Abstract
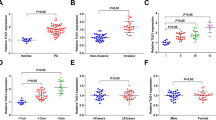
CircRNAs play important roles in a variety of biological processes by acting as microRNA sponges and protein scaffolds or by encoding functional proteins. However, their functions and underlying mechanisms remain largely unknown. Distinctive circRNA patterns were explored by comparing nonfunctioning pituitary adenomas (NFPAs) and normal pituitary tissues with a circRNA array. The biological functions of selected circRNAs were determined in vitro and in vivo. RNA-seq and circRNA pulldown assays were applied to investigate the underlying mechanisms. The circRNA profile of NFPAs is tremendously different from that of normal pituitary tissues. CircVPS13C is significantly upregulated in NFPA samples and cell lines. Gain- and loss-of-function experiments demonstrate that silencing circVPS13C inhibits the proliferation of pituitary tumor cells in vitro and in vivo. Mechanistically, circVPS13C silencing increases the expression of IFITM1 and subsequently activates its downstream genes involved in MAPK- and apoptosis-associated signaling pathways. Rescue experiments show that IFITM1 overexpression partly reverses the biological effects of circVPS13C. Further studies reveal that circVPS13C inhibits IFITM1 expression through a novel mechanism mainly by competitively interacting with RRBP1, a ribosome-binding protein of the endoplasmic reticulum membrane, and thereby alleviating the stability of IFITM1 mRNA. Clinically, circVPS13C expression is markedly higher in high-risk NFPA samples and is downregulated in patient serum 7 days post-transsphenoidal adenoma resection. Our findings suggest that circVPS13C is a critical regulator in the proliferation and development of NFPAs through a novel mechanism, whereby regulating mRNA stability via interacting with ribosome-binding proteins on the endoplasmic reticulum membrane.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Clinical information of samples and primers used in the present study can be found in supplementary tables. Other original data in our study are available upon request.
References
Agustsson TT, Baldvinsdottir T, Jonasson JG, Olafsdottir E, Steinthorsdottir V, Sigurdsson G, et al. The epidemiology of pituitary adenomas in Iceland, 1955-2012: a nationwide population-based study. Eur J Endocrinol. 2015;173:655–64.
Aflorei ED, Korbonits M. Epidemiology and etiopathogenesis of pituitary adenomas. J Neurooncol. 2014;117:379–94.
Esposito D, Olsson DS, Ragnarsson O, Buchfelder M, Skoglund T, Johannsson G. Non-functioning pituitary adenomas: indications for pituitary surgery and post-surgical management. Pituitary. 2019;22:422–34.
Mete O, Lopes MB. Overview of the 2017 WHO classification of pituitary tumors. Endocr Pathol. 2017;28:228–43.
Melmed S. Pituitary-tumor endocrinopathies. N Engl J Med. 2020; 382:937–50.
Jasim S, Alahdab F, Ahmed AT, Tamhane S, Prokop LJ, Nippoldt TB, et al. Mortality in adults with hypopituitarism: a systematic review and meta-analysis. Endocrine. 2017;56:33–42.
Song ZJ, Reitman ZJ, Ma ZY, Chen JH, Zhang QL, Shou XF, et al. The genome-wide mutational landscape of pituitary adenomas. Cell Res. 2016;26:1255–9.
Yacqub-Usman K, Richardson A, Duong CV, Clayton RN, Farrell WE. The pituitary tumour epigenome: aberrations and prospects for targeted therapy. Nat Rev Endocrinol. 2012;8:486–94.
Ng WL, Mohd MT, Shukla K. Functional role of circular RNAs in cancer development and progression. Rna Biol. 2018;15:995–1005.
Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol Cancer. 2020;19:30.
Huang A, Zheng H, Wu Z, Chen M, Huang Y. Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics. 2020;10:3503–17.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Du Q, Hu B, Feng Y, Wang Z, Wang X, Zhu D, et al. circOMA1-mediated miR-145-5p suppresses tumor growth of nonfunctioning pituitary adenomas by targeting TPT1. J Clin Endocrinol Metab. 2019;104:2419–34.
Abdelmohsen K, Panda AC, Munk R, Grammatikakis I, Dudekula DB, De S, et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol. 2017;14:361–9.
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–70.
Yang ZG, Awan FM, Du WW, Zeng Y, Lyu J, Wu D, et al. The circular RNA interacts with STAT3, increasing its nuclear translocation and wound repair by modulating Dnmt3a and miR-17 function. Mol Ther. 2017;25:2062–74.
Legnini I, Di Timoteo G, Rossi F, Morlando M, Briganti F, Sthandier O, et al. Circ-ZNF609 Is a circular RNA that can be translated and functions in myogenesis. Mol Cell. 2017;66:22–37.
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J Natl Cancer Inst. 2018;110:304–15.
Jagannathan S, Hsu JC, Reid DW, Chen Q, Thompson WJ, Moseley AM, et al. Multifunctional roles for the protein translocation machinery in RNA anchoring to the endoplasmic reticulum. J Biol Chem. 2014;289:25907–24.
Cui XA, Zhang H, Palazzo AF. p180 promotes the ribosome-independent localization of a subset of mRNA to the endoplasmic reticulum. PLoS Biol. 2012;10:e1001336.
Dhillon AS, Hagan S, Rath O, Kolch W. MAP kinase signalling pathways in cancer. Oncogene. 2007;26:3279–90.
Suojun Z, Feng W, Dongsheng G, Ting L. Targeting Raf/MEK/ERK pathway in pituitary adenomas. Eur J Cancer. 2012;48:389–95.
