Abstract
Human tumors show altered patterns of protein isoforms that can be related to the dysregulation of messenger RNA alternative splicing also observed in transformed cells. Although somatic mutations in core spliceosome components and their associated factors have been described in some cases, almost nothing is known about the contribution of distorted epigenetic patterns to aberrant splicing. Herein, we show that the splicing RNA-binding protein CELF2 is targeted by promoter hypermethylation-associated transcriptional silencing in human cancer. Focusing on the context of breast cancer, we also demonstrate that CELF2 restoration has growth-inhibitory effects and that its epigenetic loss induces an aberrant downstream pattern of alternative splicing, affecting key genes in breast cancer biology such as the autophagy factor ULK1 and the apoptotic protein CARD10. Furthermore, the presence of CELF2 hypermethylation in the clinical setting is associated with shorter overall survival of the breast cancer patients carrying this epigenetic lesion.
Similar content being viewed by others
Human cells perform their complex physiological roles having only a limited set of genes. Thus, in order to achieve homeostasis, and at the same time respond to changing environment and developmental stages, the presence of precise and dynamic mechanisms of gene regulation is required. Among these, alternative splicing allows the generation of discretely different proteins encoded by the same gene [1, 2]. It is now accepted that the vast majority of human genes undergo some form of alternative splicing [3] and that the thousands of generated protein isoforms overall exert particular activities that are critical for the cellular function. Therefore, the formation of protein diversity is often associated with the distinct skipping or inclusion of exons and other DNA sequences. RNA splicing is thus a highly regulated process that relies on both in trans-regulatory factors and cis-regulatory elements. The spliceosome, the core macromolecular machinery that organizes intron removal and exon junction, and its associated factors includes more than 300 proteins and five small nuclear ribonucleotide particles [4]. The described balanced splicing scenario of the normal cell and tissue becomes highly distorted in cancer. Beyond particular splicing defects of specific genes associated with mutations in skipping sites for those DNA sequences, the overall transcriptome of human tumors exhibits global splicing abnormalities detectable by whole transcriptome analyses [5,6,7], including inefficient exon removal or inclusion of unexpected exons, associated with protein isoforms that contribute to the transforming phenotype [5, 7]. In this regard, an increasing number of cancer-linked mutations in genes encoding spliceosomal proteins and associated RNA splicing factors have recently been reported [7,8,9]. However, although on many occasions there is loss of expression of these splicing factors in the absence of any genetic defect, the role of epigenetic lesions targeting splicing in cancer cells have not been addressed.
Herein, we have interrogated the presence of cancer-specific defects in the CELF protein family (CUGBP and ETR-3-like factors), a family of RNA-binding proteins that act as trans-acting factors enhancing or inhibiting exon inclusion into the final messenger RNA (mRNA) [10,11,12]. The CELF family consists of six members (CELF1–6) that are all characterized by three RNA recognition motifs, two N terminal and one C terminal with a linker region between them termed the “divergent domain” that is the one involved in alternative splicing [10,11,12]. To uncover candidate genetic and epigenetic changes in the CELF family in human tumors, we first data-mined a collection of more than 1000 human cancer cell lines in which we have characterized the exome sequence, gene copy number, transcriptome, and DNA methylation profiles [13]. The available genomic data did not identify the presence of CELF1–6 mutations or copy number changes in the studied cell lines (Dataset S1). Although no genetic lesions were observed in the interrogated genes, promoter CpG island hypermethylation and its associated transcriptional silencing is another relevant mechanism of gene inactivation in cancer cells [14,15,16]. The promoter-associated CpG islands of CELF1, CELF3, CELF4, CELF5, and CELF6 were mostly unmethylated in the assessed cancer cell lines (Dataset S1). However, a CELF2 promoter CpG island was commonly methylated among different cancer cell lines types, including pancreatic (74%, 20 of 27), gastric (57%, 16 of 28), and breast (46%, 19 of 41) tumors (Fig. 1a and Dataset S1). The frequency of promoter CpG island methylation status for the six members of the CELF family in the studied breast cancer cell lines is shown in Supplementary Fig. S1. The CELF2 methylated sites occurred in the CpG island located around the transcription start site of the longest isoform of CELF2 (CELF2-TV2, GRCh37/hg19, NM_006561, originating a 55.7 kDa protein) (Dataset S1). This genomic locus was found unmethylated in all the different normal tissue samples analyzed from the The Cancer Genome Atlas (TCGA) dataset (n = 730) (Dataset S2), including 98 normal breast tissues. Thus, the cancer-specific DNA methylation event at the CELF2 promoter became our focus of interest and was herein further studied in the context of breast cancer.
