Abstract
Recently, we detected a new fusion transcript LMO3-BORCS5 in a patient with Ewing sarcoma within a cohort of relapsed pediatric cancers. LMO3-BORCS5 was as highly expressed as the characteristic fusion oncogene EWS/FLI1. However, the expression level of LMO3-BORCS5 at diagnosis was very low. Sanger sequencing depicted two LMO3-BORCS5 variants leading to loss of the functional domain LIM2 in LMO3 gene, and disruption of BORCS5. In vitro studies showed that LMO3-BORCS5 (i) increases proliferation, (ii) decreases expression of apoptosis-related genes and treatment sensitivity, and (iii) downregulates genes involved in differentiation and upregulates proliferative and extracellular matrix-related pathways. Remarkably, in vivo LMO3-BORCS5 demonstrated its high oncogenic potential by inducing tumors in mouse fibroblastic NIH-3T3 cell line. Moreover, BORCS5 probably acts, in vivo, as a tumor-suppressor gene. In conclusion, functional studies of fusion oncogenes at relapse are of great importance to define mechanisms involved in tumor progression and resistance to conventional treatments.
Similar content being viewed by others
Introduction
Fusion transcripts are responsible for 20% of global cancer morbidity and have been recognized as major cancer drivers [1]. They result from the juxtaposition of two genes, usually separated, due to chromosomal rearrangements or nonchromosomal events [2]. They can affect coding or regulatory sequences which can lead to (i) the creation of a chimeric oncoprotein, (ii) aberrant overexpression of the second partner involved in the fusion, or (iii) truncation of a tumor-suppressor gene (TSG) [2].
Fusions can serve as diagnostic, prognostic, predictive biomarkers, as well as therapeutic targets [2,3,4,5,6,7,8,9,10]. Apart from the recurrent known fusion genes, the majority of newly discovered gene fusions vary extremely in their expression and their recurrence is much lower than other somatic aberrations [11]. Even though it is commonly thought that recurrence represents the first criteria to consider an alteration as driver, its impact is determined by patient and tumor’s history (e.g., genetic background, tumor type, or treatments received) and a single nucleotide variation could have different consequences on drugs sensitivity depending on cancer types [12]. Hence, nonrecurrent fusions could have a high clinical value for the individual patient. In addition, most of studies have been performed on patients at diagnosis while the analysis of tumors at relapse remains one of the challenges to enrich the repertoire of evolving driver alterations, including fusions [13].
Previously, we performed a retrospective study on RNA-sequencing data of pediatric resistant or relapsing patients included in the molecular profiling trial MOSCATO-01 conducted at Gustave Roussy [14, 15]. Interestingly, we detected a new fusion transcript, LMO3-BORCS5 at diagnosis and at relapse in the tumor of a patient with Ewing sarcoma (EwS) [16].
LMO3-BORCS5 results from a deletion of 4.2 megabases on chromosome 12 where both LMO3 and BLOC-1 Related Complex Subunit 5 (BORCS5) are located, 12p12.3 and 12p13.2, respectively. LIM-domain-only proteins (LMO) are characterized by the presence of two functional domains, LIM1 and LIM2. The family of proteins is composed of four members (LMO1–4) mediating protein–protein interactions between transcription factors and other proteins, representing important transcription co-regulators acting on both tumorigenesis and differentiation [17].
The oncogenic activity of LMO3 has been described in neuroblastoma where its expression is correlated with amplified MYCN and activation of HEN2, leading to poor prognosis in patients [18]. Larsen et al. demonstrated the ability of LMO3 to directly inhibit TP53, strengthening its oncogenic status [19]. LMO3 was also related to lung carcinoma and gastric cancer [18, 20, 21]. LMO3 is highly expressed in brain and acts as a regulator of neurogenesis and controls neural-specific transcription factors, thus being critical for cell differentiation [22].
BORCS5, also named LOH12CR1, belongs to the BORC complex involved in the regulation of cell migration and lysosomal trafficking as well as in axonal transport of synaptic vesicles [23,24,25]. Its chromosome locus contains several TSGs for which loss of heterozygosity has frequently been observed in various neoplasms [26]. Yet, no study has described the different roles of BORC subunits or the possible role of BORCS5 loss on tumorigenicity.
The use of fusions as personalized medicine tools in clinics is often limited due to the absence of functional studies [27, 28]. Therefore, the aim of our study was to understand the role and function of LMO3-BORCS5 in the development and/or resistance of tumors and to evaluate the effects of BORCS5 knockout. Taking together, our results suggest that LMO3-BORCS5 fusion oncogene plays an essential role in tumorigenesis and treatment sensitivity and could be a potential therapeutic target. Characterization of BORCS5 showed pro-oncogenic effects on proliferation and treatment sensitivity in vitro, and could act as a gene in vivo by inhibiting tumor invasiveness.
Results
LMO3-BORCS5 detected in a patient at relapse with EwS is highly coexpressed with EWS/FLI1
At relapse, LMO3-BORCS5 was highly coexpressed with the characteristic fusion oncogene of Ewing sarcoma EWS-FLI1 (Ct: 23 and 22.4 at diagnosis and relapse respectively). RT-qPCR followed by migration in gel electrophoresis detected two variant forms of the LMO3-BORCS5 fusion transcript in the EwS tumor sample at relapse (LMO3-BORCS5 v1 corresponding to an amplicon size of 264 pb and LMO3-BORCS5 v2 at 205 pb), while it was hardly detectable at diagnosis (Ct: 22.6 vs 39.7) (Fig. 1a). This particular sequence comprised a lot of (N) bases after Sanger sequencing results, probably due to the low expression level of the fusion nevertheless the fusion point was identical to LMO3-BORCS5 v1 sequence (Fig. 1b). Sanger sequencing of both amplicons showed that LMO3-BORCS5 v2 corresponds to a deletion of 59 bp at the 3′-end of LMO3 exon 3 compared with LMO3-BORCS5 v1. In both cases, the breakpoint in BORCS5 remains identical and is situated at the beginning of exon 3 (Fig. 1b, c).
Detection of LMO3-BORCS5 fusion transcripts in a patient with Ewing sarcoma, at diagnosis and relapse and reconstitution of the different LMO3-BORCS5 variants at mRNA and protein levels. a The presence of LMO3-BORCS5 and EwS-FLI1 was tested in the mRNA obtained from tumor samples at diagnosis and at relapse of a patient with EWS. After mRNA extraction and reverse transcription (RT), the presence of LMO3-BORCS5 was assessed by real-time PCR (qPCR) followed by migration of PCR products on gel. b Sanger sequencing of LMO3-BORCS5 v1 and v2, and graphical reconstitution at mRNA level: LMO3 sequence (blue), BORCS5 sequence (gray), and primers (yellow) used to precisely amplify the junction sequence. The red arrow indicates the fusion point for the variant 1 (v1), the green arrow indicates the fusion point for the variant 2 (v2). Sequencing of bacterially cloned v1 and v2 PCR product confirmed the correct sequence (not shown). c Graphical reconstitution of LMO3-BORCS5 v1 and LMO3-BORCS5 v2 caused by an interstitial deletion of 4.2 MB on chromosome 12. This fuses exons 1, 2, 3 of LMO3 to exons 3 and 4 of BORCS5 for LMO3-BORCS5 v1, while a part of exon 3 is lost for the LMO3-BORCS5 v2. d Graphical reconstitution of LMO3-BORCS5 v1 and LMO3-BORCS5 v2 protein showing that the fusion disrupts differently the regulation domain of LMO3, and loss of BORCS5 in both variants
At the protein level on the N-terminal part, variant 1 is predicted to produce a protein with complete conservation of LIM1 and loss of LIM2 domains of LMO3, while 24 amino-acids at the end of LIM1 domain are lost in variant 2 (Fig. 1d). In both cases on the C-terminal part, the fusion transcripts lead to a frameshift in the BORCS5 sequence giving a putative truncated out-of-frame BORCS5 protein with the creation of 43 amino-acids associated with unknown protein domain/function (Fig. 1d, in dark red).