Yang G, Xu Y, Chen X, Hu G. IFITM1 plays an essential role in the antiproliferative action of interferon-gamma. Oncogene. 2007;26:594–603.
Hatano H, Kudo Y, Ogawa I, Tsunematsu T, Kikuchi A, Abiko Y, et al. IFN-induced transmembrane protein 1 promotes invasion at early stage of head and neck cancer progression. Clin Cancer Res. 2008;14:6097–105.
Brass AL, Huang IC, Benita Y, John SP, Krishnan MN, Feeley EM, et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, West Nile virus, and dengue virus. Cell 2009;139:1243–54.
Pan Y, Cao F, Guo A, Chang W, Chen X, Ma W, et al. Endoplasmic reticulum ribosome-binding protein 1, RRBP1, promotes progression of colorectal cancer and predicts an unfavourable prognosis. Br J Cancer. 2015;113:763–72.
Tsai HY, Yang YF, Wu AT, Yang CJ, Liu YP, Jan YH, et al. Endoplasmic reticulum ribosome-binding protein 1 (RRBP1) overexpression is frequently found in lung cancer patients and alleviates intracellular stress-induced apoptosis through the enhancement of GRP78. Oncogene. 2013;32:4921–31.
Hyde M, Block-Alper L, Felix J, Webster P, Meyer DI. Induction of secretory pathway components in yeast is associated with increased stability of their mRNA. J Cell Biol. 2002;156:993–1001.
Piovesan D, Minervini G, Tosatto SC. The RING 2.0 web server for high quality residue interaction networks. Nucleic Acids Res. 2016;44:W367–W374.
Kristensen LS, Andersen MS, Stagsted L, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–91.
Zhang L, Hou C, Chen C, Guo Y, Yuan W, Yin D, et al. The role of N(6)-methyladenosine (m(6)A) modification in the regulation of circRNAs. Mol Cancer. 2020;19:105.
Guo JU, Agarwal V, Guo H, Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014;15:409.
Abe N, Matsumoto K, Nishihara M, Nakano Y, Shibata A, Maruyama H, et al. Rolling circle translation of circular RNA in living human cells. Sci Rep. 2015;5:16435.
Chen N, Zhao G, Yan X, Lv Z, Yin H, Zhang S, et al. A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol. 2018;19:218.
Yánez DC, Ross S, Crompton T. The IFITM protein family in adaptive immunity. Immunology. 2020;159:365–72.
Grow EJ, Flynn RA, Chavez SL, Bayless NL, Wossidlo M, Wesche DJ, et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature. 2015;522:221–5.
Deblandre GA, Marinx OP, Evans SS, Majjaj S, Leo O, Caput D, et al. Expression cloning of an interferon-inducible 17-kDa membrane protein implicated in the control of cell growth. J Biol Chem. 1995;270:23860–6.
Yu F, Xie D, Ng SS, Lum CT, Cai MY, Cheung WK, et al. IFITM1 promotes the metastasis of human colorectal cancer via CAV-1. Cancer Lett. 2015;368:135–43.
Escher TE, Lui AJ, Geanes ES, Walter KR, Tawfik O, Hagan CR, et al. Interaction between MUC1 and STAT1 drives IFITM1 overexpression in aromatase inhibitor-resistant breast cancer cells and mediates estrogen-induced apoptosis. Mol Cancer Res. 2019;17:1180–94.
Yang J, Li L, Xi Y, Sun R, Wang H, Ren Y, et al. Combination of IFITM1 knockdown and radiotherapy inhibits the growth of oral cancer. Cancer Sci. 2018;109:3115–28.
Savitz AJ, Meyer DI. 180-kD ribosome receptor is essential for both ribosome binding and protein translocation. J Cell Biol. 1993;120:853–63.
Liang X, Sun S, Zhang X, Wu H, Tao W, Liu T, et al. Expression of ribosome-binding protein 1 correlates with shorter survival in Her-2 positive breast cancer. Cancer Sci. 2015;106:740–6.
Danila DC, Zhang X, Zhou Y, Dickersin GR, Fletcher JA, Hedley-Whyte ET, et al. A human pituitary tumor-derived folliculostellate cell line. J Clin Endocrinol Metab. 2000;85:1180–7.
Acknowledgements
This research was supported by the National Natural Science Foundation of China for Young Scholars (grant number 81702479 to XJ), Guangdong Basic and Applied Basic Research Foundation (grant number 2020A151501281 to XJ), Science and Technology Program of Jiangmen, China (grant number 2018630100110019805 to YM), and the Natural Science Foundation of Jiangsu Province (grant number BK20200936 to QD).
Author information
Authors and Affiliations
Contributions
WZ, SC, QD, PB, and YC carried out experiments and participated in data analysis, statistical analysis, and manuscript preparation. YM, KS, and ZC collected samples. ZL, WZ, and SC participated in data analysis. ZZ, WZ, and XJ participated in writing and proofreading the manuscript. XJ and XF designed the study, were involved in the whole process, and were guarantors of the integrity of the entire study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Informed consent, including a tissue sharing clause, was obtained from a family member of the deceased individual before the autopsy. The use of human tissues in the study was approved by the medical ethics committee of Sun Yat-sen University Cancer Center.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, W., Chen, S., Du, Q. et al. CircVPS13C promotes pituitary adenoma growth by decreasing the stability of IFITM1 mRNA via interacting with RRBP1. Oncogene 41, 1550–1562 (2022). https://doi.org/10.1038/s41388-022-02186-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02186-0
This article is cited by
-
CircOMA1 modulates cabergoline resistance by downregulating ferroptosis in prolactinoma
Journal of Endocrinological Investigation (2023)