Abstract
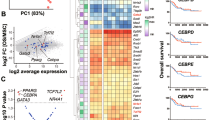
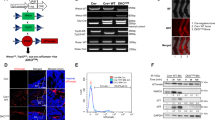
Osteosarcoma (OS) in human patients is characterized by genetic alteration of TP53. Osteoprogenitor-specific p53-deleted mice (OS mice) have been widely used to study the process of osteosarcomagenesis. However, the molecular mechanisms responsible for the development of OS upon p53 inactivation remain largely unknown. In this study, we detected prominent RUNX3/Runx3 expression in human and mouse p53-deficient OS. Myc was aberrantly upregulated by Runx3 via mR1, a consensus Runx site in the Myc promoter, in a manner dependent on p53 deficiency. Reduction of the Myc level by disruption of mR1 or Runx3 knockdown decreased the tumorigenicity of p53-deficient OS cells and effectively suppressed OS development in OS mice. Furthermore, Runx inhibitors exerted therapeutic effects on OS mice. Together, these results show that p53 deficiency promotes osteosarcomagenesis in human and mouse by allowing Runx3 to induce oncogenic Myc expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available in the main text or the supplementary materials. The ChIP-seq/ATAC-seq and RNA-seq data generated in this study were submitted to DDJB sequence read archive (DRA) with accession numbers DRA009517 and DRA011168, respectively.
References
Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA, Golub TR, et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature. 2014;505:495–501.
Bouaoun L, Sonkin D, Ardin M, Hollstein M, Byrnes G, Zavadil J, et al. TP53 Variations in Human Cancers: New Lessons from the IARC TP53 Database and Genomics Data. Hum Mutat. 2016;37:865–76.
Muller PAJ, Vousden KH. Mutant p53 in Cancer: new functions and therapeutic opportunities. Cancer Cell. 2014;25:304–17.
Kansara M, Teng MW, Smyth MJ, Thomas DM. Translational biology of osteosarcoma. Nat Rev Cancer. 2014;14:722–35.
Porter DE, Holden ST, Steel CM, Cohen BB, Wallace MR, Reid R. A significant proportion of patients with osteosarcoma may belong to Li-Fraumeni cancer families. J Bone Jt Surg Br Vol. 1992;74:883–6.
Bougeard G, Renaux-Petel M, Flaman J-M, Charbonnier C, Fermey P, Belotti M, et al. Revisiting Li-Fraumeni Syndrome From TP53 Mutation Carriers. J Clin Oncol. 2015;33:2345–52.
Chen X, Bahrami A, Pappo A, Easton J, Dalton J, Hedlund E, et al. Recurrent somatic structural variations contribute to tumorigenesis in pediatric osteosarcoma. Cell Rep. 2014;7:104–12.
Walkley CR, Qudsi R, Sankaran VG, Perry JA, Gostissa M, Roth SI, et al. Conditional mouse osteosarcoma, dependent on p53 loss and potentiated by loss of Rb, mimics the human disease. Gene Dev. 2008;22:1662–76.
Berman SD, Calo E, Landman AS, Danielian PS, Miller ES, West JC, et al. Metastatic osteosarcoma induced by inactivation of Rb and p53 in the osteoblast lineage. Proc Natl Acad Sci. 2008;105:11851–6.
Lee TI, Young RA. Transcriptional Regulation and Its Misregulation in Disease. Cell. 2013;152:1237–51.
Grigoriadis A, Schellander K, Wang Z, Wagner E. Osteoblasts are target cells for transformation in c-fos transgenic mice. J Cell Biol. 1993;122:685–701.
Gamberi G, Benassi MS, Bohling T, Ragazzini P, Molendini L, Sollazzo MR, et al. C-myc and c-fos in Human Osteosarcoma: Prognostic Value of mRNA and Protein Expression. Oncology. 1998;55:556–63.
Ito Y, Bae S-C, Chuang LSH. The RUNX family: developmental regulators in cancer. Nat Rev Cancer. 2015;15:81–95.
Levanon D, Groner Y. Structure and regulated expression of mammalian RUNX genes. Oncogene. 2004;23:4211–9.
Bauer O, Sharir A, Kimura A, Hantisteanu S, Takeda S, Groner Y. Loss of Osteoblast Runx3 Produces Severe Congenital Osteopenia. Mol Cell Biol. 2015;35:1097–109.
Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, et al. Targeted Disruption of Cbfa1 Results in a Complete Lack of Bone Formation owing to Maturational Arrest of Osteoblasts. Cell. 1997;89:755–64.
Jain M, Arvanitis C, Chu K, Dewey W, Leonhardt E, Trinh M, et al. Sustained Loss of a Neoplastic Phenotype by Brief Inactivation of MYC. Science. 2002;297:102–4.
Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, et al. CRISPR-Mediated Modular RNA-Guided Regulation of Transcription in Eukaryotes. Cell. 2013;154:442–51.
Pulikkan JA, Hegde M, Ahmad HM, Belaghzal H, Illendula A, Yu J, et al. CBFβ-SMMHC Inhibition Triggers Apoptosis by Disrupting MYC Chromatin Dynamics in Acute Myeloid Leukemia. Cell. 2018;174:172–.e21.
Cho SW, Xu J, Sun R, Mumbach MR, Carter AC, Chen YG, et al. Promoter of lncRNA Gene PVT1 Is a Tumor-Suppressor DNA Boundary Element. Cell. 2018;173:1398–.e22.
Wierstra I, Alves J. The c‐myc Promoter: Still MysterY and Challenge. Adv Cancer Res. 2008;99:113–333.
Cunningham L, Finckbeiner S, Hyde RK, Southall N, Marugan J, Yedavalli VRK, et al. Identification of benzodiazepine Ro5-3335 as an inhibitor of CBF leukemia through quantitative high throughput screen against RUNX1–CBFβ interaction. Proc Natl Acad Sci. 2012;109:14592–7.
Illendula A, Gilmour J, Grembecka J, Tirumala VSS, Boulton A, Kuntimaddi A, et al. Small Molecule Inhibitor of CBFβ-RUNX Binding for RUNX Transcription Factor Driven Cancers. Ebiomedicine. 2016;8:117–31.
Ottaviano L, Schaefer K, Gajewski M, Huckenbeck W, Baldus S, Rogel U, et al. Molecular characterization of commonly used cell lines for bone tumor research: A trans‐European EuroBoNet effort. Genes Chromosomes Cancer. 2010;49:40–51.
Yamada C, Ozaki T, Ando K, Suenaga Y, Inoue K, Ito Y, et al. RUNX3 Modulates DNA Damage-mediated Phosphorylation of Tumor Suppressor p53 at Ser-15 and Acts as a Co-activator for p53. J Biol Chem. 2010;285:16693–703.
Martin JW, Zielenska M, Stein GS, Wijnen AJ, van, Squire JA. The Role of RUNX2 in Osteosarcoma Oncogenesis. Sarcoma. 2010;2011:282745.
Shin MH, He Y, Marrogi E, Piperdi S, Ren L, Khanna C, et al. A RUNX2-Mediated Epigenetic Regulation of the Survival of p53 Defective Cancer Cells. Plos Genet. 2016;12:e1005884.
van der Deen M, Akech J, Lapointe D, Gupta S, Young DW, Montecino MA, et al. Genomic Promoter Occupancy of Runt-related Transcription Factor RUNX2 in Osteosarcoma Cells Identifies Genes Involved in Cell Adhesion and Motility*. J Biol Chem. 2012;287:4503–17.
He Y, de Castro LF, Shin MH, Dubois W, Yang HH, Jiang S, et al. p53 Loss Increases the Osteogenic Differentiation of Bone Marrow Stromal Cells. Stem Cells. 2015;33:1304–19.
Whittle MC, Izeradjene K, Rani PG, Feng L, Carlson MA, DelGiorno KE, et al. RUNX3 Controls a Metastatic Switch in Pancreatic Ductal Adenocarcinoma. Cell. 2015;161:1345–60.
Walz S, Lorenzin F, Morton J, Wiese KE, von Eyss B, Herold S, et al. Activation and repression by oncogenic MYC shape tumour-specific gene expression profiles. Nature. 2014;511:483–7.
Hosoi H, Niibori-Nambu A, Nah GSS, Bahirvani AG, Mok MMH, Sanda T, et al. Super-enhancers for RUNX3 are required for cell proliferation in EBV-infected B cell lines. Gene. 2021;774:145421.
Li Q-L, Ito K, Sakakura C, Fukamachi H, Inoue K, Chi X-Z, et al. Causal Relationship between the Loss of RUNX3 Expression and Gastric Cancer. Cell. 2002;109:113–24.
Ito K, Liu Q, Salto-Tellez M, Yano T, Tada K, Ida H, et al. RUNX3, A Novel Tumor Suppressor, Is Frequently Inactivated in Gastric Cancer by Protein Mislocalization. Cancer Res. 2005;65:7743–50.
Chuang LSH, Ito K, Ito Y. RUNX Proteins in Development and Cancer. Adv Exp Med Biol. 2017;962:299–320.
Bledsoe KL, McGee‐Lawrence ME, Camilleri ET, Wang X, Riester SM, van Wijnen AJ, et al. RUNX3 Facilitates Growth of Ewing Sarcoma Cells. J Cell Physiol. 2014;229:2049–56.
