Abstract
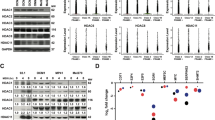
Effective therapeutic options are still lacking for uveal melanoma (UM) patients who develop metastasis. Metastatic traits of UM are linked to BRCA1-associated protein 1 (BAP1) mutations. Cell metabolism is re-programmed in UM with BAP1 mutant UM, but the underlying mechanisms and opportunities for therapeutic intervention remain unclear. BAP1 mutant UM tumors have an elevated glycolytic gene expression signature, with increased expression of pyruvate dehydrogenase (PDH) complex and PDH kinase (PDHK1). Furthermore, BAP1 mutant UM cells showed higher levels of phosphorylated PDHK1 and PDH that was associated with an upregulated glycolytic profile compared to BAP1 wild-type UM cells. Suppressing PDHK1-PDH phosphorylation decreased glycolytic capacity and cell growth, and induced cell cycle arrest of BAP1 mutant UM cells. Our results suggest that PDHK1-PDH phosphorylation is a causative factor of glycolytic phenotypes found in BAP1 mutant UM and propose a therapeutic opportunity for BAP1 mutant UM patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Krantz BA, Dave N, Komatsubara KM, Marr BP, Carvajal RD. Uveal melanoma: epidemiology, etiology, and treatment of primary disease. Clin Ophthalmol. 2017;11:279–89.
Singh N, Bergman L, Seregard S, Singh AD. Uveal melanoma: epidemiologic aspects. Ocul Oncol Pathol. 2014;18:75–84.
Chua V, Aplin AE. Novel therapeutic strategies and targets in advanced uveal melanoma. Curr Opin Oncol. 2018;30:134–41.
Heppt MV, Amaral T, Kähler KC, Heinzerling L, Hassel JC, Meissner M, et al. Combined immune checkpoint blockade for metastatic uveal melanoma: a retrospective, multi-center study. J Immunother Cancer. 2019;7:299.
Carvajal RD, Piperno-Neumann S, Kapiteijn E, Chapman PB, Frank S, Joshua AM, et al. Selumetinib in combination with dacarbazine in patients with metastatic uveal melanoma: a phase III, multicentre, randomised trial (SUMIT). J Clin Oncol. 2018;36:1232–9.
Chua V, Mattei J, Han A, Johnston L, LiPira K, Selig SM, et al. The latest on uveal melanoma research and clinical trials: updates from the cure ocular melanoma (CURE OM) science meeting (2019). Clin Cancer Res. 2021;27:28–33.
Han A, Schug ZT, Aplin AE. Metabolic alterations and therapeutic opportunities in rare forms of melanoma. Trends Cancer. 2021;7:671–81.
Nanda CS, Venkateswaran SV, Patani N, Yuneva M. Defining a metabolic landscape of tumours: genome meets metabolism. Br J Cancer. 2019:1–14.
Son J, Lyssiotis CA, Ying H, Wang X, Hua S, Ligorio M, et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature. 2013;496:101–5.
Hutton JE, Wang X, Zimmerman LJ, Slebos RJ, Trenary IA, Young JD, et al. Oncogenic KRAS and BRAF drive metabolic reprogramming in colorectal cancer. Mol Cell Proteom. 2016;15:2924–38.
Chen C-Y, Chen J, He L, Stiles BL. PTEN: tumor suppressor and metabolic regulator. Front Endocrinol. 2018;9:338.
Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA, et al. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science. 2010;330:1410–3.
Field MG, Durante MA, Anbunathan H, Cai LZ, Decatur CL, Bowcock AM, et al. Punctuated evolution of canonical genomic aberrations in uveal melanoma. Nat Commun. 2018;9:116.
Carbone M, Harbour JW, Brugarolas J, Bononi A, Pagano I, Dey A, et al. Biological mechanisms and clinical significance of BAP1 mutations in human cancer. Cancer Discov. 2020;10:1103–20.
Robertson AG, Shih J, Yau C, Gibb EA, Oba J, Mungall KL, et al. Integrative analysis identifies four molecular and clinical subsets in uveal melanoma. Cancer Cell. 2017;32:204–20.e15.
Han A, Purwin TJ, Aplin AE. Roles of the BAP1 tumor suppressor in cell metabolism. Cancer Res. 2021;81:2807–14.
