Abstract
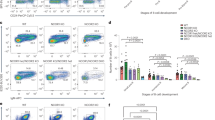
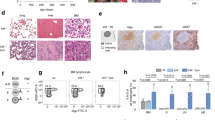
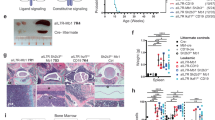
The transcription factors PAX5, IKZF1, and EBF1 are frequently mutated in B cell acute lymphoblastic leukemia (B-ALL). We demonstrate that compound heterozygous loss of multiple genes critical for B and T cell development drives transformation, including Pax5+/−xEbf1+/−, Pax5+/−xIkzf1+/−, and Ebf1+/−xIkzf1+/− mice for B-ALL, or Tcf7+/−xIkzf1+/− mice for T-ALL. To identify genetic defects that cooperate with Pax5 and Ebf1 compound heterozygosity to initiate leukemia, we performed a Sleeping Beauty (SB) transposon screen that identified cooperating partners including gain-of-function mutations in Stat5b (~65%) and Jak1 (~68%), or loss-of-function mutations in Cblb (61%) and Myb (32%). These findings underscore the role of JAK/STAT5B signaling in B cell transformation and demonstrate roles for loss-of-function mutations in Cblb and Myb in transformation. RNA-Seq studies demonstrated upregulation of a PDK1>SGK3>MYC pathway; treatment of Pax5+/−xEbf1+/− leukemia cells with PDK1 inhibitors blocked proliferation in vitro. In addition, we identified a conserved transcriptional gene signature between human and murine leukemias characterized by upregulation of myeloid genes, most notably involving the GM-CSF pathway, that resemble a B cell/myeloid mixed-lineage leukemia. Thus, our findings identify multiple mechanisms that cooperate with defects in B cell transcription factors to generate either progenitor B cell or mixed B/myeloid-like leukemias.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD, et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature. 2007;446:758–64.
Mullighan CG, Su X, Zhang J, Radtke I, Phillips LA, Miller CB, et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N Engl J Med. 2009;360:470–80.
Mullighan CG. The genomic landscape of acute lymphoblastic leukemia in children and young adults. Hematol Am Soc Hematol Educ Program. 2014;2014:174–80.
Mullighan CG, Miller CB, Radtke I, Phillips LA, Dalton J, Ma J, et al. BCR-ABL1 lymphoblastic leukaemia is characterized by the deletion of Ikaros. Nature. 2008;453:110–4.
Lin H, Grosschedl R. Failure of B-cell differentiation in mice lacking the transcription factor EBF. Nature. 1995;376:263–7.
Urbanek P, Wang ZQ, Fetka I, Wagner EF, Busslinger M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking Pax5/BSAP. Cell. 1994;79:901–12.
Chiang YJ, Kole HK, Brown K, Naramura M, Fukuhara S, Hu RJ, et al. Cbl-b regulates the CD28 dependence of T-cell activation. Nature. 2000;403:216–20.
Wang JH, Nichogiannopoulou A, Wu L, Sun L, Sharpe AH, Bigby M, et al. Selective defects in the development of the fetal and adult lymphoid system in mice with an Ikaros null mutation. Immunity. 1996;5:537–49.
Verbeek S, Izon D, Hofhuis F, Robanus-Maandag E, te Riele H, van de Wetering M, et al. An HMG-box-containing T-cell factor required for thymocyte differentiation. Nature. 1995;374:70–74.
Lee YH, Sauer B, Johnson PF, Gonzalez FJ. Disruption of the c/ebp alpha gene in adult mouse liver. Mol Cell Biol. 1997;17:6014–22.
Dupuy AJ, Rogers LM, Kim J, Nannapaneni K, Starr TK, Liu P, et al. A modified sleeping beauty transposon system that can be used to model a wide variety of human cancers in mice. Cancer Res. 2009;69:8150–6.
Temiz NA, Moriarity BS, Wolf NK, Riordan JD, Dupuy AJ, Largaespada DA, et al. RNA sequencing of Sleeping Beauty transposon-induced tumors detects transposon-RNA fusions in forward genetic cancer screens. Genome Res. 2016;26:119–29.
Beckmann PJ, Larson JD, Larsson AT, Ostergaard JP, Wagner S, Rahrmann EP, et al. Sleeping beauty insertional mutagenesis reveals important genetic drivers of central nervous system embryonal tumors. Cancer Res. 2019;79:905–17.