In order to have a representative model of LMO3-BORCS5, 27 cancer cell lines representing different pediatric tumors were screened for this fusion (Fig. S1A). It was detected at a very low level (Ct~39) in one of the five EwS cell lines (STA-ET1). As in the EwS patient’s tumor at relapse, STA-ET1 showed two variant forms after sequencing (Fig. S1A in bold). Eleven of the remaining 22 cell lines depicted ct by RT-qPCR but none of them showed a sufficient LMO3-BORCS5 expression (Ct > 35) impeding the use of these cell lines as models for further studies.
Among the 27 cell lines previously tested for LMO3-BORCS5 fusion, the relative mRNA expression of LMO3 and BORCS5 wild type has been assessed by RT-qPCR by using primers designed within regions not involved in the fusion (Table S1, Fig. S1B, C). The expression varied among the different cell lines; a high expression of LMO3 was found in neuroblastoma cell lines IGR-NB8, IGR-N91 and SH-SY5Y and the EwS cell lines STA-ET-1, TC71, and A673 (Fig. S1B). Expression of BORCS5 was present in all 27 cell lines tested with the highest expression in medulloblastoma cell line D283MED, followed by EwS (EW7 and SK-N-MC) and osteosarcoma (HOS-143B, IOR/OS18 and SAOS2) cell lines (Fig. S1C).
Due to the absence of representative model of LMO3-BORCS5 fusion transcript allowing experimental characterization of its biological relevance, we cloned both variants and tested their effects in vitro and in vivo.
LMO3-BORCS5 variant 1 displays functional relevance
For the human EwS cell line A673, two clones (C1 and C2) were obtained containing the empty vector (Fig. 2a, well 1), two clones (C1 and C2) containing LMO3-BORCS5 v1 (Fig. 2a, well 2), and two clones (C1 and C2) containing LMO3-BORCS5 v2 (Fig. 2a, well 3). Then, we investigated (i) the proliferation rate of the two clones (C1 and C2) containing LMO3-BORCS5 v1 derived from A673, (ii) their doubling time and the repartition of cells within the cell cycle, as well as (iii) their transcriptome profile. As shown in Fig. 2b, each two clones derived from the same transfection showed similar doubling time, and almost the same cell-cycle repartition (Fig. 2c). Due to their similar profile, we selected clone 1 for further studies.
Effects of LMO3-BORCS5 on cell proliferation, cell cycle, and tumorigenicity in A673 and NIH-3T3 cell lines. Empty vector pcDNA3.1, pcDNA3.1 containing LMO3-BORCS5 v1, and pcDNA3.1 containing LMO3-BORCS5 v2 were stably transfected into A673 (human EWS cell line) or NIH-3T3 (non-tumorigenic mouse fibroblasts cell line). a Presence of LMO3-BORCS5 in the clones was verified by RT-qPCR of each cell line followed by migration of the products. b Doubling time of the different clones obtained after transfection of A673 cells with empty vector (clone 1 and 2), LMO3-BORCS5 v1 (clone 1 and 2), and LMO3-BORCS5 v2 (clone 1 and 2). Cells was calculated according to an exponential regression equation, given as: y = axebx, via the formula: t1/2 = ln(2)/b. The results are presented as mean ± SD in hours of at least two independent experiments. c Cell-cycle repartition was obtained after DNA staining with propidium iodide-containing buffer followed by FACS analysis. The results are presented as mean ± SD in % of cells in each cell cycle phase of thre independent experiments. *p > 0.05, **p > 0.01; ***p > 0.001 compared with A673 empty vector C1. A673 (left) and NIH-3T3 (right) were stably transfected with the empty vector pcDNA3.1 or with pcDNA3.1 LMO3-BORCS5. Proliferation of A673 (d) or NIH-3T3 (f) cells using the IncuCyte™ video microscopy system was assessed. Doubling time of cells was calculated as previously described. Results are presented as mean ± SD in hours of three independent experiments. Cell-cycle repartition in A673 (e) or NIH-3T3 (g) cells. The results are the mean of at least three independent experiments
Studies on variant 2 did not reveal any significant difference in doubling time (Fig. 2b, conditions 5–6), and cell cycle repartition between A673 + empty vector and A673 + LMO3-BORCS5 v2 (C1 or C2) (Fig. 2c, conditions 5–6); therefore, A673 + LMO3-BORCS5 variant 2 function was not further investigated.
For the non-tumorigenic mouse fibroblast cell line NIH-3T3, one clone containing empty vector and one clone containing pcDNA3.1-LMO3-BORCS5 v1 were obtained (Fig. 2a, wells 4 and 5, respectively). One clone derived from NIH-3T3 containing pcDNA3.1-LMO3-BORCS5 v2 was obtained (Fig. 2a, well 6); however, the cells did not expand in vitro, precluding further studies.
Introduction of LMO3-BORCS5 v1 in either A673 or NIH-3T3 cell lines increased proliferation by reducing the doubling time around 20% as compared with cells expressing the empty vector (Fig. 2d, f). In A673, this was associated with a shift from G0/G1 to the G2/M phase (Fig. 2e) as compared with NIH-3T3 (Fig. 2g). No effect of LMO3-BORCS5 expression on cell migration in both A673 and NIH-3T3 was detected (not shown).
LMO3-BORCS5 v1 increases or induces tumorigenicity respectively in A673 and NIH-3T3 cells
After subcutaneous injection of A673 clones, acceleration of tumor growth was observed in xenografts expressing LMO3-BORCS5 compared with the empty vector (Fig. 3a). Hematoxylin–eosin–safranin (HES) staining showed an increase of mitotic cells (yellow arrows), and tumor infiltration in adipose (green arrows) and muscular (red arrows) tissues in the presence of LMO3-BORCS5-expressing tumors compared with controls (empty vector) (Fig. 3b, upper and lower panels). Remarkably, LMO3-BORCS5 expression in NIH-3T3 led to tumor development (Fig. 3c); HES analysis showed a high number of mitotic cells in these tumors (Fig. 3d). Of note, the presence of LMO3-BORCS5 fusion transcript was confirmed by RT-qPCR in both A673 + LMO3-BORCS5 and NIH-3T3 + LMO3-BORCS5 tumors (Ct≈22 and 27, respectively).
LMO3-BORCS5 increases tumor growth in A673 cells and induced tumorigenicity in NIH-3T3. Tumor growth was monitored after inoculation of 10 × 106 A673 cells (a) or NIH-3T3 cells (c) in nude mice (n = 5/group). Tumors were measured until reaching a size of 1000 m3. Results are presented as the tumor volume mean + SD in mm3 function of days. b Comparison of HES images in tumors obtained from A673 empty vector or A673 + LMO3-BORCS5 cell lines at ×50 (upper panel) or ×400 (lower panel) magnification. Histology revealed an increase of tumor infiltration in adipose (green arrows) and muscular (red arrows) tissues in presence of LMO3-BORCS5 in A673 cells (upper panel). Increase of mitosis was also observed in presence of the fusion (yellow arrows) in A673 cells (lower panel). d HES of tumors obtained from NIH-3T3 + LMO3-BORCS5 ×400 magnification showing cells in mitotic division (yellow arrows)
LMO3-BORCS5 decreases sensitivity to antineoplastic agents and apoptosis
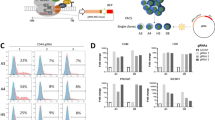
The effect of LMO3-BORCS5 on drug sensitivities of 11 anticancer drugs was screened, including five received by the patient (listed in the “Materials and methods” section). Vincristine and SN-38 showed an increase of IC50 values in A673 + LMO3-BORCS (Fig. 4a). Vincristine and SN-38 treatment showed a reduced fraction of cells in early apoptosis (delta between treated and non-treated cells of 28.8% for A673 + empty vector vs 13.7% for A673 + LMO3-BORCS5) (Fig. 4b). The same tendency, albeit not statistically significant, was observed after treatment with SN-38 (delta of 31.1% for A673 + empty vector vs 23.5% for A673 + LMO3-BORCS5) (Fig. 4b). Late apoptosis after vincristine or SN-38 treatment appeared to be unchanged in presence or absence of LMO3-BORCS5.