Bononi A, Yang H, Giorgi C, Patergnani S, Pellegrini L, Su M, et al. Germline BAP1 mutations induce a Warburg effect. Cell Death Differ. 2017;24:1694–704.
Baughman JM, Rose CM, Kolumam G, Webster JD, Wilkerson EM, Merrill AE, et al. NeuCode proteomics reveals Bap1 regulation of metabolism. Cell Rep. 2016;16:583–95.
Zhang Y, Shi J, Liu X, Feng L, Gong Z, Koppula P, et al. BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat Cell Biol. 2018;20:1181–92.
Han A, Purwin TJ, Bechtel N, Liao C, Chua V, Seifert E, et al. BAP1 mutant uveal melanoma is stratified by metabolic phenotypes with distinct vulnerability to metabolic inhibitors. Oncogene. 2021;40:618–32.
Chua V, Han A, Bechtel N, Purwin TJ, Hunter E, Liao C, et al. The AMP-dependent kinase pathway is upregulated in BAP1 mutant uveal melanoma. Pigment Cell. Melanoma Res. 2021;00:1–10. https://doi.org/10.1111/pcmr.13007.
Nguyen GK, Mellnick VM, Yim AK-Y, Salter A, Ippolito JE. Synergy of sex differences in visceral fat measured with CT and tumor metabolism helps predict overall survival in patients with renal cell carcinoma. Radiology. 2018;287:884–92.
Chattopadhyay C, Oba J, Roszik J, Marszalek JR, Chen K, Qi Y, et al. Elevated endogenous SDHA drives pathological metabolism in highly metastatic uveal melanoma. Investig Ophthalmol Vis Sci. 2019;60:4187–95.
Saunier E, Benelli C, Bortoli S. The pyruvate dehydrogenase complex in cancer: An old metabolic gatekeeper regulated by new pathways and pharmacological agents. Int J Cancer. 2016;138:809–17.
Zhang W, Zhang S-L, Hu X, Tam KY. Targeting tumor metabolism for cancer treatment: is pyruvate dehydrogenase kinases (PDKs) a viable anticancer target? Int J Biol Sci. 2015;11:1390–400.
Li X, Jiang Y, Meisenhelder J, Yang W, Hawke DH, Zheng Y, et al. Mitochondria-translocated PGK1 functions as a protein kinase to coordinate glycolysis and the TCA cycle in tumorigenesis. Mol Cell. 2016;61:705–19.
Stacpoole PW. Therapeutic targeting of the pyruvate dehydrogenase complex/pyruvate dehydrogenase kinase (PDC/PDK) axis in cancer. J Natl Cancer Inst. 2017;109:djx071. https://doi.org/10.1093/jnci/djx071.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The molecular signatures database hallmark gene set collection. Cell Syst. 2015;1:417–25.
Zhang S, Hulver MW, McMillan RP, Cline MA, Gilbert ER. The pivotal role of pyruvate dehydrogenase kinases in metabolic flexibility. Ann Nutr Metab. 2014;11:1–9.
Zachar Z, Marecek J, Maturo C, Gupta S, Stuart SD, Howell K, et al. Non-redox-active lipoate derivates disrupt cancer cell mitochondrial metabolism and are potent anticancer agents in vivo. J Mol Med. 2011;89:1137–48.
Durante MA, Rodriguez DA, Kurtenbach S, Kuznetsov JN, Sanchez MI, Decatur CL, et al. Single-cell analysis reveals new evolutionary complexity in uveal melanoma. Nat Commun. 2020;11:1–10.
Johansson PA, Brooks K, Newell F, Palmer JM, Wilmott JS, Pritchard AL, et al. Whole genome landscapes of uveal melanoma show an ultraviolet radiation signature in iris tumours. Nat Commun. 2020;11:1–8.
Kalirai H, Dodson A, Faqir S, Damato B, Coupland S. Lack of BAP1 protein expression in uveal melanoma is associated with increased metastatic risk and has utility in routine prognostic testing. Br J Cancer. 2014;111:1373–80.
Karlsson J, Nilsson LM, Mitra S, Alsén S, Shelke GV, Sah VR, et al. Molecular profiling of driver events in metastatic uveal melanoma. Nat Commun. 2020;11:1–13.
