Abstract
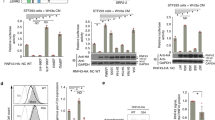
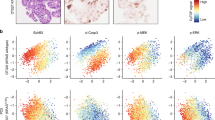
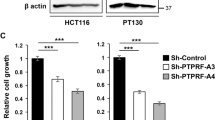
Oncogenic mutations of KRAS are found in the most aggressive human tumors, including colorectal cancer. It has been suggested that oncogenic KRAS phosphorylation at Ser181 modulates its activity and favors cell transformation. Using nonphosphorylatable (S181A), phosphomimetic (S181D), and phospho-/dephosphorylatable (S181) oncogenic KRAS mutants, we analyzed the role of this phosphorylation to the maintenance of tumorigenic properties of colorectal cancer cells. Our data show that the presence of phospho-/dephosphorylatable oncogenic KRAS is required for preserving the epithelial organization of colorectal cancer cells in 3D cultures, and for supporting subcutaneous tumor growth in mice. Interestingly, gene expression differed according to the phosphorylation status of KRAS. In DLD-1 cells, CTNNA1 was only expressed in phospho-/dephosphorylatable oncogenic KRAS-expressing cells, correlating with cell polarization. Moreover, lack of oncogenic KRAS phosphorylation leads to changes in expression of genes related to cell invasion, such as SERPINE1, PRSS1,2,3, and NEO1, and expression of phosphomimetic oncogenic KRAS resulted in diminished expression of genes involved in enterocyte differentiation, such as HNF4G. Finally, the analysis, in a public data set of human colorectal cancer, of the gene expression signatures associated with phosphomimetic and nonphosphorylatable oncogenic KRAS suggests that this post-translational modification regulates tumor progression in patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets generated during the current study are available in the GEO database repository: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE176276
References
Malumbres M, Barbacid M. RAS oncogenes: the first 30 years. Nat Rev Cancer. 2003;3:459–65.
Bourne HR, Sanders DA, McCormick F. The GTPase superfamily: conserved structure and molecular mechanism. Nature. 1991;349:117–27.
Hancock JF, Magee AI, Childs JE, Marshall CJ. All ras proteins are polyisoprenylated but only some are palmitoylated. Cell. 1989;57:1167–77.
Simanshu DK, Nissley DV, McCormick F. RAS proteins and their regulators in human disease. Cell. 2017;170:17–33.
Chandra A, Grecco HE, Pisupati V, Perera D, Cassidy L, Skoulidis F, et al. The GDI-like solubilizing factor PDEδ sustains the spatial organization and signalling of Ras family proteins. Nat Cell Biol. 2012;14:148–58.
Alvarez-Moya B, López-Alcalá C, Drosten M, Bachs O, Agell N. K-Ras4B phosphorylation at Ser181 is inhibited by calmodulin and modulates K-Ras activity and function. Oncogene. 2010;29:5911–22.
Stephen AG, Esposito D, Bagni RG, McCormick F. Dragging ras back in the ring. Cancer Cell. 2014;25:272–81.
Shalom-Feuerstein R, Plowman SJ, Rotblat B, Ariotti N, Tian T, Hancock JF, et al. K-ras nanoclustering is subverted by overexpression of the scaffold protein galectin-3. Cancer Res. 2008;68:6608–16.
Lopez-Alcalá C, Alvarez-Moya B, Villalonga P, Calvo M, Bachs O, Agell N. Identification of essential interacting elements in K-Ras/calmodulin binding and its role in K-Ras localization. J Biol Chem. 2008;283:10621–31.
Garrido E, Lázaro J, Jaumot M, Agell N, Rubio-Martinez J. Modeling and subtleties of K-Ras and calmodulin interaction. PLoS Comput Biol. 2018;14:1–19.
Villalonga P, López-Alcalá C, Bosch M, Chiloeches A, Rocamora N, Gil J, et al. Calmodulin binds to K-Ras, but not to H- or N-Ras, and modulates its downstream signaling. Mol Cell Biol. 2001;21:7345–54.
Barceló C, Etchin J, Mansour MR, Sanda T, Ginesta MM, Sanchez-Arévalo Lobo VJ. et al. Ribonucleoprotein HNRNPA2B1 interacts with and regulates oncogenic KRAS in pancreatic ductal adenocarcinoma cells. Gastroenterology. 2014;147:882–92.
Inder KL, Lau C, Loo D, Chaudhary N, Goodall A, Martin S, et al. Nucleophosmin and nucleolin regulate K-Ras plasma membrane interactions and MAPK signal transduction. J Biol Chem. 2009;284:28410–9.