Effects of LMO3-BORCS5 expression on the expression sensitivity to anticancer agents and of apoptosis-related genes. a The IC50 was determined by linear regression model after MTT assay of A673 stably transfected with pcDNA3.1 (A673 + empty vector) or with pcDNA3.1 LMO3-BORCS5 (A673 + LMO3-BORCS5) cells treated with vincristine (Vinc) and SN-38. b Early and late apoptosis detection by flow cytometry in A673 stably transfected with A673 + empty vector or with A673 + LMO3-BORCS5 cells after treatment with Vinc or SN-38. The delta represents the difference between the percentage of cells in early or late apoptosis in cells treated with Vinc or SN-38, and the percentage of cells in early or late apoptosis in nontreated cells. c Relative mRNA expression of apoptosis-related genes BAX, NOXA, TP53, and P21 by RT-qPCR in A673 and NIH-3T3 cells stably transfected with empty vector pcDNA3.1 or with pcDNA3.1 LMO3-BORCS5. *p < 0.05: by using Mann–Whitney test, a statistical difference was found for Vinc treatment between A673 + empty vector and A673 + LMO3-BORCS5 clones
The inducible decrease in sensitivity to vincristine and SN-38 led us to hypothesis that the inhibition of apoptotic gene expressions involved in cancerogenesis could favor resistance to cell death. Therefore, the expression of candidate genes (BAX, NOXA, TP53, and P21) was assessed by RT-qPCR (Fig. 4c). In A673 + LMO3-BORCS5, all genes except TP53 were significantly lower expressed compared with A673 + empty vector (Fig. 4c, upper panels). In NIH-3T3 + LMO3-BORCS5, decreased P21 expression was observed (Fig. 4c, lower panels). As the strongest impact was seen on P21, these results further support the previously obtained results on proliferation and cell cycle.
LMO3-BORCS5 downregulates genes associated with cell differentiation pathways
Microarray analysis was performed on two clones of A673 + LMO3-BORCS5 and compared with A673 empty vector. A total of 295 genes were differentially expressed [−2 < log2 (fold change > 2]. The volcano plot showed that more than 80% of differentially expressed genes were downregulated (243 genes downregulated vs 52 genes upregulated).
Using ToppGene Suite, the 12 most significantly enriched pathways of genes being upregulated can be grouped into three different cellular functions: metabolism, extracellular matrix synthesis, and proliferative signaling (Table 1, upper panel). Seven of the twelve most significantly enriched pathways in downregulated genes were related to cell differentiation, more precisely, to neural (e.g., “DCC signaling”, “NCAM1 interactions”, “TRKA signaling”), cardiac (e.g., “cardiac conduction signaling”, “muscle contraction pathway”) or renal differentiation (“aldosterone-regulated sodium reabsorption”) as well as cell pluripotency (transcriptional regulation of pluripotent stem cells pathway). Finally, apoptosis-related genes BAX, NOXA, and P21, found differentially expressed between A673 + empty vector and A673 + LMO3-BORCS5 by RT-qPCR were confirmed to be downregulated in microarray results [log2(FC) equal to −0.4, −0.6, and −1.1, respectively]. The microarray data related to the study have been submitted to the Array Express data repository at the European Bioinformatics Institute (http://www.ebi.ac.uk/arrayexpress/), under the accession number: E-MTAB-7021.
Then, we chose to validate 18 genes highly linked to cancer pathways by RT-qPCR that have been found to be differentially expressed by microarray analysis. All tested genes except ABCC3 were validated, accounting of 95% of reliability (Table 2). The majority of tested genes were found to be downregulated in the presence of LMO3-BORCS5, except Fyn-related kinase (FRK) which was upregulated. They represent genes regulating cell fate determination (FGF1 and WNT6), genes linked to neural differentiation (ALK, CUX2, FGF13, NTRK1, PAX7, and SEMA3C), IGF2, MYOD1, and RHOJ for muscle cell differentiation and NRP1 regulating cardiac differentiation as well as genes linked to cell proliferation or migration (P21, PRKCA, MMP10, and LZTS1; Table 2).
Both KO BORCS5 clones show comparable expression and cellular profiles
In order to characterize the role of BORCS5 on oncogenesis and tumorigenicity, BORCS5 gene was depleted using the CRISPR/Cas9 system in A673 cells. We compared the two derived clones obtained after KO of BORCS5 by CRISPR–Cas9 system. Both C1 and C2 did not expressed BORCS5 protein testifying the knockdown outcome (Fig. 5a), and showed the quasi same doubling time (Fig. 5b) as well as identical repartition of cells within the cell cycle (Fig. 5c). Therefore, only KO BORCS5 C1 was used for further studies, named “A673 KO BORCS5”. After Sanger sequencing a homozygous KO BORCS5 clone was identified with a compound heterozygosity leading to a deletion of 950 bp in exon four on one allele, and insertion leading to a frameshift on the other (Fig. 5d, e). The corresponding protein loss was proved by western blot (Fig. 5f).
Knockout of BORCS5 in A673 wild-type cells and comparison of the two clones obtained after KO of BORCS5 in A673 cells using CRIPR-Cas9 system. BORCS5 gene was inactivated using the CRISPR/Cas9 system targeting the last exon which contains the functional domain. a Western blot of BORCS5 protein in A673 KO BORCS5 clone 1 (C1) and A673 KO BORCS5 clone 2 (C2). b Doubling time was calculated as described previously. The results are presented as mean ± SD in hours of at least two independent experiments. c Cell-cycle repartition FACS analysis. Results are presented as mean ± SD in % of cells in each cell cycle phase of at least two independent experiments. d Deletion of the target sequence was tested by PCR which resulted in a 950-bp smaller band in KO cells compared with WT cells. e KO was confirmed by Sanger sequencing which shows an insertion of one nucleotide on one allele (upper sequence), and a 957-bp deletion on the other allele (lower sequence) between two protospacer-adjacent motifs (PAM). f Knockout of BORCS5 was further confirmed by western blot on the total protein extracts of A673 WT and A673 KO BORCS5 cells. By using an anti-BORCS5 antibody, no BORCS5 protein was detected in the KO cells
BORCS5 disruption decreases cell migration and resistance in vitro but increases tumorigenicity
Then, the A673 WT was compared with the A673 KO BORCS5 regarding proliferation, migration, sensitivity to anticancer drugs, and tumorigenicity. Significant inhibition of migration was noticed in A673 KO BORCS5 compared with A673 WT (Fig. 6a).