Bushweller JH. Targeting transcription factors in cancer — from undruggable to reality. Nat Rev Cancer. 2019;19:611–24.
Morita K, Suzuki K, Maeda S, Matsuo A, Mitsuda Y, Tokushige C, et al. Genetic regulation of the RUNX transcription factor family has antitumor effects. J Clin Investig. 2017;127:2815–28.
Date Y, Ito K. Oncogenic RUNX3: a link between p53 Deficiency and MYC Dysregulation. Mol Cells. 2020;43:176–81.
Gabay M, Li Y, Felsher DW. MYC Activation Is a Hallmark of Cancer Initiation and Maintenance. Csh Perspect Med. 2014;4:a014241.
Lancho O, Herranz D. The MYC Enhancer-ome: Long-Range Transcriptional Regulation of MYC in Cancer. Trends Cancer. 2018;4:810–22.
Chuang LSH, Ito K, Ito Y. RUNX family: Regulation and diversification of roles through interacting proteins. Int J Cancer. 2013;132:1260–71.
Sabapathy K, Lane DP. Therapeutic targeting of p53: all mutants are equal, but some mutants are more equal than others. Nat Rev Clin Oncol. 2018;15:13–30.
Taniuchi I, Osato M, Egawa T, Sunshine MJ, Bae S-C, Komori T, et al. Differential Requirements for Runx Proteins in CD4 Repression and Epigenetic Silencing during T Lymphocyte Development. Cell. 2002;111:621–33.
Qin X, Jiang Q, Nagano K, Moriishi T, Miyazaki T, Komori H, et al. Runx2 is essential for the transdifferentiation of chondrocytes into osteoblasts. Plos Genet. 2020;16:e1009169.
Naoe Y, Setoguchi R, Akiyama K, Muroi S, Kuroda M, Hatam F, et al. Repression of interleukin-4 in T helper type 1 cells by Runx/Cbfβ binding to the Il4 silencer. J Exp Med. 2007;204:1749–55.
Jonkers J, Meuwissen R, van der Gulden H, Peterse H, van der Valk M, Berns A. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat Genet. 2001;29:418–25.
de Alboran IM, O’Hagan RC, Gärtner F, Malynn B, Davidson L, Rickert R, et al. Analysis of C-MYC Function in Normal Cells via Conditional Gene-Targeted Mutation. Immunity. 2001;14:45–55.
Buenrostro JD, Wu B, Chang HY, Greenleaf WJ. ATAC-seq: A Method for Assaying Chromatin Accessibility Genome-Wide. Curr Protoc Mol Biol. 2015;109:21.29.1–9.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89.
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016;44:W160–5.
Bonev B, Cohen NM, Szabo Q, Fritsch L, Papadopoulos GL, Lubling Y, et al. Multiscale 3D Genome Rewiring during Mouse Neural Development. Cell. 2017;171:557–.e24.
Acknowledgements
We thank G. Huang for critical advice on the study; A. Berns and F. W. Alt for providing the p53 and Myc flox mouse lines, respectively; T. Kishino for generating genome-edited mouse lines; and all members of the Biomedical Research Center, Nagasaki University for maintaining mouse lines. This work was supported by KAKENHI/Japan Society for the Promotion of Science (JSPS) grants 26290040 (KI), 18H02972 (KI), and 19K22724 (KI); by the Funding Program for Next Generation World-Leading Researchers LS097 (KI); and by the JSPS Research Fellowship for Young Scientists 18J20543 (YD).
Author information
Authors and Affiliations
Contributions
KI initiated the study. YD and KI designed the experiments. SO, YD, TU, TI, SK, KO, JT, and KI conducted the experiments. YD performed bioinformatic analyses. IT and TK generated and provided animal materials. SO, YD, and KI wrote the paper. MU and TK coordinated the project. KI supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Otani, S., Date, Y., Ueno, T. et al. Runx3 is required for oncogenic Myc upregulation in p53-deficient osteosarcoma. Oncogene 41, 683–691 (2022). https://doi.org/10.1038/s41388-021-02120-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02120-w
This article is cited by
-
WWOX promotes osteosarcoma development via upregulation of Myc
Cell Death & Disease (2024)
-
SOX9 is a key component of RUNX2-regulated transcriptional circuitry in osteosarcoma
Cell & Bioscience (2023)
-
C/ebpα represses the oncogenic Runx3–Myc axis in p53-deficient osteosarcoma development
Oncogene (2023)
-
EZH2 regulates a SETDB1/ΔNp63α axis via RUNX3 to drive a cancer stem cell phenotype in squamous cell carcinoma
Oncogene (2022)