CELF2 (CUGBP and ETR-3- like factor 2) promoter CpG island hypermethylation-associated transcriptional silencing in human cancer cells. a Percentage of CELF2 methylation in the Sanger set of cancer cell lines according to tumor type. b Bisulfite genomic sequencing of the CELF2 promoter CpG island in breast cancer cell lines and normal breast tissue. CpG dinucleotides are represented as short vertical lines; the transcription start site (TSS) is represented as a long black arrow. Single clones are shown for each sample. The presence of an unmethylated or methylated cytosine is indicated by a white or black square, respectively. c DNA methylation profile of the CELF2 promoter CpG island interrogated using the 450K DNA methylation microarray. Single CpG absolute methylation levels (0–1) are shown. Green, unmethylated; red, methylated. Data from the four breast cancer cell lines and 12 normal breast samples are shown. d Expression levels of CELF2 in breast cancer cell lines determined by real-time PCR (data shown represent mean ± SD of biological triplicates) (left) and western blot (right). e Expression of the CELF2 RNA transcript (data shown represent the mean ± SD of biological triplicates) was restored in the CELF2 hypermethylated and silenced MCF7 and MDA-MB-453 cells upon use of the inhibitor of DNA methylation 5-aza-2′-deoxycytidine (AZA). *p < 0.05, **p < 0.01, and ***p < 0.001
Having found the CELF CpG island methylation patterns shown above, we assessed in detail the possible association with the loss of the CELF2 gene expression at the RNA and protein levels. We developed bisulfite genomic sequencing of multiple clones in the breast cancer cell lines MCF7, MDA-MB-453, MDA-MB-231, and MDA-MB-436 using primers that encompassed the transcription start site-linked CpG island (Supplementary Methods). We observed that the 5′ end CpG island of CELF2 in the MCF7 and MDA-MB-453 cell lines was hypermethylated in comparison with normal tissues (Fig. 1b), whereas the MDA-MB-231 and MDA-MB-436 cells were unmethylated (Fig. 1b). These data were identical to the DNA methylation profiles obtained by the microarray approach (Fig. 1c). The methylated CELF2 cell lines MCF7 and MDA-MB-453 minimally expressed the CELF2-TV2 RNA transcript and the CELF2 55.7 kDa protein, as determined by quantitative real-time PCR (RT-PCR) and western blot, respectively (Fig. 1d). Expression of CELF2 RNA and protein was found in the unmethylated cell lines (Fig. 1d). Treatment of the CELF2-hypermethylated cell lines with the DNA-demethylating agent 5′-aza-2′-deoxycytidine restored CELF2 expression (Fig. 1e). Overall, these results indicate the presence of cancer-specific promoter CpG island hypermethylation-associated loss of the CELF2 gene. Although we have herein focused in breast cancer, we found that epigenetic silencing of CELF2 also occurred in pancreatic cancer cell lines and primary tumors (Supplementary Fig. S2) and thus these results merit further exploration in future research efforts.
Once we had demonstrated the existence of CELF2 CpG island hypermethylation-linked transcriptional inactivation in human breast cancer cell lines, we studied its contribution to the tumorigenic phenotype in vitro and in vivo. Upon efficient transduction-mediated restoration of CELF2 RNA and protein in hypermethylated MCF7 breast cancer cells (Fig. 2a), we tested the potential growth-inhibitory capacity of these cells using the colony formation assay. We observed that the recovery of CELF2 expression by transduction in the MCF7 breast cancer cell line induces a significant decrease in colony formation (Fig. 2b). We also translated these findings to an in vivo mouse model where we tested the ability of these CELF2-transduced MCF7 cells to form orthotopic tumors in the mammary fat pad of nude mice compared with empty vector-transduced cells. The recovery of CELF2 expression in these breast tumors diminished their growth in comparison with empty vector-derived tumors, as observed by the continuous measurement of the tumor volume (Fig. 2c). Tumor samples obtained at the endpoint of the experimental model showed that tumors derived from CELF2-transduced cells weighed less than those tumors obtained from empty vector-transduced cells (Fig. 2c). Thus, our findings suggest that CELF2 has tumor suppressor-like features in transformed cells.