Impact of BORCS5 knockout on A673 cells in vitro and in vivo. a Effects of BORCS5 disruption on A673 cell migration by using scratch wound test to compare A673 WT and A673 KO BORCS5. Cells in monolayer were seeded in Matrigel™ (BD) then the monolayer was scratched using a 96-pin wound maker and wound confluence was monitored every 4 h IncuCyte™. Images (×10) representative of one experiment, taken by the Incucyte™ system of A673 WT or A673 KO BORCS5 cell migration after 15 h of incubation at 37 °C and 5% CO2. Results are presented as the mean time to completely close the wound per cell line of three independent experiments. b IC50 obtained by MTT assay of A673 WT and A673 KO BORCS5 cells treated with vincristine, SN-38 and mafosfamide. c Early and late apoptosis detection by flow cytometry in A673 WT and A673 KO BORCS5 cells after treatment with vincristine or SN-38. The delta represents the difference between the percentage of cells in early or late apoptosis in treated versus non-treated cells. d A total of 10 × 106 cells of A673 WT and A673 KO BORCS5 cells were injected subcutaneously in nude mice in order to monitor tumor growth and test the influence of BORCS5 disruption on tumorigenicity (n = 5/group). Results are presented as the tumor volume mean in mm3. e Comparison of tumors obtained from A673 WT or A673 KO BORCS5 cell lines at ×50 (upper panel) or ×400 (lower panel) magnification. Histology of tumors revealed a slight increase of tumor infiltration in muscular tissues (red arrows) in absence of BORCS5 in A673 cells (upper panel). No difference regarding the number of cells in mitosis was observed between A673 WT and A673 KO BORCS5 cells (yellow arrows; lower panel)
The consequence of BORCS5 loss on drug sensitivities using the previous panel of 11 anticancer drugs was tested. A significant decrease of IC50 values (60% vincristine and 40% for both SN-38 and mafosfamide) was observed in A673 KO BORCS5 compared with WT (Fig. 6b). Effects of vincristine and SN-38 on early and late apoptosis comprised a significant increase of cells in late apoptosis after vincristine treatment in A673 KO BORCS5 cells compared with A673 WT (Fig. 6c). The same tendency was observed after SN-38 treatment but did not reach significance while early apoptosis in A673 was not affected by the KO BORCS5 (Fig. 6c).
Interestingly, after xenografting of A673 WT and A673 KO BORCS5 cells, a tumor growth delay was observed for A673 KO BORCS5 (Fig. 6d). HES revealed a similar general architecture of collected tumors and no difference in mitosis frequency. (Fig. 6e, lower panel, yellow arrows). However, increased invasion into striated muscle fibers was seen in A673 KO BORCS5 tumors (Fig. 6e, upper panel, red arrows).
Discussion
This is the first study that reports the oncogenic function of a newly described fusion transcript LMO3-BORCS5. Identified in a patient with EWS/FLI1-positive EwS at relapse, this fusion transcript was already detectable at diagnosis. According to cBioPortal cancer database, LMO3 and BORCS5 are altered in 1.5% of total cancers, and fusions represent the third and second alteration type, respectively (Fig. S2). This is probably underestimated as it comprises both adult and childhood cancers and several studies do not assess the presence of fusion genes.
Both LMO3 and BORCS5 genes are situated in the short arm of chromosome 12p13.2. LMO3-BORCS5 sequence exhibited a 4.2-MB deletion within the short arm of chromosome 12 leading to the loss of the functional domain LIM2 in LMO3 gene and the disruption of the reading frame of BORCS5. LMO3-BORCS5 was detected in two variant forms (v1 and v2) both leading to a loss of the LIM2 domain after translation. Interestingly, the loss of interaction domains in fusions has been shown to be a sign of oncogenicity [29, 30]. Variant 1 of the fusion transcript LMO3-BORCS5 was abundant at relapse and barely detected at diagnosis, suggesting that the fusion transcript was not induced by prior anticancer treatments but probably selected during disease progression. Whether v1 and v2 are expressed within the same cell or in different cells, could have been investigated by a single-cell sequencing approach, but as the probability for fusion events to occur on both alleles is low, the two variants should be the product of an alternative splicing. Even if most of the cell lines tested derived from pediatric cancer types, the positive ones were of diverse histological origins. However, the expression level of LMO3-BORCS5 in these cell lines was relatively low as previously observed for other chimeras [31]. This could suggest that, a second event might be required for LMO3-BORCS5 oncogenicity as reported for several fusions [32, 33].
When assessing the expression of wild-type LMO3 and BORCS5 across 27 different tumor cell lines, LMO3 was mainly expressed in neuroblastoma and EwS [18, 34]. This result was concordant, as LMO3 is a brain-specific gene and both pathologies are thought to arise from precursors of the neural crest or neuroectoderm [33, 35]. BORCS5 was expressed in all cell lines tested. The highest expression was found in cells of EwS and osteosarcoma bone tumors, as well as medulloblastoma. Therefore, LMO3 and BORCS5 should be explored in tumors derived from neural or bone precursors.
Our first aim was to investigate the function of LMO3-BORCS5 on oncogenesis. We showed that LMO3-BORCS5 v1 plays a key role in oncogenesis: it increased proliferation in both cell lines (A673 and NIH-3T3) and increased or induced tumor growth and invasiveness in the A673 EwS and the non-tumorigenic NIH-3T3, respectively. This was strengthened by microarray analysis where LMO3-BORCS5 led to downregulation of genes involved mainly in differentiation. In fact, the LIM2 domain was shown to be critical for erythropoietic differentiation in LMO2 protein, highlighting the importance of LIM2 in cell differentiation [36]. The absence of oncogenic effects for LMO3-BORCS5 v2 suggesting that the complete conservation of LIM1 domain is required for oncogenicity.
Loss of LIM2 within LMO3-BORCS5 could be a factor of the dedifferentiation and consequently the oncogenic effects observed. Dedifferentiation is considered as one of the main characteristics of cancer cells, which are blocked at early stages of differentiation resulting in an over-proliferative state [37]. This was also supported by a shift of cells toward the mitosis phase in presence of LMO3-BORCS5 observed by FACS and tumor histological study.
In addition, decreased sensitivity to two antineoplastic agents (vincristine and SN-38) was observed in A673 cells expressing LMO3-BORCS5 which was related to inhibition of gene expression involved in apoptosis
Gene upregulation in microarray in presence of LMO3-BORCS5 affected proliferative signaling, consistent with results obtained in vitro. Some of these genes can be related to extracellular matrix synthesis, supported by our findings that in vivo as tumors with LMO3-BORCS5 were more invasive, although migration in vitro was not enhanced. This could reflect the limits of the scratch test used in vitro, a model of study that could be inadequate to our cells.
Our team previously showed that the inhibition of fusion transcripts responsible for tumor progression in thyroid or prostate cancers by injection of vectorized siRNA in vivo led to a decrease of tumor growth [38, 39]. Therefore, the use of LMO3-BORCS5 as a therapeutic target could be considered in light of our results. Since no LMO3 inhibitor exists, vectorized siRNAs could be one of the solutions to target this oncogene. Also, given the expression and the oncogenic role of LMO3 in neuroblastomas and gastric cancer, it would be of interest to test the impact of certain antineoplastic molecules used in the treatment of these pathologies. Of note, the FRK was upregulated in the presence of LMO3-BORCS5. This non-receptor tyrosine kinase leads to an activation of STAT3 pathway and inhibition of TP53, and is highly expressed in sarcomas [40]. However, sensitivity to dasatinib with or without LMO3-BORCS5 showed no difference to treatment as described for hepatocellular carcinoma cell lines [40].