Scott MC, Temiz NA, Sarver AE, LaRue RS, Rathe SK, Varshney J, et al. Comparative transcriptome analysis quantifies immune cell transcript levels, metastatic progression, and survival in osteosarcoma. Cancer Res. 2018;78:326–37.
Sarver AL, Xie C, Riddle MJ, Forster CL, Wang X, Lu H, et al. Retinoblastoma tumor cell proliferation is negatively associated with an immune gene expression signature and increased immune cells. Lab Invest. 2021;101:701–18.
Heltemes-Harris LM, Willette MJ, Ramsey LB, Qiu YH, Neeley ES, Zhang N, et al. Ebf1 or Pax5 haploinsufficiency synergizes with STAT5 activation to initiate acute lymphoblastic leukemia. The. J Exp Med. 2011;208:1135–49.
Winandy S, Wu P, Georgopoulos K. A dominant mutation in the Ikaros gene leads to rapid development of leukemia and lymphoma. Cell. 1995;83:289–99.
Papathanasiou P, Perkins AC, Cobb BS, Ferrini R, Sridharan R, Hoyne GF, et al. Widespread failure of hematolymphoid differentiation caused by a recessive niche-filling allele of the Ikaros transcription factor. Immunity. 2003;19:131–44.
Heath V, Suh HC, Holman M, Renn K, Gooya JM, Parkin S, et al. C/EBPalpha deficiency results in hyperproliferation of hematopoietic progenitor cells and disrupts macrophage development in vitro and in vivo. Blood. 2004;104:1639–47.
Prasad MA, Ungerback J, Ahsberg J, Somasundaram R, Strid T, Larsson M, et al. Ebf1 heterozygosity results in increased DNA damage in pro-B cells and their synergistic transformation by Pax5 haploinsufficiency. Blood. 2015;125:4052–9.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SA, Behjati S, Biankin AV, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Katerndahl CDS, Heltemes-Harris LM, Willette MJL, Henzler CM, Frietze S, Yang R, et al. Antagonism of B cell enhancer networks by STAT5 drives leukemia and poor patient survival. Nat Immunol. 2017;18:694–704.
Castel P, Ellis H, Bago R, Toska E, Razavi P, Carmona FJ, et al. PDK1-SGK1 signaling sustains AKT-independent mTORC1 activation and confers resistance to PI3Kalpha inhibition. Cancer Cell. 2016;30:229–42.
Tan J, Li Z, Lee PL, Guan P, Aau MY, Lee ST, et al. PDK1 signaling toward PLK1-MYC activation confers oncogenic transformation, tumor-initiating cell activation, and resistance to mTOR-targeted therapy. Cancer Discov. 2013;3:1156–71.
Najafov A, Sommer EM, Axten JM, Deyoung MP, Alessi DR. Characterization of GSK2334470, a novel and highly specific inhibitor of PDK1. Biochem J. 2011;433:357–69.
Kornblau SM, Tibes R, Qiu YH, Chen W, Kantarjian HM, Andreeff M, et al. Functional proteomic profiling of AML predicts response and survival. Blood. 2009;113:154–64.
Chen EY, Tan CM, Kou Y, Duan Q, Wang Z, Meirelles GV, et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 2013;14:128.
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–97.
Xiao X, Yang G, Bai P, Gui S, Nyuyen TM, Mercado-Uribe I, et al. Inhibition of nuclear factor-kappa B enhances the tumor growth of ovarian cancer cell line derived from a low-grade papillary serous carcinoma in p53-independent pathway. BMC Cancer. 2016;16:582.
Chen F, Castranova V. Nuclear factor-kappaB, an unappreciated tumor suppressor. Cancer Res. 2007;67:11093–8.
Oliveros JC Venny. An interactive tool for comparing lists with Venn’s diagrams, 2007–2015.
van der Weyden L, Giotopoulos G, Wong K, Rust AG, Robles-Espinoza CD, Osaki H, et al. Somatic drivers of B-ALL in a model of ETV6-RUNX1; Pax5(+/−) leukemia. BMC Cancer. 2015;15:585.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008;9:559.
Kollmann S, Grundschober E, Maurer B, Warsch W, Grausenburger R, Edlinger L, et al. Twins with different personalities: STAT5B-but not STAT5A-has a key role in BCR/ABL-induced leukemia. Leukemia. 2019;33:1583–97.
