Abstract
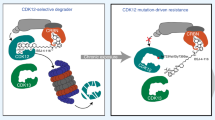
WD repeat domain 5 (WDR5), an integral component of the MLL/KMT2A lysine methyltransferase complex, is critically involved in oncogenesis and represents an attractive onco-target. Inhibitors targeting protein-protein interactions (PPIs) between WDR5 and its binding partners, however, do not inhibit all of WDR5-mediated oncogenic functions and exert rather limited antitumor effects. Here, we report a cereblon (CRBN)-recruiting proteolysis targeting chimera (PROTAC) of WDR5, MS40, which selectively degrades WDR5 and the well-established neo-substrates of immunomodulatory drugs (IMiDs):CRBN, the Ikaros zinc finger (IKZF) transcription factors IKZF1 and IKZF3. MS40-induced WDR5 degradation caused disassociation of the MLL/KMT2A complex off chromatin, resulting in decreased H3K4me2. Transcriptomic profiling revealed that targets of both WDR5 and IMiDs:CRBN were significantly repressed by treatment of MS40. In MLL-rearranged leukemias, which exhibit IKZF1 high expression and dependency, co-suppression of WDR5 and Ikaros by MS40 is superior in suppressing oncogenesis to the WDR5 PPI inhibitor, to MS40’s non-PROTAC analog controls (MS40N1 and MS40N2, which do not bind CRBN and WDR5, respectively), and to a matched VHL-based WDR5 PROTAC (MS169, which degrades WDR5 but not Ikaros). MS40 suppressed the growth of primary leukemia patient cells in vitro and patient-derived xenografts in vivo. Thus, dual degradation of WDR5 and Ikaros is a promising anti-cancer strategy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ge Z, Song EJ, Kawasawa YI, Li J, Dovat S, Song C. WDR5 high expression and its effect on tumorigenesis in leukemia. Oncotarget. 2016;7:37740–54.
Carugo A, Genovese G, Seth S, Nezi L, Rose JL, Bossi D, et al. In vivo functional platform targeting patient-derived xenografts identifies WDR5-Myc association as a critical determinant of pancreatic cancer. Cell Rep. 2016;16:133–47.
Sun Y, Bell JL, Carter D, Gherardi S, Poulos RC, Milazzo G, et al. WDR5 Supports an N-Myc Transcriptional Complex That Drives a Protumorigenic Gene Expression Signature in Neuroblastoma. Cancer Res. 2015;75:5143–54.
Miller T, Krogan NJ, Dover J, Erdjument-Bromage H, Tempst P, Johnston M, et al. COMPASS: a complex of proteins associated with a trithorax-related SET domain protein. Proc Natl Acad Sci USA. 2001;98:12902–7.
Trievel RC, Shilatifard A. WDR5, a complexed protein. Nat Struct Mol Biol. 2009;16:678–80.
Rao RC, Dou Y. Hijacked in cancer: the KMT2 (MLL) family of methyltransferases. Nat Rev Cancer. 2015;15:334–46.
Patel A, Dharmarajan V, Cosgrove MS. Structure of WDR5 bound to mixed lineage leukemia protein-1 peptide. J Biol Chem. 2008;283:32158–61.
Patel A, Vought VE, Dharmarajan V, Cosgrove MS. A conserved arginine-containing motif crucial for the assembly and enzymatic activity of the mixed lineage leukemia protein-1 core complex. J Biol Chem. 2008;283:32162–75.
Dharmarajan V, Lee JH, Patel A, Skalnik DG, Cosgrove MS. Structural basis for WDR5 interaction (Win) motif recognition in human SET1 family histone methyltransferases. J Biol Chem. 2012;287:27275–89.
Thomas LR, Wang Q, Grieb BC, Phan J, Foshage AM, Sun Q, et al. Interaction with WDR5 promotes target gene recognition and tumorigenesis by MYC. Mol Cell. 2015;58:440–52.
Guarnaccia AD, Tansey WP. Moonlighting with WDR5: a cellular multitasker. J Clin Med. 2018; 7.
