Abstract
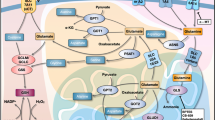
Advanced and aggressive prostate cancer (PCa) depends on glutamine for survival and proliferation. We have previously shown that inhibition of glutaminase 1, which catalyzes the rate-limiting step of glutamine catabolism, achieves significant therapeutic effect; however, therapy resistance is inevitable. Here we report that while the glutamine carbon is critical to PCa survival, a parallel pathway of glutamine nitrogen catabolism that actively contributes to pyrimidine assembly is equally important for PCa cells. Importantly, we demonstrate a reciprocal feedback mechanism between glutamine carbon and nitrogen pathways which leads to therapy resistance when one of the two pathways is inhibited. Combination treatment to inhibit both pathways simultaneously yields better clinical outcome for advanced PCa patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Rawla P. Epidemiology of Prostate Cancer. World J Oncol. 2019;10:63–89. https://doi.org/10.14740/wjon1191.
Swami U, McFarland TR, Nussenzveig R, Agarwal N, Advanced prostate cancer: treatment advances and future directions. Trends Cancer. 2020;6:702–15. https://doi.org/10.1016/j.trecan.2020.04.010.
Spetsieris N, Boukovala M, Patsakis G, Alafis I, Efstathiou E. Neuroendocrine and aggressive-variant prostate cancer. Cancers. 2020;12:ARTN 3792. https://doi.org/10.3390/cancers12123792.
Li Y, He Y, Butler W, Xu L, Chang Y, Kefeng L, et al. Targeting cellular heterogeneity with CXCR2 blockade for the treatment of therapy-resistant prostate cancer. Sci Transl Med. 2019;11. https://doi.org/10.1126/scitranslmed.aax0428.
Yin Y, Xu L, Chang Y, Zeng T, Chen X, Wang A, et al. N-Myc promotes therapeutic resistance development of neuroendocrine prostate cancer by differentially regulating miR-421/ATM pathway. Mol Cancer. 2019;18:11. https://doi.org/10.1186/s12943-019-0941-2.
Bergers G, Fendt SM. The metabolism of cancer cells during metastasis. Nat Rev Cancer. 2021. https://doi.org/10.1038/s41568-020-00320-2.
Park JH, Pyun WY, Park HW. Cancer metabolism: phenotype, signaling and therapeutic targets. Cells-Basel. 2020;9:ARTN 2308. https://doi.org/10.3390/cells9102308.
Yoo HC, Yu YC, Sung Y, Han JM. Glutamine reliance in cell metabolism. Exp Mol Med. 2020;52:1496–516. https://doi.org/10.1038/s12276-020-00504-8.
Xu L, Yin Y, Li Y, Chen X, Chang Y, Zhang H, et al. A glutaminase isoform switch drives therapeutic resistance and disease progression of prostate cancer. Proc Natl Acad Sci USA. 2021;118, https://doi.org/10.1073/pnas.2012748118.
Altman BJ, Stine ZE, Dang CV. From Krebs to clinic: glutamine metabolism to cancer therapy. Nat Rev Cancer. 2016;16:619–34. https://doi.org/10.1038/nrc.2016.71.
Biancur DE, Paulo JA, Malachowska B, Rey MQDR, Sousa CM, Wang X, et al. Compensatory metabolic networks inpancreatic cancers upon perturbation of glutamine metabolism. Nat Commun. 2017;8:15965. https://doi.org/10.1038/ncomms15965.
Yang L, Achreja A, Yeung T, Mangala L, Jiang D, Han C, et al. Targeting stromal glutamine synthetase in tumors disrupts tumor microenvironment-regulated cancer cell growth. Cell Metab. 2016;24:685–700. https://doi.org/10.1016/j.cmet.2016.10.011.
Wang Y, Bai C, Ruan Y, Liu M, Chu Q, Yang C, et al. Coordinative metabolism of glutamine carbon and nitrogen in proliferating cancer cells under hypoxia. Nat Commun. 2019;10:201. https://doi.org/10.1038/s41467-018-08033-9.