Lee S, Jeong W, Cho Y, Cha P, Yoon J, Ro EJ, et al. β‐Catenin‐RAS interaction serves as a molecular switch for RAS degradation via GSK3β. EMBO Rep. 2018;19:e46060.
Villalonga P, López-Alcalá C, Chiloeches A, Gil J, Marais R, Bachs O, et al. Calmodulin prevents activation of Ras by PKC in 3T3 fibroblasts. J Biol Chem. 2002;277:37929–35.
Barceló C, Paco N, Beckett AJ, Alvarez-Moya B, Garrido E, Gelabert M, et al. Oncogenic K-ras segregates at spatially distinct plasma membrane signaling platforms according to its phosphorylation status. J Cell Sci. 2013;126:4553–9.
Yang MH, Nickerson S, Kim ET, Liot C, Laurent G, Spang R, et al. Regulation of RAS oncogenicity by acetylation. Proc Natl Acad Sci USA. 2012;109:10843–8.
Barcelo C, Paco N, Morell M, Alvarez-Moya B, Bota-Rabassedas N, Jaumot M, et al. Phosphorylation at Ser-181 of oncogenic KRAS is required for tumor growth. Cancer Res. 2014;74:1190–9.
Wang MT, Holderfield M, Galeas J, Delrosario R, To MD, Balmain A, et al. K-Ras promotes tumorigenicity through suppression of non-canonical Wnt signaling. Cell. 2015;163:1237–51.
Bivona TG, Quatela SE, Bodemann BO, Ahearn IM, Soskis MJ, Mor A, et al. PKC regulates a farnesyl-electrostatic switch on K-Ras that promotes its association with Bcl-XL on mitochondria and induces apoptosis. Mol Cell. 2006;21:481–93.
Sasaki AT, Carracedo A, Locasale JW, Anastasiou D, Takeuchi K, Kahoud ER, et al. Ubiquitination of K-Ras enhances activation and facilitates binding to select downstream effectors. Sci Signal. 2011;4:ra13.
Vartanian S, Bentley C, Brauer MJ, Li L, Shirasawa S, Sasazuki T, et al. Identification of mutant K-Ras-dependent phenotypes using a panel of isogenic cell lines. J Biol Chem. 2013;288:2403–13.
Shirasawa S, Furuse M, Yokoyama N, Sasazuki T. Altered growth of human colon cancer cell lines disrupted at activated Ki-ras. Science. (80-). 1993;260:85. LP – 88
Brookes MJ, Hughes S, Turner FE, Reynolds G, Sharma N, Ismail T, et al. Modulation of iron transport proteins in human colorectal carcinogenesis. Gut. 2006;55:1449–60.
Hu DG, Mackenzie PI, McKinnon RA, Meech R. Genetic polymorphisms of human UDP-glucuronosyltransferase (UGT) genes and cancer risk. Drug Metab Rev. 2016;48:47–69.
Maher DM, Gupta BK, Nagata S, Jaggi M, Chauhan SC. Mucin 13: Structure, function, and potential roles in cancer pathogenesis. Mol Cancer Res. 2011;9:531–7.
Lindeboom RG, van Voorthuijsen L, Oost KC, Rodríguez‐Colman MJ, Luna‐Velez MV, Furlan C, et al. Integrative multi‐omics analysis of intestinal organoid differentiation. Mol Syst Biol. 2018;14:e8227.
Santiago L, Daniels G, Wang D, Deng M, Lee P. Wnt signaling pathway protein LEF1 in cancer, as a biomarker for prognosis and a target for treatment. Am J Cancer Res. 2017;7:1389–406.
Hou Z, Guo K, Sun X, Hu F, Chen Q, Luo X, et al. TRIB2 functions as novel oncogene in colorectal cancer by blocking cellular senescence through AP4/p21 signaling. Mol Cancer. 2018;17:1–15.
Yamamoto H, Iku S, Adachi Y, Imsumran A, Taniguchi H, Nosho K, et al. Association of trypsin expression with tumour progression and matrilysin expression in human colorectal cancer. J Pathol. 2003;199:176–84.
Li S, Wei X, He J, Tian X, Yuan S, Sun L. Plasminogen activator inhibitor-1 in cancer research. Biomed Pharmacother. 2018;105:83–94.
Chaturvedi V, Fournier-Level A, Cooper HM, Murray MJ. Loss of Neogenin1 in human colorectal carcinoma cells causes a partial EMT and wound-healing response. Sci Rep. 2019;9:1–15.
