Abstract
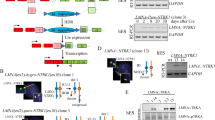
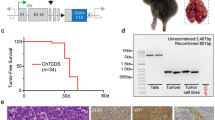
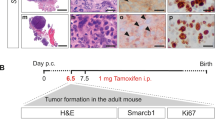
Chromosomal translocations constitute driver mutations in solid tumors and leukemias. The mechanisms of how related or even identical gene fusions drive the pathogenesis of various tumor types remain elusive. One remarkable example is the presence of EWSR1 fusions with CREB1 and ATF1, members of the CREB family of transcription factors, in a variety of sarcomas, carcinomas and mesotheliomas. To address this, we have developed in vitro models of oncogenic fusions, in particular, EWSR1-CREB1 and EWSR1-ATF1, in human embryonic stem (hES) cells, which are capable of multipotent differentiation, using CRISPR-Cas9 technology and HDR together with conditional fusion gene expression that allows investigation into the early steps of cellular transformation. We show that expression of EWSR1-CREB1/ATF1 fusion in hES cells recapitulates the core gene signatures, respectively, of angiomatoid fibrous histiocytoma (AFH) and gastrointestinal clear cell sarcoma (GI-CCS), although both fusions lead to cell lethality. Conversely, expression of the fusions in hES cells differentiated to mesenchymal progenitors is compatible with prolonged viability while maintaining the core gene signatures. Moreover, in the context of a mesenchymal lineage, the proliferation of cells expressing the EWSR1-CREB1 fusion is further extended by deletion of the tumor suppressor TP53. We expect the generation of isogenic lines carrying oncogenic fusions in various cell lineages to expand our general understanding of how those single genetic events drive tumorigenesis while providing valuable resources for drug discovery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Antonescu CR, Dal Cin P, Nafa K, Teot LA, Surti U, Fletcher CD, et al. EWSR1-CREB1 is the predominant gene fusion in angiomatoid fibrous histiocytoma. Genes Chromosomes Cancer. 2007;46:1051–60.
Antonescu CR, Tschernyavsky SJ, Woodruff JM, Jungbluth AA, Brennan MF, Ladanyi M. Molecular diagnosis of clear cell sarcoma: detection of EWS-ATF1 and MITF-M transcripts and histopathological and ultrastructural analysis of 12 cases. J Mol Diagnostics JMD. 2002;4:44–52.
Antonescu CR, Nafa K, Segal NH, Dal Cin P, Ladanyi M. EWS-CREB1: a recurrent variant fusion in clear cell sarcoma-association with gastrointestinal location and absence of melanocytic differentiation. Clin Cancer Res. 2006;12:5356–62.
Thway K, Nicholson AG, Lawson K, Gonzalez D, Rice A, Balzer B, et al. Primary pulmonary myxoid sarcoma with EWSR1-CREB1 fusion: a new tumor entity. Am J Surg Pathol. 2011;35:1722–32.
Desmeules P, Joubert P, Zhang L, Al-Ahmadie HA, Fletcher CD, Vakiani E, et al. A subset of malignant mesotheliomas in young adults are associated with recurrent EWSR1/FUS-ATF1 fusions. Am J Surg Pathol. 2017;41:980–8.
Antonescu CR, Katabi N, Zhang L, Sung YS, Seethala RR, Jordan RC, et al. EWSR1-ATF1 fusion is a novel and consistent finding in hyalinizing clear-cell carcinoma of salivary gland. Genes Chromosomes Cancer. 2011;50:559–70.
Rossi S, Szuhai K, Ijszenga M, Tanke HJ, Zanatta L, Sciot R, et al. EWSR1-CREB1 and EWSR1-ATF1 fusion genes in angiomatoid fibrous histiocytoma. Clin Cancer Res. 2007;13:7322–8.
Wang WL, Mayordomo E, Zhang W, Hernandez VS, Tuvin D, Garcia L, et al. Detection and characterization of EWSR1/ATF1 and EWSR1/CREB1 chimeric transcripts in clear cell sarcoma (melanoma of soft parts). Mod Pathol. 2009;22:1201–9.
Braunreiter CL, Hancock JD, Coffin CM, Boucher KM, Lessnick SL. Expression of EWS-ETS fusions in NIH3T3 cells reveals significant differences to Ewing’s sarcoma. Cell Cycle. 2006;5:2753–9.
Brunet E, Jasin M. Induction of chromosomal translocations with CRISPR-Cas9 and other nucleases: understanding the repair mechanisms that give rise to translocations. Adv Exp Med Biol. 2018;1044:15–25.