Heltemes-Harris LM, Larson JD, Starr TK, Hubbard GK, Sarver AL, Largaespada DA, et al. Sleeping Beauty transposon screen identifies signaling modules that cooperate with STAT5 activation to induce B-cell acute lymphoblastic leukemia. Oncogene. 2016;35:3454–64.
Shah S, Schrader KA, Waanders E, Timms AE, Vijai J, Miething C, et al. A recurrent germline PAX5 mutation confers susceptibility to pre-B cell acute lymphoblastic leukemia. Nat Genet. 2013;45:1226–31.
Churchman ML, Qian M, Te Kronnie G, Zhang R, Yang W, Zhang H, et al. Germline genetic IKZF1 variation and predisposition to childhood acute lymphoblastic leukemia. Cancer Cell. 2018;33:937–948 e938.
Zhou J, Kryczek I, Li S, Li X, Aguilar A, Wei S, et al. The ubiquitin ligase MDM2 sustains STAT5 stability to control T cell-mediated antitumor immunity. Nat Immunol. 2021;22:460–70.
Goetz CA, Harmon IR, O’Neil JJ, Burchill MA, Farrar MA. STAT5 activation underlies IL7 receptor-dependent B cell development. J Immunol. 2004;172:4770–8.
Badger-Brown KM, Gillis LC, Bailey ML, Penninger JM, Barber DL. CBL-B is required for leukemogenesis mediated by BCR-ABL through negative regulation of bone marrow homing. Leukemia. 2013;27:1146–54.
Roberts KG, Li Y, Payne-Turner D, Harvey RC, Yang YL, Pei D, et al. Targetable kinase-activating lesions in Ph-like acute lymphoblastic leukemia. N Engl J Med. 2014;371:1005–15.
Xia Y, Shen S, Verma IM. NF-kappaB, an active player in human cancers. Cancer Immunol Res. 2014;2:823–30.
Fahl SP, Crittenden RB, Allman D, Bender TP. c-Myb is required for pro-B cell differentiation. J Immunol. 2009;183:5582–92.
Acknowledgements
We thank A. Rost, for technical assistance with mouse breeding; the University of Minnesota’s Supercomputing Institute for providing computing and bioinformatic resources; Dr. Meinrad Busslinger (Pax5+/−), Dr. Rudolf Grosschedl (Ebf1−/−), Dr. Peter Johnson (Cebpa−/−), Dr. Andrew Wells (Ikzf1−/−) and Dr. David Largaespada (Rosa26LSL-SB11T2/OncxRosa26LSL-SB11) for providing the indicated mouse strains. The results published here are in part based upon data generated by the Therapeutically Applicable Research to Generate Effective Treatments (TARGET) initiative, phs000218, managed by the NCI. This work was supported by a Cancer Research Institute Investigator award, a Leukemia and Lymphoma Society Scholar award, funding from the UMN Masonic Cancer center and grants from the NIH (RO1 CA232317) to MAF. TKS was supported by grants from the Randy Shaver Cancer Research and Community Fund, NIH NCI (R21 CA216652), and the Masonic Cancer Center. ALS was supported by NCI (CA211249) and Masonic Cancer Center Support Grant (CA077598). SMK was supported by CPRIT MIRA RP 160693 and NIH/NCI P50 CA100632-09.
Author information
Authors and Affiliations
Contributions
Conception and design: L. Heltemes-Harris and M. Farrar. Development of Methodology: L. Heltemes-Harris, A. Sarver, S. Kornblau and M. Farrar. Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.) L. Heltemes-Harris, G. Hubbard, A. Sarver, S. Kornblau and M. Farrar. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): L. Heltemes-Harris, G. Hubbard, R. LaRue, T. Starr, S. Munro, Henzler, A. R. Yang, A. Sarver, S. Kornblau and M. Farrar. Writing, review, and/or revision of the manuscript: L. Heltemes-Harris, G. Hubbard, R. LaRue, T. Starr, S. Munro, C. Henzler, R. Yang, A. Sarver, S. Kornblau and M. Farrar. Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): L. Heltemes-Harris. Study Supervision: L. Heltemes-Harris, M. Farrar.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Heltemes-Harris, L.M., Hubbard, G.K., LaRue, R.S. et al. Identification of mutations that cooperate with defects in B cell transcription factors to initiate leukemia. Oncogene 40, 6166–6179 (2021). https://doi.org/10.1038/s41388-021-02012-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-02012-z