Abstract
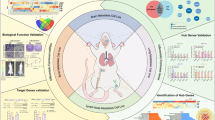
Despite the continual discovery of promising new cancer targets, drug discovery is often hampered by the poor druggability of these targets. As such, repurposing FDA-approved drugs based on cancer signatures is a useful alternative to cancer precision medicine. Here, we adopted an in silico approach based on large-scale gene expression signatures to identify drug candidates for lung cancer metastasis. Our clinicogenomic analysis identified GALNT14 as a putative driver of lung cancer metastasis, leading to poor survival. To overcome the poor druggability of GALNT14 in the control of metastasis, we utilized the Connectivity Map and identified bortezomib (BTZ) as a potent metastatic inhibitor, bypassing the direct inhibition of the enzymatic activity of GALNT14. The antimetastatic effect of BTZ was verified both in vitro and in vivo. Notably, both BTZ treatment and GALNT14 knockdown attenuated TGFβ-mediated gene expression and suppressed TGFβ-dependent metastatic genes. These results demonstrate that our in silico approach is a viable strategy for the use of undruggable targets in cancer therapies and for revealing the underlying mechanisms of these targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
All computational codes are available from the authors upon request.
Change history
11 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41388-021-01642-7
References
Spear BB, Heath-Chiozzi M, Huff J. Clinical application of pharmacogenetics. Trends Mol Med. 2001;7:201–4.
Olsen D, Jorgensen JT. Companion diagnostics for targeted cancer drugs - clinical and regulatory aspects. Front Oncol. 2014;4:105.
Kim ES, Herbst RS, Wistuba II, Lee JJ, Blumenschein GR Jr, Tsao A, et al. The BATTLE trial: personalizing therapy for lung cancer. Cancer Disco. 2011;1:44–53.
Marquart J, Chen EY, Prasad V. Estimation of the percentage of US patients with cancer who benefit from genome-driven oncology. JAMA Oncol. 2018;4:1093–8.
Lazo JS, Sharlow ER. Drugging undruggable molecular cancer targets. Annu Rev Pharmacol Toxicol. 2016;56:23–40.
Dang CV, Reddy EP, Shokat KM, Soucek L. Drugging the ‘undruggable’ cancer targets. Nat Rev Cancer. 2017;17:502–8.
Gayvert KM, Dardenne E, Cheung C, Boland MR, Lorberbaum T, Wanjala J, et al. A computational drug repositioning approach for targeting oncogenic transcription factors. Cell Rep. 2016;15:2348–56.
Nagaraj AB, Wang QQ, Joseph P, Zheng C, Chen Y, Kovalenko O, et al. Using a novel computational drug-repositioning approach (DrugPredict) to rapidly identify potent drug candidates for cancer treatment. Oncogene. 2018;37:403–14.
Kwon OS, Kim W, Cha HJ, Lee H. In silico drug repositioning: from large-scale transcriptome data to therapeutics. Arch Pharm Res. 2019;42:879–89.
Ashburn TT, Thor KB. Drug repositioning: identifying and developing new uses for existing drugs. Nat Rev Drug Discov. 2004;3:673–83.
Bertolini F, Sukhatme VP, Bouche G. Drug repurposing in oncology-patient and health systems opportunities. Nat Rev Clin Oncol. 2015;12:732–42.
Sleire L, Forde HE, Netland IA, Leiss L, Skeie BS, Enger PO. Drug repurposing in cancer. Pharmacol Res. 2017;124:74–91.
van’t Veer LJ, Bernards R. Enabling personalized cancer medicine through analysis of gene-expression patterns. Nature. 2008;452:564–70.
Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell. 2017;171:1437–e1417.
van Noort V, Scholch S, Iskar M, Zeller G, Ostertag K, Schweitzer C, et al. Novel drug candidates for the treatment of metastatic colorectal cancer through global inverse gene-expression profiling. Cancer Res. 2014;74:5690–9.
Chen B, Ma L, Paik H, Sirota M, Wei W, Chua MS, et al. Reversal of cancer gene expression correlates with drug efficacy and reveals therapeutic targets. Nature Commun. 2017;8:16022.
Lee H, Kang S, Kim W. Drug repositioning for cancer therapy based on large-scale drug-induced transcriptional signatures. PloS One. 2016;11:e0150460.
Ten Hagen KG, Fritz TA, Tabak LA. All in the family: the UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferases. Glycobiology. 2003;13:1R–16R.
Wagner KW, Punnoose EA, Januario T, Lawrence DA, Pitti RM, Lancaster K, et al. Death-receptor O-glycosylation controls tumor-cell sensitivity to the proapoptotic ligand Apo2L/TRAIL. Nat Med. 2007;13:1070–7.