Alicea-Velazquez NL, Shinsky SA, Loh DM, Lee JH, Skalnik DG, Cosgrove MS. Targeted Disruption of the Interaction between WD-40 Repeat Protein 5 (WDR5) and Mixed Lineage Leukemia (MLL)/SET1 family proteins specifically inhibits MLL1 and SETd1A methyltransferase complexes. J Biol Chem. 2016;291:22357–72.
Odho Z, Southall SM, Wilson JR. Characterization of a novel WDR5-binding site that recruits RbBP5 through a conserved motif to enhance methylation of histone H3 lysine 4 by mixed lineage leukemia protein-1. J Biol Chem. 2010;285:32967–76.
Senisterra G, Wu H, Allali-Hassani A, Wasney GA, Barsyte-Lovejoy D, Dombrovski L, et al. Small-molecule inhibition of MLL activity by disruption of its interaction with WDR5. Biochem J. 2013;449:151–9.
Bolshan Y, Getlik M, Kuznetsova E, Wasney GA, Hajian T, Poda G, et al. Synthesis, Optimization, and Evaluation of Novel Small Molecules as Antagonists of WDR5-MLL Interaction. ACS Med Chem Lett. 2013;4:353–7.
Karatas H, Townsend EC, Cao F, Chen Y, Bernard D, Liu L, et al. High-affinity, small-molecule peptidomimetic inhibitors of MLL1/WDR5 protein-protein interaction. J Am Chem Soc. 2013;135:669–82.
Cao F, Townsend EC, Karatas H, Xu J, Li L, Lee S, et al. Targeting MLL1 H3K4 methyltransferase activity in mixed-lineage leukemia. Mol Cell. 2014;53:247–61.
Grebien F, Vedadi M, Getlik M, Giambruno R, Grover A, Avellino R, et al. Pharmacological targeting of the Wdr5-MLL interaction in C/EBPalpha N-terminal leukemia. Nat Chem Biol. 2015;11:571–8.
Zhu J, Sammons MA, Donahue G, Dou Z, Vedadi M, Getlik M, et al. Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth. Nature. 2015;525:206–11.
Getlik M, Smil D, Zepeda-Velazquez C, Bolshan Y, Poda G, Wu H, et al. Structure-based optimization of a small molecule antagonist of the interaction between WD repeat-containing protein 5 (WDR5) and mixed-lineage leukemia 1 (MLL1). J Med. Chem. 2016;59:2478–96.
Li DD, Chen WL, Wang ZH, Xie YY, Xu XL, Jiang ZY, et al. High-affinity small molecular blockers of mixed lineage leukemia 1 (MLL1)-WDR5 interaction inhibit MLL1 complex H3K4 methyltransferase activity. Eur J Med Chem. 2016;124:480–9.
Karatas H, Li Y, Liu L, Ji J, Lee S, Chen Y, et al. Discovery of a highly potent, cell-permeable macrocyclic peptidomimetic (MM-589) targeting the WD repeat domain 5 protein (WDR5)-mixed lineage leukemia (MLL) protein-protein interaction. J medicinal Chem. 2017;60:4818–39.
Wang F, Jeon KO, Salovich JM, Macdonald JD, Alvarado J, Gogliotti RD. et al. Discovery of potent 2-Aryl-6,7-dihydro-5 H-pyrrolo[1,2- a]imidazoles as WDR5-WIN-site inhibitors using fragment-based methods and structure-based design. J. Med. Chem. 2018;61:5623–42..
Aho ER, Wang J, Gogliotti RD, Howard GC, Phan J, Acharya P, et al. Displacement of WDR5 from Chromatin by a WIN Site Inhibitor with Picomolar Affinity. Cell Rep. 2019;26:2916–28. e2913
Macdonald JD, Chacon Simon S, Han C, Wang F, Shaw JG, Howes JE, et al. Discovery and optimization of salicylic acid-derived sulfonamide inhibitors of the WD repeat-containing protein 5-MYC protein-protein interaction. J Med Chem. 2019;62:11232–59.
Tian J, Teuscher KB, Aho ER, Alvarado JR, Mills JJ, Meyers KM, et al. Discovery and structure-based optimization of potent and selective WD repeat domain 5 (WDR5) inhibitors containing a dihydroisoquinolinone bicyclic core. J Med Chem. 2020;63:656–75.
