Abstract
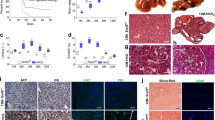
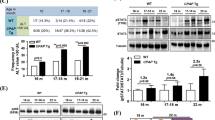
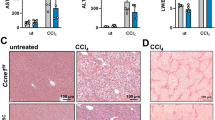
The oncofetal long noncoding RNA (lncRNA) H19 is postnatally repressed in most tissues, and re-expressed in many cancers, including hepatocellular carcinoma (HCC). The role of H19 in carcinogenesis is a subject of controversy. We aimed to examine the role of H19 in chronic inflammation-mediated hepatocarcinogenesis using the Mdr2/Abcb4 knockout (Mdr2-KO) mouse, a well-established HCC model. For this goal, we have generated Mdr2-KO/H19-KO double knockout (dKO) mice and followed spontaneous tumor development in the dKO and control Mdr2-KO mice. Cellular localization of H19 and effects of H19 loss in the liver were determined in young and old Mdr2-KO mice. Tumor incidence and tumor load were both significantly decreased in the liver of dKO versus Mdr2-KO females. The expression levels of H19 and Igf2 were variable in nontumor liver tissues of Mdr2-KO females and were significantly downregulated in most matched tumors. In nontumor liver tissue of aged Mdr2-KO females, H19 was expressed mainly in hepatocytes, and hepatocyte proliferation was increased compared to dKO females. At an early age, dKO females displayed lower levels of liver injury and B-cell infiltration, with higher percentage of binuclear hepatocytes. In human samples, H19 expression was higher in females, positively correlated with cirrhosis (in nontumor liver samples) and negatively correlated with CTNNB1 (beta-catenin) mutations and patients’ survival (in tumors). Our data demonstrate that the lncRNA H19 is pro-oncogenic during the development of chronic inflammation-mediated HCC in the Mdr2-KO mouse model, mainly by increasing liver injury and decreasing hepatocyte polyploidy in young mice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Llovet JM, Zucman-Rossi J, Pikarsky E, Sangro B, Schwartz M, Sherman M, et al. Hepatocellular carcinoma. Nat Rev Dis Prim. 2016;2:16018.
Raveh E, Matouk IJ, Gilon M, Hochberg A. The H19 long non-coding RNA in cancer initiation, progression and metastasis—a proposed unifying theory. Mol Cancer. 2015;14:184.
Yoshimizu T, Miroglio A, Ripoche MA, Gabory A, Vernucci M, Riccio A, et al. The H19 locus acts in vivo as a tumor suppressor. Proc Natl Acad Sci USA. 2008;105:12417–22.
Matouk IJ, DeGroot N, Mezan S, Ayesh S, Abu-lail R, Hochberg A, et al. The H19 non-coding RNA is essential for human tumor growth. PLoS ONE. 2007;2:e845.
Mauad TH, van Nieuwkerk CM, Dingemans KP, Smit JJ, Schinkel AH, Notenboom RG, et al. Mice with homozygous disruption of the mdr2 P-glycoprotein gene. A novel animal model for studies of nonsuppurative inflammatory cholangitis and hepatocarcinogenesis. Am J Pathol. 1994;145:1237–45.
Katzenellenbogen M, Pappo O, Barash H, Klopstock N, Mizrahi L, Olam D, et al. Multiple adaptive mechanisms to chronic liver disease revealed at early stages of liver carcinogenesis in the Mdr2-knockout mice. Cancer Res. 2006;66:4001–10.
Katzenellenbogen M, Mizrahi L, Pappo O, Klopstock N, Olam D, Jacob-Hirsch J, et al. Molecular mechanisms of liver carcinogenesis in the Mdr2-knockout mice. Mol Cancer Res. 2007;5:1159–70.
Potikha T, Stoyanov E, Pappo O, Frolov A, Mizrahi L, Olam D, et al. Interstrain differences in chronic hepatitis and tumor development in a murine model of inflammation-mediated hepatocarcinogenesis. Hepatology. 2013;58:192–204.
Ella E, Heim D, Stoyanov E, Harari-Steinfeld R, Steinfeld I, Pappo O, et al. Specific genomic and transcriptomic aberrations in tumors induced by partial hepatectomy of a chronically inflamed murine liver. Oncotarget. 2014;5:10318–31.
