Abstract
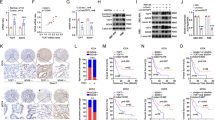
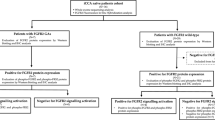
Treatment options for gallbladder carcinoma (GBC) are limited and GBC prognosis remains poor. There is no well-accepted targeted therapy to date, so effective biomarkers of GBC are urgently needed. Here we investigated the expression and correlations of fibroblast growth factor receptors (FGFR1-4) and 18 fibroblast growth factors (FGFs) in two independent patient cohorts and evaluated their prognostic significance. Consequently, we demonstrated that both FGF19 and FGFR4 were unfavorable prognostic biomarkers, and their co-expression was a more sensitive predictor. By analyzing the correlations between all 18 FGFs and FGFR4, we showed that FGF19 expression was significantly associated with FGFR4 and promoted GBC progression via stimulating FGFR4. With experiments using GBC cells, GPBAR1−/− mice models, and human subjects, we demonstrated that elevated bile acids (BAs) could increase the transcription and expression of FGF19 and FGFR4 by activating GPBAR1-cAMP-EGR1 pathway. FGF19 secreted from GBC cells promoted GBC progression by stimulating FGFR4 and downstream ERK in an autocrine manner with bile as a potential carrier. Patients with GBC had significantly higher FGF19 in serum and bile, compared to patients with cholelithiasis. BLU9931 inhibited FGFR4 and attenuated its oncogenic effects in GBC cell line. In conclusion, upregulation of BAs elevated co-expression of FGF19 and FGFR4 by activating GPBAR1-cAMP-EGR1 pathway. Co-expression of FGF19 and FGFR4 was a sensitive and unfavorable prognostic marker. GBC cells secreted FGF19 and facilitated progression by activating FGFR4 with bile as a potential carrier in an autocrine pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
13 September 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41388-023-02834-z
References
Nakamura H, Arai Y, Totoki Y, Shirota T, Elzawahry A, Kato M, et al. Genomic spectra of biliary tract cancer. Nat Genet. 2015;47:1003–10.
Xu YF, Liu ZL, Pan C, Yang XQ, Ning SL, Liu HD, et al. HMGB1 correlates with angiogenesis and poor prognosis of perihilar cholangiocarcinoma via elevating VEGFR2 of vessel endothelium. Oncogene. 2019;38:868–80.
Valle JW, Lamarca A, Goyal L, Barriuso J, Zhu AX. New horizons for precision medicine in biliary tract cancers. Cancer Discov. 2017;7:943–62.
Valle JW, Kelley RK, Nervi B, Oh DY, Zhu AX. Biliary tract cancer. Lancet. 2021;397:428–44.
Sun R, Liu Z, Qiu B, Chen T, Li Z, Zhang X, et al. Annexin10 promotes extrahepatic cholangiocarcinoma metastasis by facilitating EMT via PLA2G4A/PGE2/STAT3 pathway. EBioMedicine. 2019;47:142–55.
Mitin T, Enestvedt CK, Jemal A, Sineshaw HM. Limited use of adjuvant therapy in patients with resected gallbladder cancer despite a strong association with survival. J Natl Cancer Inst. 2017;109.
Henley SJ, Weir HK, Jim MA, Watson M, Richardson LC. Gallbladder cancer incidence and mortality, United States 1999–2011. Cancer Epidemiol, Biomark Prev. 2015;24:1319–26.
Li M, Zhang Z, Li X, Ye J, Wu X, Tan Z, et al. Whole-exome and targeted gene sequencing of gallbladder carcinoma identifies recurrent mutations in the ErbB pathway. Nat Genet. 2014;46:872–6.
Javle M, Bekaii-Saab T, Jain A, Wang Y, Kelley RK, Wang K, et al. Biliary cancer: utility of next-generation sequencing for clinical management. Cancer. 2016;122:3838–47.
Mhatre S, Wang Z, Nagrani R, Badwe R, Chiplunkar S, Mittal B, et al. Common genetic variation and risk of gallbladder cancer in India: a case–control genome-wide association study. Lancet Oncol. 2017;18:535–44.
Katoh M, Nakagama H. FGF receptors: cancer biology and therapeutics. Med Res Rev. 2014;34:280–300.
Wiedemann M, Trueb B. Characterization of a novel protein (FGFRL1) from human cartilage related to FGF receptors. Genomics. 2000;69:275–9.
Xu Y, Yang X, Li Z, Li S, Guo S, Ismail S, et al. Sprouty2 correlates with favorable prognosis of gastric adenocarcinoma via suppressing FGFR2-induced ERK phosphorylation and cancer progression. Oncotarget. 2017;8:4888–4900.
Yu C, Wang F, Kan M, Jin C, Jones RB, Weinstein M, et al. Elevated cholesterol metabolism and bile acid synthesis in mice lacking membrane tyrosine kinase receptor FGFR4. J Biol Chem. 2000;275:15482–9.