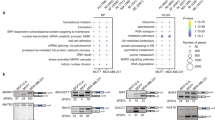
CELF2 (CUGBP and ETR-3-like factor 2) tumor growth-inhibitory properties and its impact in alternative splicing patterns. a Efficient recovery of CELF2 expression upon transduction in MCF7 cells according to quantitative real-time-PCR (qRT-PCR) (left) and western blot (right). Triplicate qRT-PCR values were analyzed and expressed as the mean ± SD. b The colony formation assay showed that MCF7 cells stably transduced with CELF2 formed significantly fewer colonies than empty vector-transduced cells. Data shown represent mean ± SD of biological triplicates. P values are those corresponding to Student’s t test. c Empty vector (EV) and CELF2-transduced MCF7 cells were injected in the mammary fat pad of nude mice to form orthotopic tumors. Tumor volume over time (left) and tumor weight upon sacrifice (right) are shown. P values are those corresponding to Student’s t test. Bars show means ± SD. d Workflow of the RNA-sequencing (RNA-seq) analysis developed to detect RNA alternative splicing changes in MCF7 cells upon CELF2 transduction. e Gene ontology analysis (GO) of the RNAs with differential splicing in MCF7 cells upon CELF2 transduction (hypergeometric test with a false discovery rate (FDR) adjusted p value < 0.05). f Validation of CELF2 splicing targets in breast cancer cells. Restoration of CELF2 expression by transduction in CELF2 hypermethylated MCF7 cells induces a shift in the splicing patterns of NPTN, ULK1, CARD10, RHBDF2, and FBXL2, determined by exon-specific qRT-PCR. P values are those corresponding to Student’s t test. Bars show means ± SD. *p < 0.05, **p < 0.01, and ***p < 0.001
We then wondered about the molecular pathways involved in the growth-inhibitory role of CELF2. In this regard, it is likely that, due to the recognized role of CELF2 in mRNA splicing [10,11,12], the epigenetic loss of CELF2 in breast cancer cells generates a downstream aberrant splicing pattern that contribute to the biology of these tumors. To assess this hypothesis, we performed RNA-sequencing (RNA-seq) to study the transcriptome in empty vector-transduced MCF7 cells (showing CELF2 DNA methylation-associated loss) compared with stably CELF2-transduced MCF7 cells to characterize mRNAs whose different transcript isoforms were CELF2-dependent (Fig. 2d). The RNA-seq data have been deposited in the Sequence Read Archive repository (https://trace.ncbi.nlm.nih.gov/Traces/sra/), under the SRA study ID: PRJNA510082.
We identified 82 events of differential splicing, reflecting distinct exon/intron usage, upon CELF2 transduction in MCF7 cells (Table S1). Most of the CELF2 splicing targeted RNA transcripts corresponded to messenger RNAs (mRNAs) (71 of 82), whereas (11 of 82) were non-coding RNAs (such as pseudogenes and antisense transcripts). Most RNAs (68%; 56 of 82) showed differences in intron retention upon the restoration of CELF2 expression, followed by changes in exon skipping (26%, 21 of 82) and at much lower frequency affected alternative 5′-end or 3′-end splicing sites (5%, 4 of 82 and 1%, 1 of 82, respectively) (Fig. 2d). In order to better characterize the described set of 82 RNAs with significantly differential splicing in CELF2-transduced MCF7 cells in comparison to empty vector-transduced cells, we performed a gene functional annotation by computing overlaps between our gene list and MSigDB (Molecular Signatures Database) gene set collections. We observed an over-representation of biological processes related to proliferation and phosphorylation (FDR < 0.05 in a hypergeometric test). The top 10 significant GO categories by gene count are shown in Fig. 2e. Among the set of transcripts derived from the RNA-seq experiment that showed different exon usage upon CELF2 recovery, and thus candidate targets of the alternative splicing activity of the protein, we found many genes with known oncogenic or antitumoral activities, such as the autophagy factor ULK1 (unc-51 like autophagy activating kinase 1) [17], the apoptotic protein CARD10 (caspase recruitment domain family member 10) [18], the activator of EGFR signaling RHBDF2 (rhomboid 5 homolog 2) [19], the PTEN competitor FBXL2 (F-box and leucine-rich repeat protein 2) [20], and the metastasis breast antigen NPTN (neuroplastin) [21] (Table S1).