Lane AN, Fan TWM. Regulation of mammalian nucleotide metabolism and biosynthesis. Nucleic Acids Res. 2015;43:2466–85. https://doi.org/10.1093/nar/gkv047.
Oizel K, Tait-Mulder J, Fernandez-de-Cossio-Diaz J, Pietzke M, Brunton H, Lilla S, et al. Formate induces a metabolic switch in nucleotide and energy metabolism. Cell Death Dis. 2020;11:310. https://doi.org/10.1038/s41419-020-2523-z.
Bott AJ, Maimouni S, Zong WX. The pleiotropic effects of glutamine metabolism in cancer. Cancers. 2019;11:770. https://doi.org/10.3390/cancers11060770.
Zhang J, Pavlova NN, Thompson CB. Cancer cell metabolism: the essential role of the nonessential amino acid, glutamine. Embo J. 2017;36:1302–15. https://doi.org/10.15252/embj.201696151.
Cluntun AA, Lukey MJ, Cerione RA, Locasale JW. Glutamine metabolism in cancer: understanding the heterogeneity. Trends Cancer. 2017;3:169–80. https://doi.org/10.1016/j.trecan.2017.01.005.
Beltran H, Prandi D, Mosquera JM, Benelli M, Puca L, Cyrta J, et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat Med. 2016;22:298–305. https://doi.org/10.1038/nm.4045.
Beltran H, Rickman DS, Park K, Chae SS, Sboner A, MacDonald TY, et al. Molecular characterization of neuroendocrine prostate cancer and identification of new drug targets. Cancer Disco. 2011;1:487–95. https://doi.org/10.1158/2159-8290.Cd-11-0130.
Grasso CS, Wu Y, Robinson D, Cao X, Dhanasekaran SM, Khan AP, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature. 2012;487:239–43. https://doi.org/10.1038/nature11125.
Quigley DA, Dang HX, Zhao SG, Lloyd P, Aggarwal R.Alumkal P, et al. Genomic hallmarks and structural variation in metastatic prostate cancer. Cell. 2018;174:758. https://doi.org/10.1016/j.cell.2018.06.039.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver B, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22. https://doi.org/10.1016/j.ccr.2010.05.026.
Ben-Sahra I, Howell JJ, Asara JM, Manning BD. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science. 2013;339:1323–8. https://doi.org/10.1126/science.1228792.
Robitaille AM, Christen S, Shimobayashi M, Cornu M, Fava LL, Moes S, et al. Quantitative phosphoproteomics reveal mTORC1 activates de novo pyrimidine synthesis. Science. 2013;339:1320–3. https://doi.org/10.1126/science.1228771.
Crumbaker M, Khoja L, Joshua AM. AR signaling and the PI3K pathway in prostate cancer. Cancers. 2017;9:34. https://doi.org/10.3390/cancers9040034.
Chen M, Zhang J, Sampieri K, Clohessy JG, Mendez L, Gonzalez-Billalabeitia E, et al. An aberrant SREBP-dependent lipogenic program promotes metastatic prostate cancer. Nat Genet. 2018;50:206. https://doi.org/10.1038/s41588-017-0027-2.
Jamaspishvili T, Berman DM, Ross AE, Scher HI, Marzo AMD, Squire JA, et al. Clinical implications of PTEN loss in prostate cancer. Nat Rev Urol. 2018;15:222–34. https://doi.org/10.1038/nrurol.2018.9.
Park JW, Lee JK, Phillips JW, Huang P, Cheng D, Huang J, et al. Prostate epithelial cell of origin determines cancer differentiation state in an organoid transformation assay. P Natl Acad Sci USA. 2016;113:4482–7. https://doi.org/10.1073/pnas.1603645113.
Wang X, Yang K, Wu Q, Kim LJY, Morton AR, Gimple RC, et al. Targeting pyrimidine synthesis accentuates molecular therapy response in glioblastoma stem cells. Sci Transl Med. 2019;11:eaau4972. https://doi.org/10.1126/scitranslmed.aau4972.