Martinez-Quetglas I, Pinyol R, Dauch D, Torrecilla S, Tovar V, Moeini A, et al. IGF2 Is up-regulated by epigenetic mechanisms in hepatocellular carcinomas and is an actionable oncogene product in experimental models. Gastroenterology. 2016;151:1192–205.
Yamamoto Y, Nishikawa Y, Tokairin T, Omori Y, Enomoto K. Increased expression of H19 non-coding mRNA follows hepatocyte proliferation in the rat and mouse. J Hepatol. 2004;40:808–14.
Pope C, Piekos SC, Chen L, Mishra S, Zhong XB. The role of H19, a long non-coding RNA, in mouse liver postnatal maturation. PLoS ONE. 2017;12:e0187557.
Shoshani O, Massalha H, Shani N, Kagan S, Ravid O, Madar S, et al. Polyploidization of murine mesenchymal cells is associated with suppression of the long noncoding RNA H19 and reduced tumorigenicity. Cancer Res. 2012;72:6403–13.
Guidotti JE, Bregerie O, Robert A, Debey P, Brechot C, Desdouets C. Liver cell polyploidization: a pivotal role for binuclear hepatocytes. J Biol Chem. 2003;278:19095–101.
Boyault S, Rickman DS, de Reynies A, Balabaud C, Rebouissou S, Jeannot E, et al. Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology. 2007;45:42–52.
Lecerf C, Le Bourhis X, Adriaenssens E. The long non-coding RNA H19: an active player with multiple facets to sustain the hallmarks of cancer. Cell Mol Life Sci. 2019;76:4673–87.
Li X, Liu R, Yang J, Sun L, Zhang L, Jiang Z, et al. The role of long noncoding RNA H19 in gender disparity of cholestatic liver injury in multidrug resistance 2 gene knockout mice. Hepatology. 2017;66:869–84.
Adriaenssens E, Lottin S, Dugimont T, Fauquette W, Coll J, Dupouy JP, et al. Steroid hormones modulate H19 gene expression in both mammary gland and uterus. Oncogene. 1999;18:4460–73.
Yang F, Bi J, Xue X, Zheng L, Zhi K, Hua J, et al. Up-regulated long non-coding RNA H19 contributes to proliferation of gastric cancer cells. FEBS J. 2012;279:3159–65.
Schultheiss CS, Laggai S, Czepukojc B, Hussein UK, List M, Barghash A, et al. The long non-coding RNA H19 suppresses carcinogenesis and chemoresistance in hepatocellular carcinoma. Cell Stress. 2017;1:37–54.
Jiang Y, Huang Y, Cai S, Song Y, Boyer JL, Zhang K, et al. H19 is expressed in hybrid hepatocyte nuclear factor 4alpha(+) periportal hepatocytes but not cytokeratin 19(+) cholangiocytes in cholestatic livers. Hepatol Commun. 2018;2:1356–68.
Chen X, Yamamoto M, Fujii K, Nagahama Y, Ooshio T, Xin B, et al. Differential reactivation of fetal/neonatal genes in mouse liver tumors induced in cirrhotic and non-cirrhotic conditions. Cancer Sci. 2015;106:972–81.
Uehara T, Ainslie GR, Kutanzi K, Pogribny IP, Muskhelishvili L, Izawa T, et al. Molecular mechanisms of fibrosis-associated promotion of liver carcinogenesis. Toxicol Sci. 2013;132:53–63.
Zhang Y, Liu C, Barbier O, Smalling R, Tsuchiya H, Lee S, et al. Bcl2 is a critical regulator of bile acid homeostasis by dictating Shp and lncRNA H19 function. Sci Rep. 2016;6:20559.
Faggioli F, Palagano E, Di Tommaso L, Donadon M, Marrella V, Recordati C, et al. B lymphocytes limit senescence-driven fibrosis resolution and favor hepatocarcinogenesis in mouse liver injury. Hepatology. 2018;67:1970–85.
