Abstract
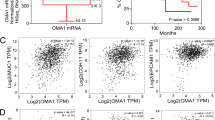
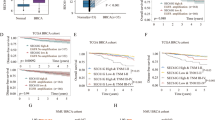
Oxidative stress-responsive kinase 1 (OSR1) plays a critical role in multiple carcinogenic signal pathways, and its overexpression has been found in various types of cancer; however, the pathophysiological role of OSR1 in breast cancer has not been evaluated. This study aims to elaborate on the role of OSR1 in breast cancer metastasis and the specific regulatory mechanism. Our results showed that OSR1 mRNA and protein were upregulated in both human breast cancer samples and cell lines. Moreover, phosphorylated OSR1 (p-OSR1) was an independent poor prognostic indicator in patients with breast cancer. OSR1 upregulation induced epithelial-to-mesenchymal transition (EMT) in normal and malignant mammary epithelial cells with the increasing metastatic capacity. In contrast, deleting OSR1 in aggressive breast cancer cells inhibited these phenotypes. OSR1 is the critical activator for transcription factors of EMT. Mechanistically, we found that OSR1 can directly interact and phosphorylate the linker region of Smad2 at Thr220 and Smad3 at Thr179. Phosphorylated Smad2/3 translocated into the nucleus to enhance transforming growth factor-β1 (TGF-β1) autocrine signalling and increase the transcription of EMT regulators. Importantly, interruption of the OSR1-Smad2/3-TGF-β1 signalling axis elicited a robust anti-EMT and anti-metastatic effect in vitro and in vivo. Taken together, we conclude that OSR1-mediated Smad2/3-TGF-β1 signalling promotes EMT and metastasis representing a promising therapeutic target in breast cancer treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
31 May 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41388-023-02736-0
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34.
Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003;3:453–8.
Massagué J, Obenauf AC. Metastatic colonization by circulating tumour cells. Nature. 2016;529:298–306.
Moustakas A, Heldin CH. The regulation of TGFbeta signal transduction. Development. 2009;136:3699–714.
Ikushima H, Miyazono K. TGF-beta signalling: a complex web in cancer progression. Nat Rev Cancer. 2010;10:415–24.
Colak S, Ten Dijke P. Targeting TGF-β signaling in cancer. Trends Cancer. 2017;3:56–71.
Akhurst RJ, Derynck R. TGF-beta signaling in cancer-a double-edged sword. Trends Cell Biol. 2001;11:S44–51.
Zavadil J, Böttinger EP. TGF-beta and epithelial-to-mesenchymal transitions. Oncogene. 2005;24:5764–74.
Drabsch Y, ten Dijke P. TGF-β signalling and its role in cancer progression and metastasis. Cancer Metastasis Rev. 2012;31:553–68.
Tang B, Vu M, Booker T, Santner SJ, Miller FR, Anver MR, et al. TGF-beta switches from tumor suppressor to prometastatic factor in a model of breast cancer progression. J Clin Investig. 2003;112:1116–24.
Elliott RL, Blobe GC. Role of transforming growth factor Beta in human cancer. J Clin Oncol. 2005;23:2078–93.
Tamari M, Daigo Y, Nakamura Y. Isolation and characterization of a novel serine threonine kinase gene on chromosome 3p22-21.3. J Hum Genet. 1999;44:116–20.
Gallolu Kankanamalage S, Karra AS, Cobb MH. WNK pathways in cancer signaling networks. Cell Commun Signal. 2018;16:72–72.
Zhu W, Begum G, Pointer K, Clark PA, Yang S-S, Lin S-H, et al. WNK1-OSR1 kinase-mediated phospho-activation of Na+-K+-2Cl− cotransporter facilitates glioma migration. Mol Cancer. 2014;13:31–31.
Conti A, Luchini A, Benassi MS, Magagnoli G, Pierini M, Piccinni-Leopardi M, et al. Circulating Candidate Biomarkers in Giant Cell Tumors of Bone. Proteom Clin Appl. 2018;12:e1800041–e1800041.
Burington B, Barlogie B, Zhan F, Crowley J, Shaughnessy JD Jr. Tumor cell gene expression changes following short-term in vivo exposure to single agent chemotherapeutics are related to survival in multiple myeloma. Clin Cancer Res. 2008;14:4821–9.