As LMO3-BORCS5 fusion leads to a frameshift of BORCS5 with the putative formation of 43 new amino acids, we further aimed to characterize the effects of BORCS5 loss on tumorigenicity. It could be demonstrated that the disruption of BORCS5 in A673 cells had both pro- or anti-tumorigenic effects, which could depend on the tumor type and microenvironment context. In vivo results favor of a tumor-suppressor role of BORCS5, consistent with the loss observed in the formation of LMO3-BORCS5 fusion. The disruption of BORCS5 in LMO3-BORCS5 could have two roles (i) its loss on one allele could serve as “second hit” or (ii) the 43 new amino-acids created in the C-terminal part of the fusion protein after LIM1 domain could enable the stabilization of the new smaller LMO3 protein, preventing from its degradation [27].
Several studies previously reported that only in-frame transcripts need to be considered as drivers [41, 42]; however, we have demonstrated the highly oncogenic properties of an out-of-frame fusion. These results lead to a paradigm shift of nonrecurrent alterations, which are commonly classified as passenger events [27, 43]. As fusions appeared to be segregated according tumor type and given the low incidence of EwS, the statistical power of our cohort might be insufficient to detect recurrence [27, 1]. In addition, it is possible that LMO3-BORCS5 was present in other patients but below the detection limit of RNA sequencing, or filtered out [27]. It is also noteworthy that previous studies, which have aimed at detecting fusion in EwS, were not focused on relapsed patients and/or used whole-genome sequencing, with a lower coverage than RNA-seq [44,45,46]. This could represent a major determinant for not recurrently detecting this fusion because rearrangements loops arise in an early EwS clone from which both the primary tumor and the lethal relapse emerged, and then evolved in parallel until clinically detected [47].
In conclusion, we discovered a new fusion oncogene LMO3-BORCS5 which emphasizes the critical role of fusion transcripts not only in tumor initiation but also in progression and resistance even in the individual patient. Their deep study could explain some consequences missed by traditional analyses. This suggests that cancer treatment should include the consideration of individual alterations next to looking for universality. Fusion transcripts being one of the most remarkable tools used in personalized medicine should be systematically integrated and investigated to new genome-based clinical trials.
Materials and methods
Patient’s history, detection, and sequencing of LMO3-BORCS5 fusion transcript
Using the primers described in Table S1, LMO3-BORCS5 was identified together with the characteristic fusion oncogene of Ewing sarcoma EWS-FLI1 after in silico RNA-seq analysis of a 20-year-old male EwS patient, included in the cohort of pediatric MOSCATO-01 (ClinicalTrials.gov: NCT01566019) patients that we previously analyzed [16]. He had relapsed 3.4 years after initial diagnosis treated with chemotherapy (vincristine, ifosfamide, doxorubicin, etoposide, dactinomycin, and cyclophosphamide). The presence of LMO3-BORCS5 fusion transcript was confirmed by RT-qPCR in tumor samples at diagnosis and relapse. The products were sequenced by Sanger technique (Eurofins, Cochin Institute, Paris, France).
Cell lines and cell culture
To assess the presence of LMO3-BORCS5 fusion transcript, 27 cell lines of pediatric tumors including 5 EwS cell lines and 22 from various histological origins were used. They were provided from the Innovative Therapies for Children with Cancer in Europe consortium cell line panel [48] (Table S2), which was responsible for cell lines authentication. For functional studies, A673 EwS cell line and the mouse embryonic fibroblast cells NIH-3T3 were used. NIH-3T3 cell line was previously purchased from ATCC collection and not retested for authentication. DMEM or RPMI medium were supplemented with penicillin (100 U/ml), streptomycin (10 μg/ml) and 10% fetal calf serum (Life Technologies™, Saint Aubin, France) for all cell lines or 10% new born calf serum for NIH-3T3. Cell cultures were maintained at 37 °C in an atmosphere of 5% CO2 and 95% humidity and were systematically tested by PCR analysis to be free of mycoplasma.
Construction and establishment of cell lines stably expressing LMO3-BORCS5
The sequences of LMO3-BORCS5 v1 or LMO3-BORCS5 v2 were obtained after reverse transcription of the patient’s tumor RNA into cDNA followed by PCR amplification using the following primers: Forward: 5′-GCAACCGAAAGATCAAGG-3′ and Reverse: 5′-CATTTGTTGGGGAAGTCTG-3′. After purification of the amplicons, the sequences were cloned under the control of the CMV promoter into the pcDNA3.1 expression vector. Two plasmids were obtained: pcDNA3.1-LMO3-BORCS5 v1 and pcDNA3.1-LMO3-BORCS5 v2.
Then, A673 and NIH-3T3 cells were transfected with empty vector (pcDNA3.1) or with either pcDNA3.1-LMO3-BORCS5 v1 or pcDNA3.1-LMO3-BORCS5 v2 to obtain stable clones as previously described and selected with neomycine [49].
CRISPR/Cas9 knockout of BORCS5
BORCS5 gene was inactivated using the CRISPR/Cas9 system targeting the last exon containing the functional domain [50]. Using CRISPOR software v4.2 [51], two couples of 20-base pairs targeting sequences were designed and synthetized (Eurofins Genomics, Ebensburg, Germany). Each guide from a pair was respectively inserted into pLentiCRISPR V2 (Addgene 52961) containing either GFP or CFP instead of Puromycine [pLentiCRISPR V2-GFP and pLentiCRISPR V2-CFP, provided by Julie Rivière (INSERM U1170, Gustave Roussy)].
A673 cells were co-transfected with both plasmids, from each pair (2 co-transfections). After 27 h, GFP and CFP-positive cells were selected using FACSAria™ Fusion cell sorter (BD Biosciences, Le Pont de Claix, France) and reseeded at a concentration of one cell/well to allow single colony formation. After 10 days, genomic DNA was extracted from individual colonies using QuickExtract™ DNA Extraction solution (Lucigen, Middleton, USA). A total of 12 clones were obtained for the first pair of guides, and six clones for the second pair. After PCR testing for deletion of the target sequence using the following primers: Forward 5′-GAGCTGTTTTCTCCTGGCCT-3′ and Reverse 5′-TTCATGCTGAAGGGCTCCAG-3′, two positive clones were obtained only for one pair of guides with the following sequences: guide 1: 5′-AGAGAGACCGCCATCGTGTT-3′ and guide 2: 5′-CTGGATCTGCTCGGCATACT-3′.
Reverse transcription-quantitative PCR (RT-qPCR)
RNA extraction, reverse transcription, and real-time PCR (qPCR) were performed as previously described [49]. Primers were either purchased from Invitrogen or Eurofins Genomics (Ebersberg, Germany) listed in Tables S1 and S3, or included in a personalized Human qPCR SignArrays® [52] from AnyGenes (Paris, France).
RT-qPCR was performed for LMO3-BORCS5 and the LMO3 and BORCS5 wild-type genes in (i) patient’s tumor sample at diagnosis and relapse, (ii) the 27 various cell lines (Table S2), and (iii) A673 or NIH-3T3 stably transfected with pcDNA3.1 (empty vector) or pcDNA3.1-LMO3-BORCS5 v1 (referred to as “+LMO3-BORCS5”). For LMO3-BORCS5, the amplicon size was verified by electrophoresis in 2% agarose gel. PCR products were purified with DNA extraction with NucleoSpin® Gel and PCR clean-up extraction kit (Macherey-Nagel, Dueren, Germany) and sequenced by Sanger technique (Eurofins, Cochin Institute, Paris, France).
Genes involved in apoptosis BAX; NOXA; P21; TP53 in A673 and those sorted out from microarray analysis for validation (ABCC3; FGF1; IGF2; PAX7; PRKCA; WNT6; ALK; CUX2; FGF13; FRK; LZTS1; MMP10; MYOD1; NRP1; NTRK1; RHOJ; SEMA3C) were also assessed (Table S3).