Zhao R, Chen X, Ma W, Zhang J, Guo J, Zhong X, et al. A GPR174-CCL21 module imparts sexual dimorphism to humoral immunity. Nature. 2020;577:416–20.
Stoyanov E, Ludwig G, Mizrahi L, Olam D, Schnitzer-Perlman T, Tasika E, et al. Chronic liver inflammation modifies DNA methylation at the precancerous stage of murine hepatocarcinogenesis. Oncotarget. 2015;6:11047–60.
Zhang S, Zhou K, Luo X, Li L, Tu HC, Sehgal A, et al. The polyploid state plays a tumor-suppressive role in the liver. Dev Cell. 2018;44:447–59.e5.
Wilkinson PD, Delgado ER, Alencastro F, Leek MP, Roy N, Weirich MP, et al. The polyploid state restricts hepatocyte proliferation and liver regeneration in mice. Hepatology. 2019;69:1242–58.
Lee JW, Stone ML, Porrett PM, Thomas SK, Komar CA, Li JH, et al. Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature. 2019;567:249–52.
Nemeth J, Stein I, Haag D, Riehl A, Longerich T, Horwitz E, et al. S100A8 and S100A9 are novel nuclear factor kappa B target genes during malignant progression of murine and human liver carcinogenesis. Hepatology. 2009;50:1251–62.
Robert O, Boujedidi H, Bigorgne A, Ferrere G, Voican CS, Vettorazzi S, et al. Decreased expression of the glucocorticoid receptor-GILZ pathway in Kupffer cells promotes liver inflammation in obese mice. J Hepatol. 2016;64:916–24.
Ricci E, Ronchetti S, Gabrielli E, Pericolini E, Gentili M, Roselletti E, et al. GILZ restrains neutrophil activation by inhibiting the MAPK pathway. J Leukoc Biol. 2019;105:187–94.
Jones SA, Toh AE, Odobasic D, Oudin MA, Cheng Q, Lee JP, et al. Glucocorticoid-induced leucine zipper (GILZ) inhibits B cell activation in systemic lupus erythematosus. Ann Rheum Dis. 2016;75:739–47.
Zhu P, Wang Y, Du Y, He L, Huang G, Zhang G, et al. C8orf4 negatively regulates self-renewal of liver cancer stem cells via suppression of NOTCH2 signalling. Nat Commun. 2015;6:7122.
Goyal N, Tiwary S, Kesharwani D, Datta M. Long non-coding RNA H19 inhibition promotes hyperglycemia in mice by upregulating hepatic FoxO1 levels and promoting gluconeogenesis. J Mol Med. 2019;97:115–26.
Li X, Liu R, Huang Z, Gurley EC, Wang X, Wang J, et al. Cholangiocyte-derived exosomal long noncoding RNA H19 promotes cholestatic liver injury in mouse and humans. Hepatology. 2018;68:599–615.
Yang JB, Zhao ZB, Liu QZ, Hu TD, Long J, Yan K, et al. FoxO1 is a regulator of MHC-II expression and anti-tumor effect of tumor-associated macrophages. Oncogene. 2018;37:1192–204.
Yang XW, Shen GZ, Cao LQ, Jiang XF, Peng HP, Shen G, et al. MicroRNA-1269 promotes proliferation in human hepatocellular carcinoma via downregulation of FOXO1. BMC Cancer. 2014;14:909.
Yang N, Zhou J, Li Q, Han F, Yu Z. miR-96 exerts carcinogenic effect by activating AKT/GSK-3beta/beta-catenin signaling pathway through targeting inhibition of FOXO1 in hepatocellular carcinoma. Cancer Cell Int. 2019;19:38.
Hao PP, Li H, Lee MJ, Wang YP, Kim JH, Yu GR, et al. Disruption of a regulatory loop between DUSP1 and p53 contributes to hepatocellular carcinoma development and progression. J Hepatol. 2015;62:1278–86.
Ying J, Srivastava G, Hsieh WS, Gao Z, Murray P, Liao SK, et al. The stress-responsive gene GADD45G is a functional tumor suppressor, with its response to environmental stresses frequently disrupted epigenetically in multiple tumors. Clin Cancer Res. 2005;11:6442–9.