Di Nunno V, Frega G, Santoni M, Gatto L, Fiorentino M, Montironi R, et al. BAP1 in solid tumors. Future Oncol. 2019;15:2151–62.
Wang A, Papneja A, Hyrcza M, Al-Habeeb A, Ghazarian D. Gene of the month: BAP1. J Clin Pathol. 2016;69:750–3.
Hebert L, Bellanger D, Guillas C, Campagne A, Dingli F, Loew D, et al. Modulating BAP1 expression affects ROS homeostasis, cell motility and mitochondrial function. Oncotarget. 2017;8:72513–27.
Nie H, Ju H, Fan J, Shi X, Cheng Y, Cang X, et al. O-GlcNAcylation of PGK1 coordinates glycolysis and TCA cycle to promote tumor growth. Nat Commun. 2020;11:1–14.
Forma E, Jóźwiak P, Bryś M, Krześlak A. The potential role of O-GlcNAc modification in cancer epigenetics. Cell Mol Biol Lett. 2014;19:438–60.
Ruan H-B, Han X, Li M-D, Singh JP, Qian K, Azarhoush S, et al. O-GlcNAc transferase/host cell factor C1 complex regulates gluconeogenesis by modulating PGC-1α stability. Cell Metab. 2012;16:226–37.
Kennedy BE, Murphy JP, Clements DR, Konda P, Holay N, Kim Y, et al. Inhibition of pyruvate dehydrogenase kinase enhances the antitumor efficacy of oncolytic reovirus. Cancer Res. 2019;79:3824–36.
DeBerardinis RJ, Chandel NS. Fundamentals of cancer metabolism. Sci Adv. 2016;2:e1600200.
Kreuzaler P, Panina Y, Segal J, Yuneva M. Adapt and conquer: metabolic flexibility in cancer growth, invasion and evasion. Mol Metab. 2020;33:83–101.
Shain AH, Bagger MM, Yu R, Chang D, Liu S, Vemula S, et al. The genetic evolution of metastatic uveal melanoma. Nat Genet. 2019;51:1123–30.
Eldredge-Hindy H, Ohri N, Anne PR, Eschelman D, Gonsalves C, Intenzo C, et al. Yttrium-90 microsphere brachytherapy for liver metastases from uveal melanoma: clinical outcomes and the predictive value of fluorodeoxyglucose positron emission tomography. J Clin Oncol. 2016;39:189–95.
Cai Z, Li CF, Han F, Liu C, Zhang A, Hsu CC, et al. Phosphorylation of PDHA by AMPK drives TCA cycle to promote cancer metastasis. Mol Cell. 2020;80:263–78 e7.
Wei Q, Qian Y, Yu J, Wong CC. Metabolic rewiring in the promotion of cancer metastasis: mechanisms and therapeutic implications. Oncogene. 2020;39(39):6139–56.
Pascual G, Domínguez D, Benitah SA. The contributions of cancer cell metabolism to metastasis. Dis Model Mech. 2018;11:dmm032920.
Meir T, Dror R, Yu X, Qian J, Simon I, Pe’er J, et al. Molecular characteristics of liver metastases from uveal melanoma. Investig Ophthalmol Vis Sci. 2007;48:4890–6.
Dupuy F, Tabariès S, Andrzejewski S, Dong Z, Blagih J, Annis MG, et al. PDK1-dependent metabolic reprogramming dictates metastatic potential in breast cancer. Cell Metab. 2015;22:577–89.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Mootha VK, Lindgren CM, Eriksson K-F, Subramanian A, Sihag S, Lehar J, et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–73.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. PNAS. 2005;102:15545–50.
Liu J, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, et al. An integrated TCGA Pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell. 2018;173:400–16 e11.
Attrill H, Gaudet P, Huntley RP, Lovering RC, Engel SR, Poux S, et al. Annotation of gene product function from high-throughput studies using the Gene Ontology. Database. 2019;2019.
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36:411–20.
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM III, et al. Comprehensive integration of single-cell data. Cell. 2019;177(7):1888–902. e21.