Sugiyama N, Varjosalo M, Meller P, Lohi J, Hyytiainen M, Kilpinen S, et al. Fibroblast growth factor receptor 4 regulates tumor invasion by coupling fibroblast growth factor signaling to extracellular matrix degradation. Cancer Res. 2010;70:7851–61.
Taylor JGT, Cheuk AT, Tsang PS, Chung JY, Song YK, Desai K, et al. Identification of FGFR4-activating mutations in human rhabdomyosarcomas that promote metastasis in xenotransplanted models. J Clin Investig. 2009;119:3395–407.
Ye YW, Zhou Y, Yuan L, Wang CM, Du CY, Zhou XY, et al. Fibroblast growth factor receptor 4 regulates proliferation and antiapoptosis during gastric cancer progression. Cancer. 2011;117:5304–13.
Hagel M, Miduturu C, Sheets M, Rubin N, Weng W, Stransky N, et al. First selective small molecule inhibitor of FGFR4 for the treatment of hepatocellular carcinomas with an activated FGFR4 signaling pathway. Cancer Discov. 2015;5:424–37.
Xu YF, Yang XQ, Lu XF, Guo S, Liu Y, Iqbal M, et al. Fibroblast growth factor receptor 4 promotes progression and correlates to poor prognosis in cholangiocarcinoma. Biochem Biophys Res Commun. 2014;446:54–60.
Inagaki T, Choi M, Moschetta A, Peng L, Cummins CL, McDonald JG, et al. Fibroblast growth factor 15 functions as an enterohepatic signal to regulate bile acid homeostasis. Cell Metab. 2005;2:217–25.
Choi M, Moschetta A, Bookout AL, Peng L, Umetani M, Holmstrom SR, et al. Identification of a hormonal basis for gallbladder filling. Nat Med. 2006;12:1253–5.
Elzi DJ, Song M, Blackman B, Weintraub ST, Lopez-Terrada D, Chen Y, et al. FGF19 functions as autocrine growth factor for hepatoblastoma. Genes Cancer. 2016;7:125–35.
Song KH, Li T, Owsley E, Strom S, Chiang JY. Bile acids activate fibroblast growth factor 19 signaling in human hepatocytes to inhibit cholesterol 7alpha-hydroxylase gene expression. Hepatology. 2009;49:297–305.
Zweers SJ, Booij KA, Komuta M, Roskams T, Gouma DJ, Jansen PL, et al. The human gallbladder secretes fibroblast growth factor 19 into bile: towards defining the role of fibroblast growth factor 19 in the enterobiliary tract. Hepatology. 2012;55:575–83.
Li T, Apte U. Bile acid metabolism and signaling in cholestasis, inflammation, and cancer. Adv Pharm. 2015;74:263–302.
Keitel V, Stindt J, Haussinger D. Bile acid-activated receptors: GPBAR1 (TGR5) and other G protein-coupled receptors. Handb Exp Pharmacol. 2019;256:19–49.
Yang F, Mao C, Guo L, Lin J, Ming Q, Xiao P, et al. Structural basis of GPBAR activation and bile acid recognition. Nature. (2020);587:499–504.
Li J, Dawson PA. Animal models to study bile acid metabolism. Biochim Biophys Acta Mol Basis Dis.2019;1865:895–911.
Keitel V, Haussinger D. TGR5 in cholangiocytes. Curr Opin Gastroenterol. 2013;29:299–304.
Dienstmann R, Rodon J, Prat A, Perez-Garcia J, Adamo B, Felip E, et al. Genomic aberrations in the FGFR pathway: opportunities for targeted therapies in solid tumors. Ann Oncol. 2014;25:552–63.
Qiu B, Chen T, Sun R, Liu Z, Zhang X, Li Z, et al. Sprouty4 correlates with favorable prognosis in perihilar cholangiocarcinoma by blocking the FGFR-ERK signaling pathway and arresting the cell cycle. EBioMedicine. 2019;50:166–77.
Wu YM, Su F, Kalyana-Sundaram S, Khazanov N, Ateeq B, Cao X, et al. Identification of targetable FGFR gene fusions in diverse cancers. Cancer Discov. 2013;3:636–47.
Goyal L, Saha SK, Liu LY, Siravegna G, Leshchiner I, Ahronian LG, et al. Polyclonal secondary FGFR2 mutations drive acquired resistance to FGFR inhibition in patients with FGFR2 fusion-positive cholangiocarcinoma. Cancer Discov. 2017;7:252–63.
Heinzle C, Erdem Z, Paur J, Grasl-Kraupp B, Holzmann K, Grusch M, et al. Is fibroblast growth factor receptor 4 a suitable target of cancer therapy? Curr Pharm Des. 2014;20:2881–98.
Beenken A, Mohammadi M. The FGF family: biology, pathophysiology and therapy. Nat Rev Drug Discov. 2009;8:235–53.
Zhou M, Wang X, Phung V, Lindhout DA, Mondal K, Hsu JY, et al. Separating tumorigenicity from bile acid regulatory activity for endocrine hormone FGF19. Cancer Res. 2014;74:3306–16.