Chacon Simon S, Wang F, Thomas LR, Phan J, Zhao B, Olejniczak ET, et al. Discovery of WD repeat-containing protein 5 (WDR5)-MYC inhibitors using fragment-based methods and structure-based design. J Med Chem. 2020;63:4315–33.
Lai AC, Crews CM. Induced protein degradation: an emerging drug discovery paradigm. Nat Rev Drug Discov. 2017;16:101–14.
Schapira M, Calabrese MF, Bullock AN, Crews CM. Targeted protein degradation: expanding the toolbox. Nat Rev Drug Discov. 2019;18:949–63.
Dale B, Cheng M, Park KS, Kaniskan HU, Xiong Y, Jin J. Advancing targeted protein degradation for cancer therapy. Nat Rev Cancer. 2021;21:638–54.
Yu X, Li D, Kottur J, Shen Y, Kim HS, Park KS, et al. A selective WDR5 degrader inhibits acute myeloid leukemia in patient-derived mouse models. Sci Transl Med. 2021;13:eabj1578.
Dolle A, Adhikari B, Kramer A, Weckesser J, Berner N, Berger LM, et al. Design, synthesis, and evaluation of WD-repeat-containing protein 5 (WDR5) degraders. J Med Chem. 2021;64:10682–710.
Akuffo AA, Alontaga AY, Metcalf R, Beatty MS, Becker A, McDaniel JM, et al. Ligand-mediated protein degradation reveals functional conservation among sequence variants of the CUL4-type E3 ligase substrate receptor cereblon. J Biol Chem. 2018;293:6187–200.
Mori T, Ito T, Liu S, Ando H, Sakamoto S, Yamaguchi Y, et al. Structural basis of thalidomide enantiomer binding to cereblon. Sci Rep. 2018;8:1294.
Lu G, Middleton RE, Sun H, Naniong M, Ott CJ, Mitsiades CS, et al. The myeloma drug lenalidomide promotes the cereblon-dependent destruction of Ikaros proteins. Science. 2014;343:305–9.
Kronke J, Fink EC, Hollenbach PW, MacBeth KJ, Hurst SN, Udeshi ND, et al. Lenalidomide induces ubiquitination and degradation of CK1alpha in del(5q) MDS. Nature. 2015;523:183–8.
Matyskiela ME, Lu G, Ito T, Pagarigan B, Lu CC, Miller K, et al. A novel cereblon modulator recruits GSPT1 to the CRL4(CRBN) ubiquitin ligase. Nature. 2016;535:252–7.
Ishoey M, Chorn S, Singh N, Jaeger MG, Brand M, Paulk J, et al. Translation termination factor GSPT1 is a phenotypically relevant off-target of heterobifunctional phthalimide degraders. ACS Chem Biol. 2018;13:553–60.
An J, Ponthier CM, Sack R, Seebacher J, Stadler MB, Donovan KA, et al. pSILAC mass spectrometry reveals ZFP91 as IMiD-dependent substrate of the CRL4(CRBN) ubiquitin ligase. Nat Commun. 2017;8:15398.
Xia R, Cheng Y, Han X, Wei Y, Wei X. Ikaros proteins in tumor: current perspectives and new developments. Front Mol Biosci. 2021;8:788440.
Park I, Phan TM, Fang J. Novel molecular mechanism of lenalidomide in myeloid malignancies independent of deletion of chromosome 5q. Cancers (Basel) 2021; 13.
Dou Y, Milne TA, Tackett AJ, Smith ER, Fukuda A, Wysocka J, et al. Physical association and coordinate function of the H3 K4 methyltransferase MLL1 and the H4 K16 acetyltransferase MOF. Cell. 2005;121:873–85.
Steward MM, Lee JS, O’Donovan A, Wyatt M, Bernstein BE, Shilatifard A. Molecular regulation of H3K4 trimethylation by ASH2L, a shared subunit of MLL complexes. Nat Struct Mol Biol. 2006;13:852–4.
Patel A, Vought VE, Dharmarajan V, Cosgrove MS. A novel non-SET domain multi-subunit methyltransferase required for sequential nucleosomal histone H3 methylation by the mixed lineage leukemia protein-1 (MLL1) core complex. J Biol Chem. 2011;286:3359–69.