Chua V, Orloff M, Teh JL, Sugase T, Liao C, Purwin TJ, et al. Stromal fibroblast growth factor 2 reduces the efficacy of bromodomain inhibitors in uveal melanoma. EMBO Mol Med. 2019;11:e9081.
Lapadula D, Farias E, Randolph CE, Purwin TJ, McGrath D, Charpentier TH, et al. Effects of Oncogenic Galphaq and Galpha11 inhibition by FR900359 in uveal melanoma. Mol Cancer Res. 2019;17:963–73.
Amirouchene-Angelozzi N, Nemati F, Gentien D, Nicolas A, Dumont A, Carita G, et al. Establishment of novel cell lines recapitulating the genetic landscape of uveal melanoma and preclinical validation of mTOR as a therapeutic target. Mol Oncol. 2014;8:1508–20.
Tibes R, Qiu Y, Lu Y, Hennessy B, Andreeff M, Mills GB, et al. Reverse phase protein array: validation of a novel proteomic technology and utility for analysis of primary leukemia specimens and hematopoietic stem cells. Mol Cancer Ther. 2006;5:2512–21.
Cheng H, Chua V, Liao C, Purwin TJ, Terai M, Kageyama K, et al. Co-targeting HGF/cMET signaling with MEK inhibitors in metastatic uveal melanoma. Mol Cancer Ther. 2017;16:516–28.
Agarwal E, Altman BJ, Seo JH, Ghosh JC, Kossenkov AV, Tang H-Y, et al. Myc-mediated transcriptional regulation of the mitochondrial chaperone TRAP1 controls primary and metastatic tumor growth. J Biol Chem. 2019;294:10407–14.
Li J, Agarwal E, Bertolini I, Seo JH, Caino MC, Ghosh JC, et al. The mitophagy effector FUNDC1 controls mitochondrial reprogramming and cellular plasticity in cancer cells. Sci Signal. 2020;13:eaaz8240.
Acknowledgements
The Cancer Center Support Grant 5P30CA056036-17 to the SKCC supported the Microscopy Shared Resource facility. We thank Dr. Bruce Ksander (Schepens Eye Research Institute, Boston, MA), Dr. Martine Jager (Leiden University, Leiden, The Netherlands), and Dr. Sergio Roman-Roman (Institute Curie, Paris, France) for cell lines. We are grateful to Dr. Michael Durante (University of Miami Miller School of Medicine, Miami, FL) for his help with single-cell RNA-Seq data and Dr. Hsin-Yao Tang (The Wistar Institute, Philadelphia, PA) for metabolomic analyses. This work was supported by National Institutes of Health (NIH)/National Cancer Institute (NCI) grants R01 CA196278, R01 CA253977, and P01 CA114046 to AEA. This work also was supported by a Melanoma Research Alliance team science award (#559058) to AEA and JWH. The Wistar Proteomics and Metabolomics Facility was supported by P30CA010815, P01CA140043, and S10OD023586. Further support was from the American Association for Cancer Research (AACR)/Ocular Melanoma Foundation (OMF) awarded to AH.
Author information
Authors and Affiliations
Contributions
Conceptualization: AH; Formal analysis: AH, UB, and TJP; Investigations and interpretations: AH, TJP, UB, NB, VC, EH, MT, and ZTS; Resources: AH, ES, DWS, and JWH; Writing (original draft): AH; Writing (review and editing): AH, VC, UB, ES, ZTS, DWS JWH, and AEA; Funding acquisition: AH, DWS, JWH, and AEA; Supervision: AEA.
Corresponding author
Ethics declarations
Competing interests
AEA reports receiving a commercial research grant from Pfizer Inc. (2013–2017) and has ownership interest in patent number 9880150. JWH is the inventor of intellectual property related to prognostic testing for uveal melanoma. He is a paid consultant for Castle Biosciences, licensee of this intellectual property, and he receives royalties from its commercialization. The other authors disclose no potential conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Han, A., Chua, V., Baqai, U. et al. Pyruvate dehydrogenase inactivation causes glycolytic phenotype in BAP1 mutant uveal melanoma. Oncogene 41, 1129–1139 (2022). https://doi.org/10.1038/s41388-021-02154-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02154-0
This article is cited by
-
RNF40 epigenetically modulates glycolysis to support the aggressiveness of basal-like breast cancer
Cell Death & Disease (2023)