for their interaction. NRF2 formed a complex with SMN and Gemin2 in nuclear gems and Cajal bodies. Furthermore, the NRF2–SMN interaction regulated RNA splicing by expressing SMN in NRF2-knockout HeLa cells, reverting some of the altered RNA splicing. Moreover, SMN overexpression was significantly associated with alterations in the NRF2 pathway in patients with lung squamous cell carcinoma from The Cancer Genome Atlas. Taken together, our findings suggest a novel therapeutic strategy for cancers involving an aberrant NRF2 pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this paper and its supplementary files. The RNA-Seq and ChIP-seq data of raw and aligned files have been deposited in NCBI Gene Expression Omnibus, accession number GSE164720 and 141497, respectively.
References
Urbanski LM, Leclair N, Anczukow O. Alternative-splicing defects in cancer: splicing regulators and their downstream targets, guiding the way to novel cancer therapeutics. Wiley Interdiscip Rev RNA. 2018;9:e1476.
Wang BD, Lee NH. Aberrant RNA splicing in cancer and drug resistance. Cancers. 2018;10:458.
Furukawa M, Xiong Y. BTB protein Keap1 targets antioxidant transcription factor Nrf2 for ubiquitination by the Cullin 3-Roc1 ligase. Mol Cell Biol. 2005;25:162–71.
McMahon M, Itoh K, Yamamoto M, Hayes JD. Keap1-dependent proteasomal degradation of transcription factor Nrf2 contributes to the negative regulation of antioxidant response element-driven gene expression. J Biol Chem. 2003;278:21592–600.
Tong KI, Kobayashi A, Katsuoka F, Yamamoto M. Two-site substrate recognition model for the Keap1-Nrf2 system: a hinge and latch mechanism. Biol Chem. 2006;387:1311–20.
Dinkova-Kostova AT, Holtzclaw WD, Cole RN, Itoh K, Wakabayashi N, Katoh Y, et al. Direct evidence that sulfhydryl groups of Keap1 are the sensors regulating induction of phase 2 enzymes that protect against carcinogens and oxidants. Proc Natl Acad Sci USA. 2002;99:11908–13.
Itoh K, Wakabayashi N, Katoh Y, Ishii T, Igarashi K, Engel JD, et al. Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes Dev. 1999;13:76–86.
Hayes JD, Dinkova-Kostova AT. The Nrf2 regulatory network provides an interface between redox and intermediary metabolism. Trends Biochem Sci. 2014;39:199–218.
Hayes JD, McMahon M. NRF2 and KEAP1 mutations: permanent activation of an adaptive response in cancer. Trends Biochem Sci. 2009;34:176–88.
Cancer Genome Atlas Research N. Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511:543–50.
Namani A, Cui QQ, Wu Y, Wang H, Wang XJ, Tang X. NRF2-regulated metabolic gene signature as a prognostic biomarker in non-small cell lung cancer. Oncotarget. 2017;8:69847–62.
Raimer AC, Gray KM, Matera AG. SMN - a chaperone for nuclear RNP social occasions? RNA Biol. 2017;14:701–11.
Nilsen TW. The spliceosome: the most complex macromolecular machine in the cell? Bioessays. 2003;25:1147–9.
Will CL, Luhrmann R. Spliceosomal UsnRNP biogenesis, structure and function. Curr Opin Cell Biol. 2001;13:290–301.
Neugebauer KM. Special focus on the Cajal Body. RNA Biol. 2017;14:669–70.
Dominguez CE, Cunningham D, Chandler DS. SMN regulation in SMA and in response to stress: new paradigms and therapeutic possibilities. Hum Genet. 2017;136:1173–91.
Tang X, Wang H, Fan L, Wu X, Xin A, Ren H, et al. Luteolin inhibits Nrf2 leading to negative regulation of the Nrf2/ARE pathway and sensitization of human lung carcinoma A549 cells to therapeutic drugs. Free Radic Biol Med. 2011;50:1599–609.
Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci USA. 2014;111:E5593–601.
Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Thomas-Chollier M, Hufton A, Heinig M, O'Keeffe S, Masri NE, Roider HG, et al. Transcription factor binding predictions using TRAP for the analysis of ChIP-seq data and regulatory SNPs. Nat Protoc. 2011;6:1860–9.
Martin R, Gupta K, Ninan NS, Perry K, Van, Duyne GD. The survival motor neuron protein forms soluble glycine zipper oligomers. Structure. 2012;20:1929–39.
Seng CO, Magee C, Young PJ, Lorson CL, Allen JP. The SMN structure reveals its crucial role in snRNP assembly. Hum Mol Genet. 2015;24:2138–46.
Burghes AH, Beattie CE. Spinal muscular atrophy: why do low levels of survival motor neuron protein make motor neurons sick? Nat Rev Neurosci. 2009;10:597–609.
DeNicola GM, Karreth FA, Humpton TJ, Gopinathan A, Wei C, Frese K, et al. Oncogene-induced Nrf2 transcription promotes ROS detoxification and tumorigenesis. Nature. 2011;475:106–9.
Mitsuishi Y, Taguchi K, Kawatani Y, Shibata T, Nukiwa T, Aburatani H, et al. Nrf2 redirects glucose and glutamine into anabolic pathways in metabolic reprogramming. Cancer Cell. 2012;22:66–79.