Zhao S, Allis CD, Wang GG. The language of chromatin modification in human cancers. Nat Rev Cancer. 2021;21:413–30.
Chi P, Allis CD, Wang GG. Covalent histone modifications-miswritten, misinterpreted and mis-erased in human cancers. Nat Rev Cancer. 2010;10:457–69.
Dawson MA. The cancer epigenome: concepts, challenges, and therapeutic opportunities. Science. 2017;355:1147–52.
Roguev A, Schaft D, Shevchenko A, Pijnappel WW, Wilm M, Aasland R, et al. The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J. 2001;20:7137–48.
Dobrovolsky D, Wang ES, Morrow S, Leahy C, Faust T, Nowak RP, et al. Bruton tyrosine kinase degradation as a therapeutic strategy for cancer. Blood. 2019;133:952–61.
Ciceri P, Muller S, O’Mahony A, Fedorov O, Filippakopoulos P, Hunt JP, et al. Dual kinase-bromodomain inhibitors for rationally designed polypharmacology. Nat Chem Biol. 2014;10:305–12.
Romanelli A, Stazi G, Fioravanti R, Zwergel C, Di Bello E, Pomella S, et al. Design of first-in-class dual EZH2/HDAC inhibitor: biochemical activity and biological evaluation in cancer cells. ACS Med Chem Lett. 2020;11:977–83.
Jin J, Wang G, Liu J, Yu X, Li D. WD40 REPEAT DOMAIN PROTEIN 5 (WDR5) DEGRADATION/DISRUPTION COMPOUNDS AND METHODS OF USE. WO2019246570; 2019.
Gadd MS, Testa A, Lucas X, Chan KH, Chen W, Lamont DJ, et al. Structural basis of PROTAC cooperative recognition for selective protein degradation. Nat Chem Biol. 2017;13:514–21.
Li J, Galbo PM Jr., Gong W, Storey AJ, Tsai YH, Yu X, et al. ZMYND11-MBTD1 induces leukemogenesis through hijacking NuA4/TIP60 acetyltransferase complex and a PWWP-mediated chromatin association mechanism. Nat Commun. 2021;12:1045.
Lu R, Wang P, Parton T, Zhou Y, Chrysovergis K, Rockowitz S, et al. Epigenetic perturbations by Arg882-mutated DNMT3A potentiate aberrant stem cell gene-expression program and acute leukemia development. Cancer Cell. 2016;30:92–107.
Cai L, Tsai YH, Wang P, Wang J, Li D, Fan H, et al. ZFX mediates non-canonical oncogenic functions of the androgen receptor splice variant 7 in castrate-resistant prostate cancer. Mol cell. 2018;72:341–54. e346
Egan B, Yuan CC, Craske ML, Labhart P, Guler GD, Arnott D, et al. An alternative approach to chip-seq normalization enables detection of genome-wide changes in histone H3 lysine 27 trimethylation upon EZH2 Inhibition. PLoS ONE. 2016;11:e0166438.
Ahn JH, Davis ES, Daugird TA, Zhao S, Quiroga IY, Uryu H, et al. Phase separation drives aberrant chromatin looping and cancer development. Nature. 2021;595:591–5.
Xu J, Li L, Xiong J, DenDekker A, Ye A, Karatas H, et al. MLL1 and MLL1 fusion proteins have distinct functions in regulating leukemic transcription program. Cell Discov. 2016;2:16008.
Fan H, Lu J, Guo Y, Li D, Zhang ZM, Tsai YH, et al. BAHCC1 binds H3K27me3 via a conserved BAH module to mediate gene silencing and oncogenesis. Nat Genet. 2020;52:1384–96.
Ren Z, Ahn JH, Liu H, Tsai YH, Bhanu NV, Koss B, et al. PHF19 promotes multiple myeloma tumorigenicity through PRC2 activation and broad H3K27me3 domain formation. Blood. 2019;134:1176–89.
Li W, Notani D, Ma Q, Tanasa B, Nunez E, Chen AY, et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature. 2013;498:516–20.
Xu B, On DM, Ma A, Parton T, Konze KD, Pattenden SG, et al. Selective inhibition of EZH2 and EZH1 enzymatic activity by a small molecule suppresses MLL-rearranged leukemia. Blood. 2015;125:346–57.