Vanoli F, Tomishima M, Feng W, Lamribet K, Babin L, Brunet E, et al. CRISPR-Cas9-guided oncogenic chromosomal translocations with conditional fusion protein expression in human mesenchymal cells. Proc Natl Acad Sci USA. 2017;114:3696–701.
Hisaoka M, Ishida T, Kuo TT, Matsuyama A, Imamura T, Nishida K, et al. Clear cell sarcoma of soft tissue: a clinicopathologic, immunohistochemical, and molecular analysis of 33 cases. Am J Surg Pathol. 2008;32:452–60.
Stockman DL, Miettinen M, Suster S, Spagnolo D, Dominguez-Malagon H, Hornick JL, et al. Malignant gastrointestinal neuroectodermal tumor: clinicopathologic, immunohistochemical, ultrastructural, and molecular analysis of 16 cases with a reappraisal of clear cell sarcoma-like tumors of the gastrointestinal tract. Am J Surg Pathol. 2012;36:857–68.
Karamchandani JR, Nielsen TO, van de Rijn M, West RB. Sox10 and S100 in the diagnosis of soft-tissue neoplasms. Appl Immunohistochem Mol Morphol. 2012;20:445–50.
Pierce AJ, Hu P, Han M, Ellis N, Jasin M. Ku DNA end-binding protein modulates homologous repair of double-strand breaks in mammalian cells. Genes Dev. 2001;15:3237–42.
Vriend LE, Prakash R, Chen CC, Vanoli F, Cavallo F, Zhang Y, et al. Distinct genetic control of homologous recombination repair of Cas9-induced double-strand breaks, nicks and paired nicks. Nucleic Acids Res. 2016;44:5204–17.
Jakubauskas A, Valceckiene V, Andrekute K, Seinin D, Kanopka A, Griskevicius L. Discovery of two novel EWSR1/ATF1 transcripts in four chimerical transcripts-expressing clear cell sarcoma and their quantitative evaluation. Exp Mol Pathol. 2011;90:194–200.
Panagopoulos I, Mertens F, Debiec-Rychter M, Isaksson M, Limon J, Kardas I, et al. Molecular genetic characterization of the EWS/ATF1 fusion gene in clear cell sarcoma of tendons and aponeuroses. Int J Cancer. 2002;99:560–7.
Toguchida J, Yamaguchi T, Ritchie B, Beauchamp RL, Dayton SH, Herrera GE, et al. Mutation spectrum of the p53 gene in bone and soft tissue sarcomas. Cancer Res. 1992;52:6194–9.
Tirode F, Laud-Duval K, Prieur A, Delorme B, Charbord P, Delattre O. Mesenchymal stem cell features of Ewing tumors. Cancer Cell. 2007;11:421–9.
Liang Q, Monetti C, Shutova MV, Neely EJ, Hacibekiroglu S, Yang H, et al. Linking a cell-division gene and a suicide gene to define and improve cell therapy safety. Nature. 2018;563:701–4.
Gerald WL, Rosai J, Ladanyi M. Characterization of the genomic breakpoint and chimeric transcripts in the EWS-WT1 gene fusion of desmoplastic small round cell tumor. Proc Natl Acad Sci USA. 1995;92:1028–32.
Lee SB, Kolquist KA, Nichols K, Englert C, Maheswaran S, Ladanyi M, et al. The EWS-WT1 translocation product induces PDGFA in desmoplastic small round-cell tumour. Nat Genet. 1997;17:309–13.
Bertoli C, Skotheim JM, de Bruin RA. Control of cell cycle transcription during G1 and S phases. Nat Rev Mol cell Biol. 2013;14:518–28.
Gartel AL, Tyner AL. The role of the cyclin-dependent kinase inhibitor p21 in apoptosis. Mol Cancer Ther. 2002;1:639–49.
Abbas T, Dutta A. p21 in cancer: intricate networks and multiple activities. Nat Rev Cancer. 2009;9:400–14.
Mertens F, Antonescu CR, Mitelman F. Gene fusions in soft tissue tumors: recurrent and overlapping pathogenetic themes. Genes Chromosomes Cancer. 2016;55:291–310.
Haldar M, Hancock JD, Coffin CM, Lessnick SL, Capecchi MR. A conditional mouse model of synovial sarcoma: insights into a myogenic origin. Cancer Cell. 2007;11:375–88.
Qin H, Yu T, Qing T, Liu Y, Zhao Y, Cai J, et al. Regulation of apoptosis and differentiation by p53 in human embryonic stem cells. J Biol Chem. 2007;282:5842–52.
Filion TM, Qiao M, Ghule PN, Mandeville M, van Wijnen AJ, Stein JL, et al. Survival responses of human embryonic stem cells to DNA damage. J Cell Physiol. 2009;220:586–92.