For all experiments, gene expression was determined by quantification-comparative 2−ΔΔCt method [53] and normalized to GAPDH levels. The results are expressed as relative mRNA levels and represent at least three independent experiments.
Doubling time and cell-cycle analysis
Cell-cycle analysis by flow cytometry and doubling time of the different clones were assessed as previously described [49]. All samples were tested in triplicates and at least n = 3 experiments are represented.
Cell migration assays
Scratch test was performed to evaluate cell migration capacity as previously described [49]. The results are presented as wound closure time in hours per cell line. All samples were tested in triplicates and n = 3 experiments were performed.
Cell survival assays
Cell line A673 was seeded in 96-well plates at 25,000 cells/well. After 72 h of incubation with increased concentrations from 0 to 10 µM of vincristine, cyclophosphamide, mafosfamide, etoposide, actinomycin, dasatinib, SN-38, doxorubicin, bortezomib, pazopanib, or everolimus, MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide] test was performed as previously described [39]. The half maximal inhibitory concentration (IC50) of cell viability was determined by linear regression model. Results are the mean ± SD of at least two independent experiments containing three replicates for each condition.
Annexin V apoptosis assay
Apoptosis was determined by flow cytometry using Annexin V apoptosis detection kit (BD Biosciences, Le Pont de Claix, France) according to the manufacturer’s instructions. Each cell line was seeded in 6-well plates (3 × 105 cells/well) and treated with vincristine (10 nM) or SN-38 (20 µM). After 48 h, cells were collected and analyzed as previously described [49]. n = 3 experiments were performed in triplicate and data represent the percentage of apoptotic cells compared with non-treated cells.
Microarray analysis
The comparison of gene expressions between the two clones (C1 and C2) obtained from A673 + empty vector and A673 + LMO3-BORCS5 was assessed by microarray analysis as described previously [39]. Four independent RNA extractions were performed for each clone using RNeasy mini-kit (Qiagen, Courtaboeuf, France) following the manufacturer’s instructions.
The mRNA was labeled using fluorescent low-input linear amplification kit (Agilent Technologies, Massy, France). Briefly, reverse transcription was performed using M-MLV reverse transcriptase. Then, cyanine 3-labeled cDNAs were generated using the one-color Agilent labeling kit (Low-Input Quick Amp Labeling Kit 5190–2306) adapted for small amount of total RNA (100 ng total RNA per reaction). Hybridization was then performed on microarray using 800 ng of linearly amplified cRNA labeled, following the manufacturer protocol (Agilent SureHyb Chamber; 800 ng of labeled extract; duration of hybridization of 17 h; 40 µL per array; Temperature of 65 °C) on Agilent® SurePrint G3 Human GE 8 × 60 K Microarray (Agilent Technologies, AMADID 39494). After washing in acetonitrile, slides were scanned by using an Agilent G2565 C DNA microarray scanner with defaults parameters (100° PMT, 3 µm resolution, at 20 °C in free ozone concentration environment. Microarray images were analyzed by using Feature Extraction software version (10.7.3.1) from Agilent technologies. Defaults settings were used.
The microarray data have been submitted to the Array Express data repository at the European Bioinformatics Institute (http://www.ebi.ac.uk/arrayexpress/), under the accession number: E-MTAB-7021.
The “Linear Model for Microarray” (Lima) methodology was used for the analysis [54]. P-values were adjusted using Benjamin and Hochberg procedure to control the false discovery rate [55]. We defined up- or down-regulation as ratios greater than twofold between A673 + empty vector and A673 + LMO3-BORCS5, and an adjusted p-value (FDR) < 0.05. To interpret the biological meaning genomic data, we used the ToppGene Suite (https://toppgene.cchmc.org/enrichment.jsp) and Ingenuity (www.ingenuity.com) software.
Immunoblotting
Total proteins were extracted from A673 WT and A673 KO BORCS5 cell lines as previously described [49]. The following primary antibodies were used: monoclonal rabbit BORCS5 (1:1000, PAS-5821, Invitrogen, Carlsbad, CA, USAs) and GAPDH-HRP as internal control (1:1,000; Sigma-Aldrich Chemicals Co., Saint Quentin Fallavier, France). Blots were washed and incubated with corresponding secondary antibodies: anti-rabbit-HRP (1:3,000; Cell Signaling technology, Saint Quentin en Yvelines, France). Bands were revealed by enhanced chemiluminescence reagent (HRP system, Clarity™; Bio-Rad Laboratories, Marnes-la-Coquette, France) using ChemiDoc™ Imaging System (Bio-Rad, Hercules, CA, USA).
Animal studies and tumorigenicity tests
All animal experiments were carried out according to French laws (Directive 2010/63/UE) and approved by the institutional Ethics Committee of Animal Experimentation (CEEA): authorization number CEEA IRCIV/IGR no. 26: 94–226, no. 2011–09.
Six-week-old nude nu/nu female mice were used to test the tumorigenicity of (i) A673 empty vector versus A673 + LMO3-BORCS5 v1; (ii) NIH-3T3 + empty vector versus NIH-3T3 + LMO3-BORCS5 v1; (iii) A673 WT versus A673 KO BORCS5.
All cell lines were diluted in 100 μl of PBS and injected subcutaneously into the flank of mice (n = 5/group) at a rate of 10 × 106 cells/mouse. Groups of mice were randomly selected for the different injections. Mice were monitored (unblinded test) for tumor growth and sacrificed when tumors reached a volume of ~1000 mm3. Half of each tumor was immediately frozen in liquid nitrogen for RT-qPCR, the other half was conserved in FineFix Formalin-Free Fixative (Fisher Scientific, Illkirch, France) for histochemistry analysis. For this latter analysis, n = 2 tumor tissues per group were embedded in paraffin, sections (4-μm thick) were prepared and stained with HES. Sections were examined with Zeiss-Axiophot microscope (Microscopy and Imaging center, Texas, USA).
Statistical analysis
Data are presented as mean ± SD.
For in vitro studies, the experiments have been repeated at least three times and each experiment was done in duplicate or triplicate.
For animal experimentation, analysis of sample size against power indicated that we would have needed at least five mice per group to reach an acceptable power of 0.9 (analysis performed by Statistica V12). Therefore, we included five animals per group, thanks to the small intragroup variations.
By using GraphPad Prism 4 software, Mann–Whitney test was used to compare two groups of treatments and Kruskal–Wallis test followed by Dunn’s test for multiple treatments. Data meet the assumptions of the tests. The estimate of variation within each group of data was <10% and the variance was similar between the groups statistically compared. Doubling time was calculated according to an exponential regression equation, given as: y = axebx, via the formula: t1/2 = ln(2)/b. p < 0.05 was considered as statistically significant. *p < 0.05; **p < 0.01; ***p < 0.001.
References
Mitelman F, Johansson B, Mertens F. The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer. 2007;7:233–45.
Dupain C, Harttrampf AC, Urbinati G, Geoerger B, Massaad-Massade L. Relevance of fusion genes in pediatric cancers: toward precision medicine. Mol Ther Nucleic Acids. 2017;6:315–26.
Arpinati M, Tolomelli G, Bochicchio MT, Castagnetti F, Amabile M, Bandini G, et al. Molecular monitoring of BCR-ABL transcripts after allogeneic stem cell transplantation for chronic myeloid leukemia. Biol Blood Marrow Transplant. 2013;19:735–40.
Cachia D, Wani K, Penas-Prado M, Olar A, McCutcheon IE, Benjamin RS, et al. C11orf95–RELA fusion present in a primary supratentorial ependymoma and recurrent sarcoma. Brain Tumor Pathol. 2015;32:105–11.