The identification of these transcripts in our RNA-seq approach, which can likely contribute to breast tumorigenesis, motivated us to further validate the role of CELF2 epigenetic loss in their proposed dysregulated alternative splicing. In this regard, we confirmed by exon-specific quantitative RT-PCR (Supplementary Methods) that the restoration of CELF2 expression in MCF7 cells significantly enriched intron 12 retention for FBXL2, whereas retention of the intron 17 for ULK1 and CARD10, retention of the intron 2 for RHBDF2, and exon 2 skipping in NPTN were diminished upon transduction-mediated recovery of CELF2 (Fig. 2f). These were the same splicing events detected in our “omics” strategy (Fig. 2d) for these genes (Table S1). We also developed a second model of recovery of CELF2 expression in addition to MCF7. Using the MDA-MB-453 cell line, originally harboring DNA methylation-associated CELF2 silencing (Fig. 1), we performed transduction-mediated recovery of CELF2 expression. (Supplementary Fig. S3). We observed that the intron retention splicing patterns of CELF2-transduced MDA-MB-453 cells (Supplementary Fig. S3) mimicked those observed in CELF2-transduced MCF7 cells (Fig. 2f). We also validated the CELF2-transduced MCF7 in vivo tumor growth data in the new CELF2-transduced MDA-MB-453 model, where the recovery of CELF2 expression inhibited tumor growth (Supplementary Fig. S3). We also created the opposite model by obtaining a stably CELF2-depleted cell line. In this regard, we used the breast cancer cell line DU4475, unmethylated for the CELF2 promoter and expressing its transcript and protein (Supplementary Fig. S4), where we obtained by the short hairpin RNA (shRNA) approach a cell line with depleted levels of the CELF2 transcript and protein (Supplementary Fig. S4). We observed that the induced loss of CELF2 completely reversed the differential intron retention patterns of the target genes (Supplementary Fig. S4) in comparison to the restoration of CELF2 activity in MCF7 (Fig. 2f) or MDA-MB-453 (Supplementary Fig. S3) cells. To further demonstrate the contribution of the differential splicing events mediated by CELF2 loss to breast cancer, we studied one case in detail. Based on our observation that exon 2 retention in NPTN was increased upon transduction-mediated recovery of CELF2 in hypermethylated MCF7 cells (Fig. 2f), we specifically depleted by shRNA the exon 2 retained isoform of NPTN in these CELF2 stably transduced cells (Supplementary Fig. S5). We found that this intervention reversed the observed CELF2-mediated growth inhibition phenotype (Fig. 2b), inducing now an increase in colony formation (Supplementary Fig. S5). Overall, these data support that the identified genes are bona fide targets of CELF2-mediated alternative splicing and that the epigenetic loss of this last factor in breast cancer is associated with their imbalanced isoform content in transformed cells.
Finally, we demonstrated that CELF2 hypermethylation-associated silencing was not exclusively an in vitro cell phenomenon by translating our observations to human primary tumors. Data mining of the human primary tumor collection of TCGA project (https://cancergenome.nih.gov/), studied by the same DNA methylation microarray used herein [22], demonstrated the presence of CELF2 CpG island hypermethylation in a wide spectrum of tumor types (Fig. 3a) that resembled the one described in the cancer cell line cohort (Fig. 1a). Due to our interest and models for CELF2 in breast cancer, we further studied this tumor type in the primary context, observing that the CpG island methylation of CELF2 was found in 39% (263 of 679) of primary breast tumors included in the TCGA dataset (Fig. 3a). Using the available RNA-seq data from the TCGA in breast cancer, we observed that CELF2 CpG island promoter hypermethylation was associated with downregulation of its transcript (Fig. 3b). The link between CELF2 hypermethylation and gene inactivation was further strengthened by data-mining confirmation in early and late passages of PDXs established from human primary breast tumors [23] of the association of CELF2 promoter methylation with the loss of the corresponding transcript (Supplementary Fig. S6). Furthermore, data-mining DNA methylation microarray data available for ductal carcinoma in situ (DCIS) samples [24], we observed that CELF2 hypermethylation occurred early in breast tumorigenesis, and was already present in 11 of 40 (28%) DCIS cases (Supplementary Fig. S7).