Aladjem MI, Spike BT, Rodewald LW, Hope TJ, Klemm M, Jaenisch R, et al. ES cells do not activate p53-dependent stress responses and undergo p53-independent apoptosis in response to DNA damage. Curr Biol. 1998;8:145–55.
Lin T, Chao C, Saito S, Mazur SJ, Murphy ME, Appella E, et al. p53 induces differentiation of mouse embryonic stem cells by suppressing Nanog expression. Nat Cell Biol. 2005;7:165–71.
Zilfou JT, Lowe SW. Tumor suppressive functions of p53. Cold Spring Harb Perspect Biol. 2009;1:a001883.
Straessler KM, Jones KB, Hu H, Jin H, van de Rijn M, Capecchi MR. Modeling clear cell sarcomagenesis in the mouse: cell of origin differentiation state impacts tumor characteristics. Cancer Cell. 2013;23:215–27.
Komura S, Ito K, Ohta S, Ukai T, Kabata M, Itakura F, et al. Cell-type dependent enhancer binding of the EWS/ATF1 fusion gene in clear cell sarcomas. Nat Commun. 2019;10:3999.
Hajdu M, Singer S, Maki RG, Schwartz GK, Keohan ML, Antonescu CR. IGF2 over-expression in solitary fibrous tumours is independent of anatomical location and is related to loss of imprinting. J Pathol. 2010;221:300–7.
Antonescu CR, Yoshida A, Guo T, Chang NE, Zhang L, Agaram NP, et al. KDR activating mutations in human angiosarcomas are sensitive to specific kinase inhibitors. Cancer Res. 2009;69:7175–9.
Antonescu CR, Le Loarer F, Mosquera JM, Sboner A, Zhang L, Chen CL, et al. Novel YAP1-TFE3 fusion defines a distinct subset of epithelioid hemangioendothelioma. Genes Chromosomes Cancer. 2013;52:775–84.
Feng W, Jasin M. BRCA2 suppresses replication stress-induced mitotic and G1 abnormalities through homologous recombination. Nat Commun. 2017;8:525.
Vanoli F, Jasin M. Generation of chromosomal translocations that lead to conditional fusion protein expression using CRISPR-Cas9 and homology-directed repair. Methods 2017;121-122:138–45.
Zhang Y, Vanoli F, LaRocque JR, Krawczyk PM, Jasin M. Biallelic targeting of expressed genes in mouse embryonic stem cells using the Cas9 system. Methods 2014;69:171–8.
Pfeifer A, Brandon EP, Kootstra N, Gage FH, Verma IM. Delivery of the Cre recombinase by a self-deleting lentiviral vector: efficient gene targeting in vivo. Proc Natl Acad Sci USA. 2001;98:11450–5.
Kao YC, Sung YS, Zhang L, Chen CL, Vaiyapuri S, Rosenblum MK, et al. EWSR1 fusions with CREB family transcription factors define a novel myxoid mesenchymal tumor with predilection for intracranial location. Am J Surg Pathol. 2017;41:482–90.
Acknowledgements
We thank Chew-Li Soh and Xian Zhang for technical assistance and reagents; members of the Jasin Laboratory for discussions and suggestions; and Rohit Prakash, Travis White and Pei Xin Lim for critical reading of the manuscript. This work was supported in part by: R35 CA253174 (MJ), P50 CA140146 (CRA), P50 CA217694 (CRA), Cycle for Survival (CRA), Kristin Ann Carr Foundation (CRA), St Baldrick Foundation (CRA). Core facilities at MSK are supported by the Cancer Center Support Grant (NIH P30 CA008748).
Author information
Authors and Affiliations
Contributions
FV, MJ and CRA conceived the project, designed experiments, and supervised the research. FV, BM, LH, WM, YF, SS and LZ performed experiments. YSS analyzed the Affymetrix U133A and the RNA sequencing data. DH provided reagents. FV, MJ and CRA wrote the paper with input from DH.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vanoli, F., Meskauskaite, B., Herviou, L. et al. Generation of human embryonic stem cell models to exploit the EWSR1-CREB fusion promiscuity as a common pathway of transformation in human tumors. Oncogene 40, 5095–5104 (2021). https://doi.org/10.1038/s41388-021-01843-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01843-0
This article is cited by
-
Generating in vitro models of NTRK-fusion mesenchymal neoplasia as tools for investigating kinase oncogenic activation and response to targeted therapy
Oncogenesis (2023)
-
Comprehensive genomic profiling of EWSR1/FUS::CREB translocation-associated tumors uncovers prognostically significant recurrent genetic alterations and methylation-transcriptional correlates
Modern Pathology (2022)
-
EWSR1-ATF1 dependent 3D connectivity regulates oncogenic and differentiation programs in Clear Cell Sarcoma
Nature Communications (2022)