Graham RP, Jin L, Knutson DL, Kloft-Nelson SM, Greipp PT, Waldburger N, et al. DNAJB1-PRKACA is specific for fibrolamellar carcinoma. Mod Pathol. 2015;28:822–9.
Han S, Brenner JC, Sabolch A, Jackson W, Speers C, Wilder-Romans K, et al. Targeted radiosensitization of ETS fusion-positive prostate cancer through PARP1 inhibition. Neoplasia. 2013;15:1207–17.
Hartmaier RJ, Trabucco SE, Priedigkeit N, Chung JH, Parachoniak CA, Vanden Borre P, et al. Recurrent hyperactive ESR1 fusion proteins in endocrine therapy-resistant breast cancer. Ann Oncol. 2018;29:872–80.
Kalinova M, Krskova L, Brizova H, Kabickova E, Kepak T, Kodet R. Quantitative PCR detection of NPM/ALK fusion gene and CD30 gene expression in patients with anaplastic large cell lymphoma–residual disease monitoring and a correlation with the disease status. Leuk Res. 2008;32:25–32.
Kumar-Sinha C, Kalyana-Sundaram S, Chinnaiyan AM. Landscape of gene fusions in epithelial cancers: seq and ye shall find. Genome Med. 2015;7:129.
PANAGOPOULOS I, GORUNOVA L, BJERKEHAGEN B, BOYE K, HEIM S. Chromosome aberrations and HEY1-NCOA2 fusion gene in a mesenchymal chondrosarcoma. Oncol Rep. 2014;32:40–44.
Lawrence MS, Stojanov P, Mermel CH, Robinson JT, Garraway LA, Golub TR, et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature. 2014;505:495–501.
Dancey Janet E, Bedard Philippe L, Onetto N, Hudson Thomas J. The genetic basis for cancer treatment decisions. Cell. 2012;148:409–20.
Albanese A, Tang PS, Chan WC. The effect of nanoparticle size, shape, and surface chemistry on biological systems. Annu Rev Biomed Eng. 2012;14:1–16.
Harttrampf AC, Lacroix L, Deloger M, Deschamps F, Puget S, Auger N, et al. Molecular screening for cancer treatment optimization (MOSCATO-01) in pediatric patients: a single institutional prospective molecular stratification trial. Clin Cancer Res. 2017;23:6101–12.
Massard C, Michiels S, Ferté C, Le Deley M-C, Lacroix L, Hollebecque A, et al. High-throughput genomics and clinical outcome in hard-to-treat advanced cancers: results of the MOSCATO 01 trial. Cancer Discov. 2017;7:586–95.
Dupain C, Harttrampf AC, Boursin Y, Lebeurrier M, Rondof W, Robert-Siegwald G, et al. Discovery of new fusion transcripts in a cohort of pediatric solid cancers at relapse and relevance for personalized medicine. Mol Ther. 2019;27:200–18.
Sang M, Ma L, Sang M, Zhou X, Gao W, Geng C. LIM-domain-only proteins: multifunctional nuclear transcription coregulators that interacts with diverse proteins. Mol Biol Rep. 2014;41:1067–73.
Aoyama M, Ozaki T, Inuzuka H, Tomotsune D, Hirato J, Okamoto Y, et al. LMO3 interacts with neuronal transcription factor, HEN2, and acts as an oncogene in neuroblastoma. Cancer Res. 2005;65:4587–97.
Larsen S, Yokochi T, Isogai E, Nakamura Y, Ozaki T, Nakagawara A. LMO3 interacts with p53 and inhibits its transcriptional activity. Biochem Biophys Res Commun. 2010;392:252–7.
Qiu Y-S, Jiang N-N, Zhou Y, Yu K-Y, Gong H-Y, Liao G-J. LMO3 promotes gastric cancer cell invasion and proliferation through Akt-mTOR and Akt-GSK3β signaling. Int J Mol Med. 2018;41:2755–63.
Song Y-F, Hong J-F, Liu D-L, Lin Q-A, Lan X-P, Lai G-X. miR-630 targets LMO3 to regulate cell growth and metastasis in lung cancer. Am J Transl Res. 2015;7:1271–9.
Bao J, Talmage DA, Role LW, Gautier J. Regulation of neurogenesis by interactions between HEN1 and neuronal LMO proteins. Development. 2000;127:425–35.
Langemeyer L, Ungermann C. BORC and BLOC-1: shared subunits in trafficking complexes. Dev Cell. 2015;33:121–2.
Niwa S, Tao L, Lu SY, Liew GM, Feng W, Nachury MV, et al. BORC regulates the axonal transport of synaptic vesicle precursors by activating ARL-8. Curr Biol. 2017;27:2569–.e2564.
Pu J, Schindler C, Jia R, Jarnik M, Backlund P, Bonifacino Juan S. BORC, a multisubunit complex that regulates lysosome positioning. Dev Cell. 2015;33:176–88.
Montpetit A, Boily G, Sinnett D. A detailed transcriptional map of the chromosome 12p12 tumour suppressor locus. Eur J Hum Genet. 2002;10:62–71.
Latysheva NS, Babu MM. Discovering and understanding oncogenic gene fusions through data intensive computational approaches. Nucleic Acids Res. 2016;44:4487–503.
Mertens F, Johansson B, Fioretos T, Mitelman F. The emerging complexity of gene fusions in cancer. Nat Rev Cancer. 2015;15:371–81.
Shugay M, Ortiz de Mendíbil I, Vizmanos JL, Novo FJ. Genomic hallmarks of genes involved in chromosomal translocations in hematological cancer. PLoS Comput Biol. 2012;8:e1002797.
Shugay M, Ortiz de Mendíbil I, Vizmanos JL, Novo FJ. Oncofuse: a computational framework for the prediction of the oncogenic potential of gene fusions. Bioinformatics. 2013;29:2539–46.
Frenkel-Morgenstern M, Lacroix V, Ezkurdia I, Levin Y, Gabashvili A, Prilusky J, et al. Chimeras taking shape: potential functions of proteins encoded by chimeric RNA transcripts. Genome Res. 2012;22:1231–42.
Keller C, Arenkiel BR, Coffin CM, El-Bardeesy N, DePinho RA, Capecchi MR. Alveolar rhabdomyosarcomas in conditional Pax3:Fkhr mice: cooperativity of Ink4a/ARF and Trp53 loss of function. Genes Dev. 2004;18:2614–26.
Lin PP, Pandey MK, Jin F, Xiong S, Deavers M, Parant JM, et al. EWS-FLI1 induces developmental abnormalities and accelerates sarcoma formation in a transgenic mouse model. Cancer Res. 2008;68:8968–75.
Isogai E, Ohira M, Ozaki T, Oba S, Nakamura Y, Nakagawara A. Oncogenic LMO3 collaborates with HEN2 to enhance neuroblastoma cell growth through transactivation of Mash1. PloS ONE. 2011;6:e19297.
Tsubota S, Kadomatsu K. Origin and mechanism of neuroblastoma. Oncoscience. 2017;4:70–72.
Terano T, Zhong Y, Toyokuni S, Hiai H, Yamada Y. Transcriptional control of fetal liver hematopoiesis: dominant negative effect of the overexpression of the LIM domain mutants of LMO2. Exp Hematol. 2005;33:641–51.
Cooper GM. The development and causes of cancer. The cell: a molecular approach. 2nd ed. 2000. Sunderland, MA: Sinauer Associates.
Ali HM, Urbinati G, Chapuis H, Desmaele D, Bertrand J-R, Couvreur P, et al. Effects of siRNA on RET/PTC3 junction oncogene in papillary thyroid carcinoma: from molecular and cellular studies to preclinical investigations. PloS ONE. 2014;9:e95964.