Marisa L, de Reyniès A, Duval A, Selves J, Gaub MP, Vescovo L, et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med. 2013;10:e1001453.
Yin N, Liu Y, Khoor A, Wang X, Thompson EA, Leitges M, et al. Protein kinase Cι and Wnt/β-Cateninsignaling: alternative pathways to Kras/Trp53-Driven lung adenocarcinoma. Cancer Cell. 2019;36:156–.e7.
Mouradov D, Sloggett C, Jorissen RN, Love CG, Li S, Burgess AW, et al. Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. Cancer Res. 2014;74:3238–47.
Berg KCG, Eide PW, Eilertsen IA, Johannessen B, Bruun J, Danielsen SA, et al. Multi-omics of 34 colorectal cancer cell lines - a resource for biomedical studies. Mol Cancer. 2017;16:1–16.
Román-Fernández A, Bryant DM. Complex polarity: building multicellular tissues through apical membrane. Traffic Traffic. 2016;17:1244–61.
Maiden SL, Hardin J. The secret life of α-catenin: moonlighting in morphogenesis. J Cell Biol. 2011;195:543–52.
Compton CC. Colorectal carcinoma: diagnostic, prognostic, and molecular features. Mod Pathol. 2003;16:376–88.
Shibata H, Takano H, Ito M, Shioya H, Hirota M, Matsumoto H, et al. Alpha-Catenin is essential in intestinal adenoma formation. Proc Natl Acad Sci. 2007;104:18199–204.
Short SP, Kondo J, Smalley-Freed WG, Takeda H, Dohn MR, Powell AE, et al. p120-Catenin is an obligate haploinsufficient tumor suppressor in intestinal neoplasia. J Clin Invest. 2017;127:4462–76.
Elia AEH, Wang DC, Willis NA, Boardman AP, Hajdu I, Adeyemi RO, et al. RFWD3-dependent ubiquitination of RPA regulates repair at stalled replication forks. Mol Cell. 2015;60:280–93.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Lee GY, Kenny PA, Lee EH, Bissell MJ. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat Methods. 2007;4:359–65.
Gogarten SM, Bhangale T, Conomos MP, Laurie CA, McHugh CP, Painter I, et al. GWASTools: An R/Bioconductor package for quality control and analysis of genome-wide association studies. Bioinformatics. 2012;28:3329–31.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Cortazar AR, Torrano V, Martín-Martín N, Caro-Maldonado A, Camacho L, Hermanova I, et al. Cancertool: A visualization and representation interface to exploit cancer datasets. Cancer Res. 2018;78:6320–8.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods. 2001;25:402–8.
Humphries BA, Buschhaus JM, Chen YC, Haley HR, Qyli T, Chiang B, et al. Plasminogen activator inhibitor 1 (PAI1) promotes actin cytoskeleton reorganization and glycolytic metabolism in triple-negative breast cancer. Mol Cancer Res. 2019;17:1142–54.
Wang J, Zhang J, Xu L, Zheng Y, Ling D, Yang Z. Expression of HNF4G and its potential functions in lung cancer. Oncotarget. 2018;9:18018–28.
Acknowledgements
We thank the personnel of the Advanced Microscopy Unit of CCiT-UB (Campus Clinic) for help in setting up image acquisition and analysis. This work was supported by grants from the Ministerio de Economia y Competitividad and co funded by FEDER funds—a way to build Europe− (SAF2016-76239-R and PID2019-105483RB-I00 to NA, SAF2015-68016-R to GC), CIBERONC and the Government of Catalonia (grants 2017SGR1282 and PERIS SLT002/16/0037), and from Instituto de Salud Carlos III (grant PI17/01304 to MC); a FPU fellowship from Ministerio de Educación, Cultura y Deporte for DC, a FI fellowship from Generalitat de Catalunya for NP, and a Predoc-UB from University of Barcelona to BA; CR is supported by the Serra Húnter Program (Generalitat de Catalunya).
Author information
Authors and Affiliations
Contributions
DC and SB contributed equally to this work; DC, SB, NP, MMG, BA, and NG-S conducted the experiments and data analysis; CB, MC, JME, MB, CR, GC, MJ, and NA conducted data analysis and interpretation; TR-F, GP, and CR provided technical set-up and support. MJ and NA designed the study; MJ, DC, and NA wrote the paper. All authors read and approve the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cabot, D., Brun, S., Paco, N. et al. KRAS phosphorylation regulates cell polarization and tumorigenic properties in colorectal cancer. Oncogene 40, 5730–5740 (2021). https://doi.org/10.1038/s41388-021-01967-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01967-3