Li Y, Qin J, Wu J, Dai X, Xu J. High expression of OSR1 as a predictive biomarker for poor prognosis and lymph node metastasis in breast cancer. Breast Cancer Res Treat. 2020;182:35–46.
Tu SW, Bugde A, Luby-Phelps K, Cobb MH. WNK1 is required for mitosis and abscission. Proc Natl Acad Sci USA. 2011;108:1385–90.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thürlimann B, Senn HJ. Strategies for subtypes-dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–47.
Nieto MA, Huang RY-J, Jackson RA, Thiery JP. EMT: 2016. Cell. 2016;166:21–45.
Barrios-Rodiles M, Brown KR, Ozdamar B, Bose R, Liu Z, Donovan RS, et al. High-throughput mapping of a dynamic signaling network in mammalian cells. Science. 2005;307:1621–5.
Singh A, Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010;29:4741–51.
Massagué J, Seoane J, Wotton D. Smad transcription factors. Genes Dev. 2005;19:2783–810.
Wang G, Matsuura I, He D, Liu F. Transforming growth factor-{beta}-inducible phosphorylation of Smad3. J Biol Chem. 2009;284:9663–73.
Wrighton KH, Feng X-H. To (TGF)beta or not to (TGF)beta: fine-tuning of Smad signaling via post-translational modifications. Cell Signal. 2008;20:1579–91.
Alarcón C, Zaromytidou A-I, Xi Q, Gao S, Yu J, Fujisawa S, et al. Nuclear CDKs drive Smad transcriptional activation and turnover in BMP and TGF-beta pathways. Cell. 2009;139:757–69.
Aragón E, Goerner N, Zaromytidou A-I, Xi Q, Escobedo A, Massagué J, et al. A Smad action turnover switch operated by WW domain readers of a phosphoserine code. Genes Dev. 2011;25:1275–88.
Zhou Y, Sun W, Chen N, Xu C, Wang X, Dong K, et al. Discovery of NKCC1 as a potential therapeutic target to inhibit hepatocellular carcinoma cell growth and metastasis. Oncotarget. 2017;8:66328–42.
Chen Z, Zhang Z, Gu Y, Bai C. Impaired migration and cell volume regulation in aquaporin 5-deficient SPC-A1 cells. Respir Physiol Neurobiol. 2011;176:110–7.
Ernest NJ, Weaver AK, Van Duyn LB, Sontheimer HW. Relative contribution of chloride channels and transporters to regulatory volume decrease in human glioma cells. Am J Physiol Cell Physiol. 2005;288:C1451–C1460.
Solomon A, Bandhakavi S, Jabbar S, Shah R, Beitel GJ, Morimoto RI. Caenorhabditis elegans OSR-1 regulates behavioral and physiological responses to hyperosmotic environments. Genetics. 2004;167:161–70.
AlAmri MA, Kadri H, Alderwick LJ, Simpkins NS, Mehellou Y. Rafoxanide and closantel inhibit SPAK and OSR1 kinases by binding to a highly conserved allosteric site on their C-terminal domains. ChemMedChem. 2017;12:639–45.
Shi X, Li H, Shi A, Yao H, Ke K, Dong C, et al. Discovery of rafoxanide as a dual CDK4/6 inhibitor for the treatment of skin cancer. Oncol Rep. 2018;40:1592–1600.
Chen W, Yazicioglu M, Cobb MH. Characterization of OSR1, a member of the mammalian Ste20p/germinal center kinase subfamily. J Biol Chem. 2004;279:11129–36.
Krueger EM, Miranpuri GS, Resnick DK. Emerging role of WNK1 in pathologic central nervous system signaling. Ann Neurosci. 2011;18:70–75.
Zeng G, Gao L, Yu RK. Reduced cell migration, tumor growth and experimental metastasis of rat F-11 cells whose expression of GD3-synthase is suppressed. Int J cancer. 2000;88:53–57.
Shyamasundar S, Lim JP, Bay BH. miR-93 inhibits the invasive potential of triple-negative breast cancer cells in vitro via protein kinase WNK1. Int J Oncol. 2016;49:2629–36.
Nishida H, Sohara E, Nomura N, Chiga M, Alessi DR, Rai T, et al. Phosphatidylinositol 3-kinase/Akt signaling pathway activates the WNK-OSR1/SPAK-NCC phosphorylation cascade in hyperinsulinemic db/db mice. Hypertension. 2012;60:981–90.