Xu G, Zhang L, Ma A, Qian Y, Ding Q, Liu Y, et al. SIP1 is a downstream effector of GADD45G in senescence induction and growth inhibition of liver tumor cells. Oncotarget. 2015;6:33636–47.
Hoppstadter J, Ammit AJ. Role of dual-specificity phosphatase 1 in glucocorticoid-driven anti-inflammatory responses. Front Immunol. 2019;10:1446.
Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B, et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 2008;68:6779–88.
Zhang J, Han C, Ungerleider N, Chen W, Song K, Wang Y, et al. A transforming growth factor-beta and H19 signaling axis in tumor-initiating hepatocytes that regulates hepatic carcinogenesis. Hepatology. 2019;69:1549–63.
Zhang L, Yang F, Yuan JH, Yuan SX, Zhou WP, Huo XS, et al. Epigenetic activation of the MiR-200 family contributes to H19-mediated metastasis suppression in hepatocellular carcinoma. Carcinogenesis. 2013;34:577–86.
Acknowledgements
The authors thank Sharona Elgavish, Yuval Nevo, and Hadar Benyamini (Bioinformatics Unit of the I-CORE Computation Center at The Hebrew University and Hadassah, Jerusalem, Israel) for single-cell transcriptomic (SE & YN) and gene set enrichment (HB) analyses, Zakharia Manevich (The Core Research Facility at The Faculty of Medicine, Ein Kerem, The Hebrew University, Jerusalem, Israel) for advice with confocal microscopy, Luisa Dandolo (Institut Cochin, Paris, France) for providing the 129Sv H19Δ3 mutant mice, Shalev Itzkovitz (Weizmann Institute of Science, Rehovot, Israel) for useful discussions and help with smRNA-FISH, Ilan Stein and Yoganathan Krishnamoorthy (Department of Pathology, Hebrew University-Hadassah Medical School) for advice with immunohistochemistry, and Hilla Giladi (The Goldyne Savad Institute of Gene Therapy) for critical reading of the manuscript.
Funding
Robert H. Benson Living Trust and Selma Kron Foundation to student fellowships. DSG is supported by the Kamea Scientific Foundation of the Israeli Government. JHA was supported by ISF 923/14. SC was supported by a funding from Labex OncoImunology and CARPEM. The JZR group was supported by INSERM, Ligue Nationale contre le Cancer (Equipe Labellisée), Labex OncoImmunology (investissement d’avenir), grant IREB, Coup d’Elan de la Fondation Bettencourt-Schueller, the SIRIC CARPEM, Raymond Rosen Award from the Fondation pour le Recherche Médicale, Prix René and Andrée Duquesne—Comité de Paris Ligue Contre le Cancer and Fondation Mérieux. The work of EG was supported by the: ERC advance—GA No. 786575—RxmiRcanceR, Deutsche Forschungsgemeinschaft (DFG) SFB841 project C3, NIH CA197081-02, MOST, ISF collaboration with Canada (2473/2017), personal ISF (486/2017), ICORE–ISF (41/2011), and by DKFZ-MOST.
Author information
Authors and Affiliations
Contributions
LG and LM: main part of the experimental work and discussions; TF and EZ: data acquisition; NR: data acquisition (single-cell analysis); OP: data analysis (pathology); DO: assistance with mice and liver samples processing; KBH: help with smRNA-FISH and discussions; SC and JZ-R: human HCC data analysis and discussions; JHA: help with mice, discussions, manuscript editing; EG: study idea and discussions, manuscript editing; DSG: study design, data acquisition and analysis, manuscript writing, and discussions.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Gamaev, L., Mizrahi, L., Friehmann, T. et al. The pro-oncogenic effect of the lncRNA H19 in the development of chronic inflammation-mediated hepatocellular carcinoma. Oncogene 40, 127–139 (2021). https://doi.org/10.1038/s41388-020-01513-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01513-7
This article is cited by
-
Integrated bioinformatics and validation reveal SOX12 as potential biomarker in colon adenocarcinoma based on an immune infiltration-related ceRNA network
Journal of Cancer Research and Clinical Oncology (2023)
-
LncRNA H19 is a potential biomarker and correlated with immune infiltration in thyroid carcinoma
Clinical and Experimental Medicine (2022)