Wu C, Shan Y, Liu X, Song W, Wang J, Zou M, et al. GalNAc-T14 may be involved in regulating the apoptotic action of IGFBP-3. J Biosci (Res Support, Non-U S Gov’t). 2009;34:389–95.
Song KH, Park MS, Nandu TS, Gadad S, Kim SC, Kim MY. GALNT14 promotes lung-specific breast cancer metastasis by modulating self-renewal and interaction with the lung microenvironment. Nat Commun. 2016;7:13796.
Huanna T, Tao Z, Xiangfei W, Longfei A, Yuanyuan X, Jianhua W, et al. GALNT14 mediates tumor invasion and migration in breast cancer cell MCF-7. Mol Carcinog. 2015;54:1159–71.
Wang ZQ, Bachvarova M, Morin C, Plante M, Gregoire J, Renaud MC, et al. Role of the polypeptide N-acetylgalactosaminyltransferase 3 in ovarian cancer progression: possible implications in abnormal mucin O-glycosylation. Oncotarget. 2014;5:544–60.
Kwon OS, Oh E, Park JR, Lee JS, Bae GY, Koo JH, et al. GalNAc-T14 promotes metastasis through Wnt dependent HOXB9 expression in lung adenocarcinoma. Oncotarget. 2015;6:41916–28. https://doi.org/10.18632/oncotarget.6019.
Shan J, Liu Y, Wang Y, Li Y, Yu X, Wu C. GALNT14 involves the regulation of multidrug resistance in breast cancer cells. Transl Oncol. 2018;11:786–93.
De Mariano M, Gallesio R, Chierici M, Furlanello C, Conte M, Garaventa A, et al. Identification of GALNT14 as a novel neuroblastoma predisposition gene. Oncotarget. 2015;6:26335–46.
Soria JC, Mark Z, Zatloukal P, Szima B, Albert I, Juhasz E, et al. Randomized phase II study of dulanermin in combination with paclitaxel, carboplatin, and bevacizumab in advanced non-small-cell lung cancer. J Clin Oncol. 2011;29:4442–51.
Yeh CT, Liang KH, Lin CC, Chang ML, Hsu CL, Hung CF. A single nucleotide polymorphism on the GALNT14 gene as an effective predictor of response to chemotherapy in advanced hepatocellular carcinoma. Int J Cancer. 2014;134:1214–24.
Liang KH, Lin CL, Chen SF, Chiu CW, Yang PC, Chang ML, et al. GALNT14 genotype effectively predicts the therapeutic response in unresectable hepatocellular carcinoma treated with transcatheter arterial chemoembolization. Pharmacogenomics. 2016;17:353–66.
Tsherniak A, Vazquez F, Montgomery PG, Weir BA, Kryukov G, Cowley GS, et al. Defining a cancer dependency map. Cell. 2017;170:564–76 e516.
Stern HM, Padilla M, Wagner K, Amler L, Ashkenazi A. Development of immunohistochemistry assays to assess GALNT14 and FUT3/6 in clinical trials of dulanermin and drozitumab. Clin Cancer Res (Valid Stud). 2010;16:1587–96.
Gross BJ, Swoboda JG, Walker S. A strategy to discover inhibitors of O-linked glycosylation. J Am Chem Soc. 2008;130:440–1.
Hang HC, Yu C, Ten Hagen KG, Tian E, Winans KA, Tabak LA, et al. Small molecule inhibitors of mucin-type O-linked glycosylation from a uridine-based library. Chem Biol. 2004;11:337–45.
Lee JS, Park JR, Kwon OS, Lee TH, Nakano I, Miyoshi H, et al. SIRT1 is required for oncogenic transformation of neural stem cells and for the survival of “cancer cells with neural stemness” in a p53-dependent manner. Neuro Oncol. 2015;17:95–106.
Gandolfi S, Laubach JP, Hideshima T, Chauhan D, Anderson KC, Richardson PG. The proteasome and proteasome inhibitors in multiple myeloma. Cancer Metastasis Rev. 2017;36:561–84.
Maggiora G, Vogt M, Stumpfe D, Bajorath J. Molecular similarity in medicinal chemistry. J Med Chem. 2014;57:3186–204.
Groll M, Berkers CR, Ploegh HL, Ovaa H. Crystal structure of the boronic acid-based proteasome inhibitor bortezomib in complex with the yeast 20S proteasome. Structure. 2006;14:451–6.
Massague J. How cells read TGF-beta signals. Nat Rev Mol Cell Biol (Rev). 2000;1:169–78.
Wakefield LM, Roberts AB. TGF-beta signaling: positive and negative effects on tumorigenesis. Curr Opin Genet Dev. 2002;12:22–9.
Padua D, Massague J. Roles of TGFbeta in metastasis. Cell Res. 2009;19:89–102.