Townsend EC, Murakami MA, Christodoulou A, Christie AL, Koster J, DeSouza TA, et al. The public repository of xenografts enables discovery and randomized phase II-like trials in mice. Cancer Cell. 2016;29:574–86.
Acknowledgements
We graciously thank Drs. J Bradner, W Kaelin, Y Dou, H Wen, X Shi, and H Jiang for providing reagents and cells used in the study and the Wang and Jin Laboratory members for helpful discussion and technical support. This work was supported in part by the US National Institutes of Health grants R01GM122749 (to JJ), P30CA196521 (to JJ), R01CA211336 (to GGW), R01CA215284 (to GGW), R01CA268384 (to JJ and GGW), R35GM131780 (to AKA), R24GM137786 (to AJT), P20GM121293 (to AJT), R01CA236209 (to AJT), P20GM103429 (to AJT), an NIH/Office of the Director Grant S10OD018445 (to SGM), R01GM137009 (to ZL), U54 CA217297/PRJ001 (to ZL), endowed professorships from the Icahn School of Medicine at Mount Sinai (to JJ and AKA), and grants/awards from Gabrielle’s Angel Foundation for Cancer Research (to GGW), When Everyone Survives (WES) Leukemia Research Foundation (to GGW) and UNC Lineberger Cancer Center UCRF Stimulus Initiative Grants (to GGW and LC). G.G.W. is an American Cancer Society Research Scholar, a Leukemia and Lymphoma Society Scholar, and an American Society of Hematology Scholar in Basic Science. This work utilized the NMR Spectrometer Systems at Mount Sinai acquired with funding from National Institutes of Health SIG grants 1S10OD025132 and 1S10OD028504. We thank UNC’s facilities, including High-throughput Sequencing Facility (HTSF), Bioinformatics Core, Tissue Culture Facility, Animal Studies Core, and UNC Tissue Procurement Facility, for their professional assistance in this work. The cores affiliated with the UNC Cancer Center are supported in part by the UNC Lineberger Comprehensive Cancer Center Core Support Grant P30CA016086.
Author information
Authors and Affiliations
Contributions
DL and XY led biological/genomic and chemical biology studies, respectively, under the guidance of GGW and JJ. JK conducted protein purification and ITC experiments under the supervision of AKA and JJ. AJS, SGM, RDE, and SDB performed mass spectrometry-based proteomics analyses under the supervision of AJT. ZZ and ZL performed GRO-seq and analyzed the data. WG and Y-HT conducted RNA-seq data analysis under the supervision of LC and GGW. HU and DL analyzed the ChIP-seq data. DL, XY, LC, JJ, and GGW analyzed and interpreted experimental data. GGW, JJ, and JL conceived the project. GGW and JJ organized and led the study. DL, XY, JJ, and GGW wrote the manuscript with input from all other authors.
Corresponding authors
Ethics declarations
Competing interests
JJ, GW, JL, XY, and DL are inventors of a patent application filed by the Icahn School of Medicine at Mount Sinai and University of North Carolina at Chapel Hill. The Jin laboratory received research funds from Celgene Corporation, Levo Therapeutics, Inc., Cullgen, Inc. and Cullinan Oncology, Inc. JJ is a cofounder, scientific advisory board member and equity shareholder in Cullgen, Inc. and a consultant for Cullgen, Inc., EpiCypher, Inc. and Accent Therapeutics, Inc.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, D., Yu, X., Kottur, J. et al. Discovery of a dual WDR5 and Ikaros PROTAC degrader as an anti-cancer therapeutic. Oncogene 41, 3328–3340 (2022). https://doi.org/10.1038/s41388-022-02340-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02340-8
This article is cited by
-
Luciferase- and HaloTag-based reporter assays to measure small-molecule-induced degradation pathway in living cells
Nature Protocols (2024)
-
Sequential deregulation of histone marks, chromatin accessibility and gene expression in response to PROTAC-induced degradation of ASH2L
Scientific Reports (2023)
-
SET-PP2A complex as a new therapeutic target in KMT2A (MLL) rearranged AML
Oncogene (2023)