Wu X, Li Y. Therapeutic utilities of fibroblast growth factor 19. Expert Opin Ther Targets. 2011;15:1307–16.
Li F, Li Z, Han Q, Cheng Y, Ji W, Yang Y, et al. Enhanced autocrine FGF19/FGFR4 signaling drives the progression of lung squamous cell carcinoma, which responds to mTOR inhibitor AZD2104. Oncogene. 2020;39:3507–21.
Tiong KH, Tan BS, Choo HL, Chung FF, Hii LW, Tan SH, et al. Fibroblast growth factor receptor 4 (FGFR4) and fibroblast growth factor 19 (FGF19) autocrine enhance breast cancer cells survival. Oncotarget. 2016;7:57633–50.
Gao L, Lang L, Zhao X, Shay C, Shull AY, Teng Y. FGF19 amplification reveals an oncogenic dependency upon autocrine FGF19/FGFR4 signaling in head and neck squamous cell carcinoma. Oncogene. 2019;38:2394–404.
Li J, Dawson PA. Animal models to study bile acid metabolism. Biochim Biophys Acta. 2019;1865:895-911.
Plevris JN, Bouchier IA. Defective acid base regulation by the gall bladder epithelium and its significance for gall stone formation. Gut. 1995;37:127–31.
Schaap FG, van der Gaag NA, Gouma DJ, Jansen PL. High expression of the bile salt-homeostatic hormone fibroblast growth factor 19 in the liver of patients with extrahepatic cholestasis. Hepatology. 2009;49:1228–35.
Reue K, Lee JM, Vergnes L. Regulation of bile acid homeostasis by the intestinal Diet1-FGF15/19 axis. Curr Opin Lipidol. 2014;25:140–7.
Tsilidis KK, Kasimis JC, Lopez DS, Ntzani EE, Ioannidis JP. Type 2 diabetes and cancer: umbrella review of meta-analyses of observational studies. BMJ. 2015;350:g7607.
Camilleri M, Malhi H, Acosta A. Gastrointestinal complications of obesity. Gastroenterology. 2017;152:1656–70.
Giovannucci E, Harlan DM, Archer MC, Bergenstal RM, Gapstur SM, Habel LA, et al. Diabetes and cancer: a consensus report. Diabetes Care. 2010;33:1674–85.
Sonne DP, van Nierop FS, Kulik W, Soeters MR, Vilsboll T, Knop FK. Postprandial plasma concentrations of individual bile acids and FGF-19 in patients with Type 2 diabetes. J Clin Endocrinol Metab. 2016;101:3002–9.
Ge H, Zhang J, Gong Y, Gupte J, Ye J, Weiszmann J, et al. Fibroblast growth factor receptor 4 (FGFR4) deficiency improves insulin resistance and glucose metabolism under diet-induced obesity conditions. J Biol Chem. 2014;289:30470–80.
Funding
Our study was supported by the National Natural Science Foundation of China (Grant No. 82072676), the China Postdoctoral Science Foundation (Grant No. 2020M682190), Shandong University Multidisciplinary Research and Innovation Team of Young Scholars (Grant No. 2020QNQT002), Shandong Province Major Research and Design Program (Grant No. 2018GSF118169), Natural Science Foundation of Shandong Province (ZR2019MH008), Jinan City Science and Technology Development Program (Grant No. 201805017, 201805013), the Major Project of Shandong University Clinical Study (Grant No. 2020SDUCRCA018), the Project of Clinical New Techniques of Qilu Hospital affiliated to Shandong Universtiy, Clinical Research Innovation Fund Project (CXPJJH11800001-2018240) and Hengrui Hepatobiliary and Pancreatic Foundation (Grant No.Y-2017-144).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, T., Liu, H., Liu, Z. et al. FGF19 and FGFR4 promotes the progression of gallbladder carcinoma in an autocrine pathway dependent on GPBAR1-cAMP-EGR1 axis. Oncogene 40, 4941–4953 (2021). https://doi.org/10.1038/s41388-021-01850-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01850-1
This article is cited by
-
FGF19 promotes nasopharyngeal carcinoma progression by inducing angiogenesis via inhibiting TRIM21-mediated ANXA2 ubiquitination
Cellular Oncology (2024)
-
Bromocriptine sensitivity in bromocriptine-induced drug-resistant prolactinomas is restored by inhibiting FGF19/FGFR4/PRL
Journal of Endocrinological Investigation (2024)
-
High expression of homeobox B2 predicts poor survival of colon adenocarcinoma by enhancing tumor proliferation and invasion
Irish Journal of Medical Science (1971 -) (2023)
-
Aberrant expression of WNK lysine deficient protein kinase 1 is associated with poor prognosis of colon adenocarcinoma
Irish Journal of Medical Science (1971 -) (2023)
-
DLGAP5 promotes gallbladder cancer migration and tumor-associated macrophage M2 polarization by activating cAMP
Cancer Immunology, Immunotherapy (2023)