Levy L, Hill CS. Smad4 dependency defines two classes of transforming growth factor {beta} (TGF-{beta}) target genes and distinguishes TGF-{beta}-induced epithelial-mesenchymal transition from its antiproliferative and migratory responses. Mol Cell Biol. 2005;25:8108–25.
Li AM, Tian AX, Zhang RX, Ge J, Sun X, Cao XC. Protocadherin-7 induces bone metastasis of breast cancer. Biochem Biophys Res Commun. 2013;436:486–90.
Moon YW, Rao G, Kim JJ, Shim HS, Park KS, An SS, et al. LAMC2 enhances the metastatic potential of lung adenocarcinoma. Cell Death Differ (Res Support, N. I H, Extramural). 2015;22:1341–52.
Zhou X, Updegraff BL, Guo Y, Peyton M, Girard L, Larsen JE, et al. Dissecting the Role of PCDH7, an Oncogenic Cell Surface Receptor, in Non–Small Cell Lung Cancer. J Thorac Oncol. 2017;12:S1542.
Huang D, Du C, Ji D, Xi J, Gu J. Overexpression of LAMC2 predicts poor prognosis in colorectal cancer patients and promotes cancer cell proliferation, migration, and invasion. Tumour Biol. 2017;39:1010428317705849.
Gupta GP, Massague J. Cancer metastasis: building a framework. Cell. 2006;127:679–95.
Jahchan NS, Dudley JT, Mazur PK, Flores N, Yang D, Palmerton A, et al. A drug repositioning approach identifies tricyclic antidepressants as inhibitors of small cell lung cancer and other neuroendocrine tumors. Cancer Disco. 2013;3:1364–77.
Zeniya M, Mori T, Yui N, Nomura N, Mandai S, Isobe K, et al. The proteasome inhibitor bortezomib attenuates renal fibrosis in mice via the suppression of TGF-beta1. Sci Rep. 2017;7:13086.
Chang TP, Poltoratsky V, Vancurova I. Bortezomib inhibits expression of TGF-beta1, IL-10, and CXCR4, resulting in decreased survival and migration of cutaneous T cell lymphoma cells. J Immunol. 2015;194:2942–53.
Padua D, Zhang XH, Wang Q, Nadal C, Gerald WL, Gomis RR, et al. TGFbeta primes breast tumors for lung metastasis seeding through angiopoietin-like 4. Cell. 2008;133:66–77.
Wagle MC, Kirouac D, Klijn C, Liu B, Mahajan S, Junttila M, et al. A transcriptional MAPK Pathway Activity Score (MPAS) is a clinically relevant biomarker in multiple cancer types. Npj Precis Oncol. 2018;2:7.
Argyriou AA, Iconomou G, Kalofonos HP. Bortezomib-induced peripheral neuropathy in multiple myeloma: a comprehensive review of the literature. Blood. 2008;112:1593–9.
Oprea TI, Overington JP. Computational and practical aspects of drug repositioning. Assay Drug Dev Technol. 2015;13:299–306.
Iorio F, Rittman T, Ge H, Menden M, Saez-Rodriguez J. Transcriptional data: a new gateway to drug repositioning? Drug Disco Today. 2013;18:350–7.
Acknowledgements
We appreciate Jeong-Hwan Kim, Seon-Young Kim and Dong-Uk Kim at Korea Research Institute of Bioscience and Biotechnology (KRIBB) for helpful discussions. This work was supported by a grant from the National Research Foundation of Korea (NRF-2017M3C9A5028691 from HJC, NRF-2019R1C1C1008710 from OSK and NRF- 2017M3C9A5028690 from WK).
Author information
Authors and Affiliations
Contributions
HJC and WK conceived the overall study design and led the experiments. OSK and HL mainly conducted the experiments, data analysis, and critical discussion of the results. HJK, JEP, EJK and WL conducted the mouse xenograft experiments. SK, JHK and MK generated and analyzed RNAseq data. All authors contributed to paper writing and revising, and endorsed the final paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kwon, OS., Lee, H., Kong, HJ. et al. Connectivity map-based drug repositioning of bortezomib to reverse the metastatic effect of GALNT14 in lung cancer. Oncogene 39, 4567–4580 (2020). https://doi.org/10.1038/s41388-020-1316-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-1316-2
This article is cited by
-
5-aminosalicylic acid suppresses osteoarthritis through the OSCAR-PPARγ axis
Nature Communications (2024)
-
Epigenetic repression of CHCHD2 enhances survival from single cell dissociation through attenuated Rho A kinase activity
Cellular and Molecular Life Sciences (2024)
-
LPCAT1 overexpression promotes the progression of hepatocellular carcinoma
Cancer Cell International (2021)