Urbinati G, Ali HM, Rousseau Q, Chapuis H, Desmaële D, Couvreur P, et al. AntIneoplastic effects of siRNA against TMPRSS2-ERG junction oncogene in prostate cancer. PloS ONE. 2015;10:e0125277.
Goel RK, Lukong KE. Understanding the cellular roles of Fyn-related kinase (FRK): implications in cancer biology. Cancer Metastas Rev. 2016;35:179–99.
Teles Alves I, Hartjes T, McClellan E, Hiltemann S, Böttcher R, Dits N, et al. Next-generation sequencing reveals novel rare fusion events with functional implication in prostate cancer. Oncogene. 2015;34:568–77.
Abate F, Zairis S, Ficarra E, Acquaviva A, Wiggins CH, Frattini V, et al. Pegasus: a comprehensive annotation and prediction tool for detection of driver gene fusions in cancer. BMC Syst Biol. 2014;8:97.
Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C. The landscape of kinase fusions in cancer. Nat Commun. 2014;5:4846.
Tirode F, Surdez D, Ma X, Parker M, Le Deley MC, Bahrami A, et al. Genomic landscape of ewing sarcoma defines an aggressive subtype with co-association of STAG2 and TP53 mutations. Cancer Discov. 2014;4:1342–53.
Crompton BD, Stewart C, Taylor-Weiner A, Alexe G, Kurek KC, Calicchio ML, et al. The genomic landscape of pediatric ewing sarcoma. Cancer Discov. 2014. CD-13-1037. https://doi.org/10.1158/2159-8290.CD-13-1037.
Gao Q, Liang W-W, Foltz SM, Mutharasu G, Jayasinghe RG, Cao S, et al. Driver fusions and their implications in the development and treatment of human cancers. Cell Rep. 2018;23:227–.e223.
Anderson ND, de Borja R, Young MD, Fuligni F, Rosic A, Roberts ND, et al. Rearrangement bursts generate canonical gene fusions in bone and soft tissue tumors. Science. 2018;361:eaam8419. pii.
Daudigeos-Dubus E, Le Dret L, Lanvers-Kaminsky C, Bawa O, Opolon P, Vievard A, et al. Regorafenib: antitumor activity upon mono and combination therapy in preclinical pediatric malignancy models. PloS ONE. 2015;10:e0142612.
Dupain C, Ali HM, Mouhoub TA, Urbinati G, Massaad-Massade L. Induction of TTF-1 or PAX-8 expression on proliferation and tumorigenicity in thyroid carcinomas. Int J Oncol. 2016;49:1248–58.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23.
Haeussler M, Schönig K, Eckert H, Eschstruth A, Mianné J, Renaud J-B, et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 2016;17:148.
Buart S, Terry S, Noman MZ, Lanoy E, Boutros C, Fogel P, et al. Transcriptional response to hypoxic stress in melanoma and prognostic potential of GBE1 and BNIP3. Oncotarget. 2017;8:108786–801.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25:402–8.
Jeanmougin M, de Reynies A, Marisa L, Paccard C, Nuel G, Guedj M. Should we abandon the t-test in the analysis of gene expression microarray data: a comparison of variance modeling strategies. PloS ONE. 2010;5:e12336.
Benjamini Y, Heller R. Screening for partial conjunction hypotheses. Biometrics. 2008;64:1215–22.
Zhao Y, et al. ABCC3 as a marker for multidrug resistance in non-small cell lung cancer. Sci Rep. 2013;3:3120.
Yau NKM, et al. A Pan-Cancer Review of ALK Mutations: Implications for Carcinogenesis and Therapy. Curr Cancer Drug Targets. 2015;15:327–36.
Duan C, et al. Whole exome sequencing reveals novel somatic alterations in neuroblastoma patients with chemotherapy. Cancer Cell Int. 2018;18:21.
Li J, et al. Clinicopathological significance of fibroblast growth factor 1 in non-small cell lung cancer. Hum Pathol. 2015;46:1821–8.
Okada T, et al. Upregulated expression of FGF13/FHF2 mediates resistance to platinum drugs in cervical cancer cells. Sci Rep. 2013;3:2899.
Goel RK, Lukong KE. Understanding the cellular roles of Fyn-related kinase (FRK): implications in cancer biology. Cancer Metastasis Rev. 2016;35:179–99.
Brouwer-Visser J, Huang GS. IGF2 signaling and regulation in cancer. Cytokine Growth Factor Rev. 2015;26:371–7.
Zhou W, et al. The tumor-suppressor gene LZTS1 suppresses colorectal cancer proliferation through inhibition of the AKT-mTOR signaling pathway. Cancer Lett. 2015;360:68–75.
Justilien V, et al. Matrix metalloproteinase-10 is required for lung cancer stem cell maintenance, tumor initiation and metastatic potential. PLoS ONE. 2012;7:e35040.
Rekhi B, Upadhyay P, Ramteke MP, Dutt A. MYOD1 (L122R) mutations are associated with spindle cell and sclerosing rhabdomyosarcomas with aggressive clinical outcomes. Mod Pathol. 2016;29:1532–40.
Chaudhary B, Khaled YS, Ammori BJ, Elkord E. Neuropilin 1: function and therapeutic potential in cancer. Cancer Immunol Immunother. 2014;63:81–99.
Vaishnavi A, et al. Oncogenic and drug-sensitive NTRK1 rearrangements in lung cancer. Nat Med. 2013;19:1469–72.
Parveen A, Akash MSH, Rehman K, Kyunn WW. Dual Role of p21 in the Progression of Cancer and Its Treatment. Crit Rev Eukaryot Gene Expr. 2016;26:49–62.
Barr FG. Gene fusions involving PAX and FOX family members in alveolar rhabdomyosarcoma. Oncogene. 2001;20:5736–46.
Bridge JA, et al. Identification of a novel, recurrent SLC44A1-PRKCA fusion in papillary glioneuronal tumor. Brain Pathol. 2013;23:121–8.
Kim C, et al. Rho GTPase RhoJ is Associated with Gastric Cancer Progression and Metastasis. J Cancer. 2016;7:1550–6.
Man J, et al. Sema3C promotes the survival and tumorigenicity of glioma stem cells through Rac1 activation. Cell Rep. 2014;9:1812–26.
Zhang L, et al. Increased WNT6 expression in tumor cells predicts unfavorable survival in esophageal squamous cell carcinoma patients. Int J Clin Exp Pathol. 2015;8:11421–7.
Acknowledgements
This study was supported by Université Paris-Sud (CD) and CNRS and microarray analysis was supported by the “Taxe d’apprentissage Gustave Roussy - 2017 – CD”. We acknowledge Drs Nathalie Gaspar and Estelle Daudigeos-Dubus as well as the ITCC (Innovative Therapies for Children with Cancer) consortium for providing the pediatric cell lines and antineoplastic molecules used in the study, as well as Catherine Richon (technical assistance). We acknowledge Dr Giorgia Urbinati and Pr. Karim Benihoud for the valuable comments as well as Léna Zig and Linda Ramdani for their precious technical help for the cloning of LMO3-BORCS5. We also acknowledge Dr Guillaume Meurice for the analysis of microarray data. The animal studies were performed within the animal facility of Gustave Roussy and authors gratefully thank the technical support of Olivia Bawa and expertize of Dr Paule Opolon (histocytopathology platform) for the preparation and analysis of histological slides.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dupain, C., Gracia, C., Harttrampf, A.C. et al. Newly identified LMO3-BORCS5 fusion oncogene in Ewing sarcoma at relapse is a driver of tumor progression. Oncogene 38, 7200–7215 (2019). https://doi.org/10.1038/s41388-019-0914-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0914-3