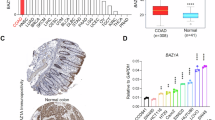
CELF2 (CUGBP and ETR-3-like factor 2) promoter hypermethylation in human breast cancer is associated with poor clinical outcome. a Percentage of CELF2 methylation in the The Cancer Genome Atlas (TCGA) dataset of primary tumors according to tumor type. b CELF2 promoter CpG island methylation is associated with the loss of the CELF2 transcript in primary breast tumors from the TCGA dataset. c Kaplan–Meier analysis of cancer-specific survival in 423 primary breast tumors according to CELF2 methylation status determined by pyrosequencing. The p value corresponds to the log-rank test. Results of the univariate Cox regression analysis are represented by the hazard ratio (HR) and 95% confidence interval (CI). CELF2 hypermethylation is associated with shorter cancer-free survival. d Multivariate Cox regression analysis of cancer-specific survival, represented by a forest plot, taking into account the clinical characteristics of the cohort of breast cancer patients. Values of p < 0.05 were considered statistically significant. In multivariate analyses, significant covariates are considered independent prognostic factors of clinical outcome; as it occurred for CELF2 hypermethylation. *p < 0.05, ***p < 0.001
We then wondered whether CELF2 methylation was of any prognostic value with respect to the growth-inhibitory capacity observed in the in vitro (Fig. 2b) and in vivo (Fig. 2c) models upon CELF2 restoration. To analyze this issue, we studied a cohort of 423 primary breast tumors in which we assessed the CELF2 methylation status by pyrosequencing (Supplementary Methods). We found CELF2 CpG island hypermethylation in 34% (143 of 423) of the primary breast tumor cases, in line with the observed frequency in the TCGA dataset (39%) (Fig. 3a). CELF2 methylation status was associated with other clinical variables and biomarkers such as younger age (Fisher’s exact test p < 0.001), the luminal subtype (χ2 test p = 0.007) and positivity for estrogen receptor/progesterone receptor (ER/PR) status (χ2 test p = 0.018) (Table S2). Importantly, these clinicopathological parameters were also significantly associated between them (younger age/ER/PR positivity, χ2 test p < 0.001; luminal type/ER/PR positivity, χ2 test p < 0.001) (Table S2). Most relevant, the presence of the CELF2 epigenetic alteration was associated with shorter cancer-specific survival (log-rank; p = 0.015; hazard ratio (HR) = 1.48, confidence interval (95% CI) = 1.08–2.04) (Fig. 3c). These observations indicate that the epigenetic loss of the splicing factor CELF2 constitutes a candidate prognostic marker of poor clinical outcome in breast cancer patients. Furthermore, multivariate Cox regression analysis showed that CELF2 hypermethylation was an independent predictor of shorter cancer-specific survival in breast cancer (HR = 1.40, p = 0.04; 95% CI = 1.02–1.93), in comparison to all other patient characteristics (Fig. 3d).
Overall, our data indicate that CELF2, a RNA-binding protein involved in alternative splicing, undergoes promoter CpG island hypermethylation-associated transcriptional silencing in human tumors where we have herein particularly focused on the case of breast cancer. From a mechanistic standpoint, the epigenetic loss of CELF2 is associated with an altered downstream pattern of exon usage in a set of target genes. Most importantly from a cellular and clinical view, the epigenetic loss of CELF2 enhances the growth of breast tumors, and it is associated with those breast cancer patients with the worst outcomes.
References
Nilsen TW, Graveley BR. Expansion of the eukaryotic proteome by alternative splicing. Nature. 2010;463:457–63.
Kalsotra A, Cooper TA. Functional consequences of developmentally regulated alternative splicing. Nat Rev Genet. 2011;12:715–29.
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008;40:1413–5.
Papasaikas P, Valcárcel J. The Spliceosome: the ultimate RNA chaperone and sculptor. Trends Biochem Sci. 2016;41:33–45.
Dvinge H, Bradley RK. Widespread intron retention diversifies most cancer transcriptomes. Genome Med. 2015;7:45.
Jung H, Lee D, Lee J, Park D, Kim YJ, Park W-Y, et al. Intron retention is a widespread mechanism of tumor-suppressor inactivation. Nat Genet. 2015;47:1242–8.