Davidson SM, Papagiannakopoulos T, Olenchock BA, Heyman JE, Keibler MA, Luengo A, et al. Environment impacts the metabolic dependencies of ras-driven non-small cell lung cancer. Cell Metab. 2016;23:517–28. https://doi.org/10.1016/j.cmet.2016.01.007.
Kodama M, Oshikawa K, Shimizu H, Yoshioka S, Takahashi M, Izumi Y, et al. A shift in glutamine nitrogen metabolism contributes to the malignant progression of cancer. Nat Commun. 2020;11:1320. https://doi.org/10.1038/s41467-020-15136-9.
Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23:27–47. https://doi.org/10.1016/j.cmet.2015.12.006.
Wang Z, Liu F, Fan N, Zhou C, Li D, Macvicar T, et al. Targeting glutaminolysis: new perspectives to understand cancer development and novel strategies for potential target therapies. Front Oncol. 2020;10:589508. https://doi.org/10.3389/fonc.2020.589508.
Yu X, Ma R, Wu YS, Zhai YS, Li SS. Reciprocal regulation of metabolic reprogramming and epigenetic modifications in cancer. Front Genet. 2018;9:394. https://doi.org/10.3389/fgene.2018.00394.
Carver BS, Chapinski C, Wongvipat J, Hieronymus H, Chen Y, Chandarlapaty S, et al. Reciprocal feedback regulation of PI3K and androgen receptor signaling in PTEN-deficient prostate cancer. Cancer Cell. 2011;19:575–86. https://doi.org/10.1016/j.ccr.2011.04.008.
Mulholland DJ, Tran LM, Li Y, Cai H, Morim A, Wang S, et al. Cell autonomous role of PTEN in regulating castration-resistant prostate cancer growth. Cancer Cell. 2011;19:792–804. https://doi.org/10.1016/j.ccr.2011.05.006.
Liu C, Lou W, Zhu Y, Nadiminty N, Schwartz CT, Evans CP, et al. Niclosamide inhibits androgen receptor variants expression and overcomes enzalutamide resistance in castration-resistant prostate cancer. Clin Cancer Res. 2014;20:3198–210. https://doi.org/10.1158/1078-0432.CCR-13-3296.
Acknowledgements
This work was supported by DOD-W81XWH-19-1-0411 (JH), DOD-W81XWH2110034 (LX), K99-K99CA237618 (XG), Prostate Cancer Foundation Movember Valor Challenge Award (2018).
Author information
Authors and Affiliations
Contributions
LX and JH conceived and designed the study and relevant experiments. LX, BZ, WB, HX, NS, SH, HZ, DM and JH performed experiments, analyzed and interpreted the data. XG performed metabolic profiling and tracing experiments. JG conducted immunuhistochemical staining. QY and YZ performed bioinformatics and statistical analyses. XC and YH assisted animal studies. LX and JH supervised the study, interpreted the data and co-wrote the manuscript. All authors discussed the results, revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
JH is a consultant for or owns shares in the following companies: Kingmed, MoreHealth, OptraScan, Genetron, Omnitura, Vetonco, York Biotechnology, Genecode, VIVA Biotechnology and Sisu Pharma.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xu, L., Zhao, B., Butler, W. et al. Targeting glutamine metabolism network for the treatment of therapy-resistant prostate cancer. Oncogene 41, 1140–1154 (2022). https://doi.org/10.1038/s41388-021-02155-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02155-z
This article is cited by
-
Heat shock factor 1 directly regulates transsulfuration pathway to promote prostate cancer proliferation and survival
Communications Biology (2024)
-
Cancer metabolites: promising biomarkers for cancer liquid biopsy
Biomarker Research (2023)
-
Molecular and metabolic alterations of 2,3-dihydroquinazolin-4(1H)-one derivatives in prostate cancer cell lines
Scientific Reports (2022)