Stamm S, Ben-Ari S, Rafalska I, Tang Y, Zhang Z, Toiber D, et al. Function of alternative splicing. Gene. 2005;344:1–20.
Ghigna C, Valacca C, Biamonti G. Alternative splicing and tumor progression. Curr Genom. 2008;9:556–70.
Venables JP. Unbalanced alternative splicing and its significance in cancer. Bioessays. 2006;28:378–86.
Venables JP, Klinck R, Koh C, Gervais-Bird J, Bramard A, Inkel L, et al. Cancer-associated regulation of alternative splicing. Nat Struct Mol Biol. 2009;16:670–6.
David CJ, Manley JL. Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev. 2010;24:2343–64.
Rambout X, Dequiedt F, Maquat LE. Beyond transcription: roles of transcription factors in Pre-mRNA splicing. Chem Rev. 2018;118:4339–64.
Hsu TY, Simon LM, Neill NJ, Marcotte R, Sayad A, Bland CS, et al. The spliceosome is a therapeutic vulnerability in MYC-driven cancer. Nature. 2015;525:384–8.
Nizzardo M, Nardini M, Ronchi D, Salani S, Donadoni C, Fortunato F, et al. Beta-lactam antibiotic offers neuroprotection in a spinal muscular atrophy model by multiple mechanisms. Exp Neurol. 2011;229:214–25.
Lefebvre S, Burglen L, Reboullet S, Clermont O, Burlet P, Viollet L, et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell. 1995;80:155–65.
Lorson CL, Strasswimmer J, Yao JM, Baleja JD, Hahnen E, Wirth B, et al. SMN oligomerization defect correlates with spinal muscular atrophy severity. Nat Genet. 1998;19:63–6.
Chen Y, Liu K, Zhang J, Hai Y, Wang P, Wang H, et al. c-Jun NH2 -Terminal protein kinase phosphorylates the Nrf2-ECH homology 6 domain of nuclear factor erythroid 2-related factor 2 and downregulates cytoprotective genes in acetaminophen-induced liver injury in mice. Hepatology. 2020;71:1787–801.
Li J, Wang H, Zheng Z, Luo L, Wang P, Liu K, et al. Mkp-1 cross-talks with Nrf2/Ho-1 pathway protecting against intestinal inflammation. Free Radic Biol Med. 2018;124:541–9.
Luo L, Chen Y, Wang H, Wang S, Liu K, Li X, et al. Mkp-1 protects mice against toxin-induced liver damage by promoting the Nrf2 cytoprotective response. Free Radic Biol Med. 2018;115:361–70.
Wang H, Liu K, Geng M, Gao P, Wu X, Hai Y, et al. RXRalpha inhibits the NRF2-ARE signalling pathway through a direct interaction with the Neh7 domain of NRF2. Cancer Res. 2013;73:3097–108.
Bruns AF, van Bergeijk J, Lorbeer C, Nolle A, Jungnickel J, Grothe C, et al. Fibroblast growth factor-2 regulates the stability of nuclear bodies. Proc Natl Acad Sci USA. 2009;106:12747–52.
Nolle A, Zeug A, van Bergeijk J, Tonges L, Gerhard R, Brinkmann H, et al. The spinal muscular atrophy disease protein SMN is linked to the rho-kinase pathway via profilin. Hum Mol Genet. 2011;20:4865–78.
Acknowledgements
We thank Prof. Peter Claus (Institute of Neuroanatomy, Hannover Medical School, Hannover, Germany) for providing the plasmids pEGFP-N2-SMN-1-294 and pET41a- SMN-1-294, and pEGFP-N2-Coilin, Prof. Qiming Sun (Zhejiang University, China) for providing Furipw vector, Prof. Hong-He Zhang and Prof. Yin-Jie Wang (Zhejiang University, China) for comments on the MS. We thank Yanwei Li, Guifeng Xiao, Wei Yin, Zhaoxiaonan Lin and Sanhua Fang from the Core Facilities, Zhejiang University School of Medicine for their technical support. We thank GENERGY BIO-TECHNOLOGY (SHANGHAI) CO., LTD. for RNA-Seq and data analysis. We also thank Lab members Miss Yi-jiao Liao, Miss Yu Tian and Mr. Yemin Shan for technical help. This work was supported by grants from the National Natural Science Foundation of China (31971188 to XT, 91643110 to XJW and 31700690 to YW).
Author information
Authors and Affiliations
Contributions
Conceptualization: XT and XJW; Methodology: QC, WW, AN, HW, AH, PH, YG, ME, YW, XJW and XT; Investigation: QC, WW, AN, HW, AH, YG, ME, XJW and XT; Writing—original draft: XT and XJW; Writing—revision: XT; Funding Acquisition: XT, XJW, PH and YW. Supervision: XT.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cui, Q., Wang, W., Namani, A. et al. NRF2 has a splicing regulatory function involving the survival of motor neuron (SMN) in non-small cell lung cancer. Oncogene 42, 2751–2763 (2023). https://doi.org/10.1038/s41388-023-02799-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-023-02799-z