Sveen A, Kilpinen S, Ruusulehto A, Lothe RA, Skotheim RI. Aberrant RNA splicing in cancer; expression changes and driver mutations of splicing factor genes. Oncogene. 2016;35:2413–27.
Dvinge H, Kim E, Abdel-Wahab O, Bradley RK. RNA splicing factors as oncoproteins and tumour suppressors. Nat Rev Cancer. 2016;16:413–30.
Carey KT, Wickramasinghe VO. Regulatory potential of the RNA processing machinery: implications for human disease. Trends Genet. 2018;34:279–90.
Han J. Identification of CELF splicing activation and repression domains in vivo. Nucleic Acids Res. 2005;33:2769–80.
Vlasova IA, Bohjanen PR. Posttranscriptional regulation of gene networks by GU-rich elements and CELF proteins. RNA Biol. 2008;5:201–7.
Dasgupta T, Ladd AN. The importance of CELF control: molecular and biological roles of the CUG-BP, Elav-like family of RNA-binding proteins. Wiley Inter Rev RNA. 2012;3:104–21.
Iorio F, Knijnenburg TA, Vis DJ, Bignell GR, Menden MP, Schubert M, et al. A Landscape of Pharmacogenomic Interactions in cancer. Cell. 2016;166:740–54.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–59.
Baylin SB, Jones PA. A decade of exploring the cancer epigenome—biological and translacional implications. Nat Rev Cancer. 2011;11:726–34.
Feinberg AP. The key role of epigenetics in human disease prevention and mitigation. N Engl J Med. 2018;378:1323–34.
Zhang L, Ouyang L, Guo Y, Zhang J, Liu B. UNC-51-like kinase 1: from an autophagic initiator to multifunctional drug target. J Med Chem. 2018;61:6491–500.
Pan D, Zhu Y, Zhou Z, Wang T, You H, Jiang C, et al. The CBM complex underwrites NF-B activation to promote HER2-associated tumor malignancy. Mol Cancer Res. 2016;14:93–102.
Hosur V, Johnson KR, Burzenski LM, Stearns TM, Maser RS, Shultz LD. Rhbdf2 mutations increase its protein stability and drive EGFR hyperactivation through enhanced secretion of amphiregulin. Proc Natl Acad Sci. 2014;111:E2200–9.
Kuchay S, Giorgi C, Simoneschi D, Pagan J, Missiroli S, Saraf A, et al. PTEN counteracts FBXL2 to promote IP3R3- and Ca2+-mediated apoptosis limiting tumour growth. Nature. 2017;546:554–8.
Rodriguez-Pinto D, Sparkowski J, Keough MP, Phoenix KN, Vumbaca F, Han DK, et al. Identification of novel tumor antigens with patient-derived immune-selected antibodies. Cancer Immunol Immunother. 2009;58:221–34.
Sandoval J, Heyn H, Moran S, Serra-Musach J, Pujana MA, Bibikova M, et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics. 2011;6:692–702.
Gómez-Miragaya J, Palafox M, Paré L, Yoldi G, Ferrer I, Vila S, et al. Resistance to taxanes in triple-negative breast cancer associates with the dynamics of a CD49f+ tumor-initiating population. Stem Cell Rep. 2017;8:1392–407.
Johnson KC, Koestler DC, Fleischer T, Chen P, Jenson EG, Marotti JD, et al. DNA methylation in ductal carcinoma in situ related with future development of invasive breast cancer. Clin Epigenet. 2015;7:75.
Acknowledgements
We thank CERCA Program/Generalitat de Catalunya for institutional support. This work was supported by the Health Department PERIS-project no. SLT/002/16/00374 and AGAUR-projects nos. 2017SGR1080, 2014SGR633, and 2009SGR1315 of the Catalan Government (Generalitat de Catalunya); the Spanish Institute of Health Carlos III (ISCIII) project no. DTS16/00153 and Ministerio de Economía y Competitividad (MINECO) project no. SAF2014–55000-R, co-financed by the European Development Regional Fund, “A way to achieve Europe” ERDF; the Cellex Foundation; and “la Caixa” Banking Foundation (LCF/PR/PR15/11100003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Piqué, L., Martinez de Paz, A., Piñeyro, D. et al. Epigenetic inactivation of the splicing RNA-binding protein CELF2 in human breast cancer. Oncogene 38, 7106–7112 (2019). https://doi.org/10.1038/s41388-019-0936-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0936-x