Ruan HY, Yang C, Tao XM, He J, Wang T, Wang H, et al. Downregulation of ACSM3 promotes metastasis and predicts poor prognosis in hepatocellular carcinoma. Am J Cancer Res. 2017;7:543–53.
Hung JY, Yen MC, Jian SF, Wu CY, Chang WA, Liu KT, et al. Secreted protein acidic and rich in cysteine (SPARC) induces cell migration and epithelial mesenchymal transition through WNK1/snail in non-small cell lung cancer. Oncotarget. 2017;8:63691–702.
Brabletz T, Jung A, Reu S, Porzner M, Hlubek F, Kunz-Schughart LA, et al. Variable beta-catenin expression in colorectal cancers indicates tumor progression driven by the tumor environment. Proc Natl Acad Sci USA. 2001;98:10356–61.
Vega S, Morales AV, Ocaña OH, Valdés F, Fabregat I, Nieto MA. Snail blocks the cell cycle and confers resistance to cell death. Genes Dev. 2004;18:1131–43.
Ju HR, Jung U, Sonn CH, Yoon SR, Jeon JH, Yang Y, et al. Aberrant signaling of TGF-beta1 by the mutant Smad4 in gastric cancer cells. Cancer Lett. 2003;196:197–206.
Lampropoulos P, Zizi-Sermpetzoglou A, Rizos S, Kostakis A, Nikiteas N, Papavassiliou AG. TGF-beta signalling in colon carcinogenesis. Cancer Lett. 2012;314:1–7.
Yeh HW, Hsu EC, Lee SS, Lang YD, Lin YC, Chang CY, et al. PSPC1 mediates TGF-β1 autocrine signalling and Smad2/3 target switching to promote EMT, stemness and metastasis. Nat cell Biol. 2018;20:479–91.
Niimi H, Pardali K, Vanlandewijck M, Heldin CH, Moustakas A. Notch signaling is necessary for epithelial growth arrest by TGF-beta. J Cell Biol. 2007;176:695–707.
Galliher AJ, Neil JR, Schiemann WP. Role of transforming growth factor-beta in cancer progression. Future Oncol. 2006;2:743–63.
Gomis RR, Alarcón C, He W, Wang Q, Seoane J, Lash A, et al. A FoxO-Smad synexpression group in human keratinocytes. Proc Natl Acad Sci USA. 2006;103:12747–52.
Seoane J, Le H-V, Shen L, Anderson SA, Massagué J. Integration of Smad and forkhead pathways in the control of neuroepithelial and glioblastoma cell proliferation. Cell. 2004;117:211–23.
Gomis RR, Alarcón C, Nadal C, Van Poznak C, Massagué J. C/EBPbeta at the core of the TGFbeta cytostatic response and its evasion in metastatic breast cancer cells. Cancer Cell. 2006;10:203–14.
David CJ, Huang Y-H, Chen M, Su J, Zou Y, Bardeesy N, et al. TGF-β Tumor Suppression through a lethal EMT. Cell. 2016;164:1015–30.
Pattabiraman DR, Weinberg RA. Tackling the cancer stem cells - what challenges do they pose? Nat Rev Drug Discov. 2014;13:497–512.
Steeg PS. Targeting metastasis. Nat Rev Cancer. 2016;16:201–18.
Acknowledgements
This work was supported by the Excellent Youth Medical Talents Program of Shanghai General Hospital (06N1702011).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Li, L., Qin, J. et al. OSR1 phosphorylates the Smad2/3 linker region and induces TGF-β1 autocrine to promote EMT and metastasis in breast cancer. Oncogene 40, 68–84 (2021). https://doi.org/10.1038/s41388-020-01499-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-020-01499-2
This article is cited by
-
Notch3 restricts metastasis of breast cancers through regulation of the JAK/STAT5A signaling pathway
BMC Cancer (2023)
-
Tripartite motif–containing 9 promoted proliferation and migration of bladder cancer cells through CEACAM6-Smad2/3 axis
Journal of Cell Communication and Signaling (2023)
-
Construction and validation of a risk prediction model for clinical axillary lymph node metastasis in T1–2 breast cancer
Scientific Reports (2022)
-
An update regarding the role of WNK kinases in cancer
Cell Death & Disease (2022)