Abstract
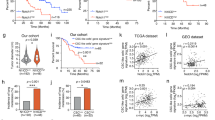
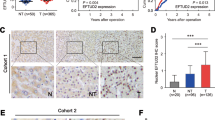
The poorly differentiated hepatocellular carcinoma (HCC) cells are usually characterized by immature hepatic progenitor cell-like properties, such as enhanced self-renewal ability, resistance to chemotherapeutic drugs, and a loss of mature hepatocyte proteins. However, the molecular mechanisms governing this process still remain unclear. In this study, we found the lymphoid enhancer-binding factor-1 (LEF1), a transcriptional factor, was frequently overexpressed in HCCs, which was significantly associated with poor prognosis and tumor cell differentiation. Functional studies have found that LEF1 enhanced cell growth, foci formation, colony formation in soft agar, and tumor formation in nude mice. Different from its canonical roles in the WNT signaling pathway, we found that LEF1 could activate the critical members (e.g., NOTCH1 and NOTCH2) of the NOTCH signaling pathway through directly binding to their promoter regions. Further studies have found that LEF1 could enhance the self-renewal ability, drug resistance, dedifferentiation, and invasion of HCC cells. The oncogenic functions and the effects of LEF1 on cancer stemness could be effectively inhibited by NOTCH inhibitor. Further characterization of LEF1 may lead to the development of novel therapeutic strategies for HCC treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Tong MJ, Chavalitdhamrong D, Lu DS, Raman SS, Gomes A, Duffy JP, et al. Survival in Asian Americans after treatments for hepatocellular carcinoma: a seven-year experience at UCLA. J Clin Gastroenterol. 2010;44:e63–70.
Dhanasekaran R, Bandoh S, Roberts LR. Molecular pathogenesis of hepatocellular carcinoma and impact of therapeutic advances. F1000Res. 2016;5:879.
Okamura RM, Sigvardsson M, Galceran J, Verbeek S, Clevers H, Grosschedl R. Redundant regulation of T cell differentiation and TCRalpha gene expression by the transcription factors LEF-1 and TCF-1. Immunity. 1998;8:11–20.
Reya T, O’Riordan M, Okamura R, Devaney E, Willert K, Nusse R, et al. Wnt signaling regulates B lymphocyte proliferation through a LEF-1 dependent mechanism. Immunity. 2000;13:15–24.
Wu W, Zhu H, Fu Y, Shen W, Miao K, Hong M, et al. High LEF1 expression predicts adverse prognosis in chronic lymphocytic leukemia and may be targeted by ethacrynic acid. Oncotarget. 2016;7:21631–43.
Petropoulos K, Arseni N, Schessl C, Stadler CR, Rawat VP, Deshpande AJ, et al. A novel role for Lef-1, a central transcription mediator of Wnt signaling, in leukemogenesis. J Exp Med. 2008;205:515–22.
Staal FJ, Luis TC, Tiemessen MM. WNT signalling in the immune system: WNT is spreading its wings. Nat Rev Immunol. 2008;8:581–93.
Clevers H. Wnt/beta-catenin signaling in development and disease. Cell. 2006;127:469–80.
McGregor SM The role of Lef1 in T cell development and lymphomagenesis[J]. Dissertations & Theses - Gradworks, 2009.
Visvader JE, Lindeman GJ. Cancer stem cells: current status and evolving complexities. Cell Stem Cell. 2012;10:717–28.
Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009;137:216–33.
Ungerback J, Elander N, Grunberg J, Sigvardsson M, Soderkvist P. The Notch-2 gene is regulated by Wnt signaling in cultured colorectal cancer cells. PLoS ONE 2011;6:e17957.
Heitzler P, Bourouis M, Ruel L, Carteret C, Simpson P. Genes of the Enhancer of split and achaete-scute complexes are required for a regulatory loop between Notch and Delta during lateral signalling in Drosophila. Development. 1996;122:161–71.
Leimeister C, Externbrink A, Klamt B, Gessler M. Hey genes: a novel subfamily of hairy- and Enhancer of split related genes specifically expressed during mouse embryogenesis. Mech Dev. 1999;85:173–7.
Kabos P, Kabosova A, Neuman T. Blocking HES1 expression initiates GABAergic differentiation and induces the expression ofp21(CIP1/WAF1) in human neural stem cells. J Biol Chem. 2002;277:8763–6.
Weng AP, Millholland JM, Yashiro-Ohtani Y, Arcangeli ML, Lau A, Wai C, et al. c-Myc is an important direct target of Notch1 in T-cell acute lymphoblastic leukemia/lymphoma. Genes Dev. 2006;20:2096–109.
Oswald F, Liptay S, Adler G, Schmid RM. NF-kappaB2 is a putative target gene of activated Notch-1 via RBP-Jkappa. Mol Cell Biol. 1998;18:2077–88.
Nakahara F, Kitaura J, Uchida T, Nishida C, Togami K, Inoue D, et al. Hes1 promotes blast crisis in chronic myelogenous leukemia through MMP-9 upregulation in leukemic cells. Blood. 2014;123:3932–42.
Lai EC. Notch signaling: control of cell communication and cell fate. Development. 2004;131:965–73.
Kopan R. Notch: a membrane-bound transcription factor. J Cell Sci. 2002;115:1095–7.
Kohn A, Rutkowski TP, Liu Z, Mirando AJ, Zuscik MJ, O’Keefe RJ, et al. Notch signaling controls chondrocyte hypertrophy via indirect regulation of Sox9. Bone Res. 2015;3:15021.
Sekiya S, Suzuki A. Intrahepatic cholangiocarcinoma can arise from Notch-mediated conversion of hepatocytes. J Clin Invest. 2012;122:3914–8.
Valenti L, Mendoza RM, Rametta R, Maggioni M, Kitajewski C, Shawber CJ, et al. Hepatic notch signaling correlates with insulin resistance and nonalcoholic fatty liver disease. Diabetes. 2013;62:4052–62.
Si-Tayeb K, Lemaigre FP, Duncan SA. Organogenesis and development of the liver. Dev Cell. 2010;18:175–89.
Mishra L, Banker T, Murray J, Byers S, Thenappan A, He AR, et al. Liver stem cells and hepatocellular carcinoma. Hepatology. 2009;49:318–29.
Park HJ, Choi YJ, Kim JW, Chun HS, Im I, Yoon S, et al. Differences in the epigenetic regulation of cytochrome P450 genes between human embryonic stem cell-derived hepatocytes and primary hepatocytes. PLoS ONE 2015;10:e0132992.
Liu M, Chen L, Ma NF, Chow RK, Li Y, Song Y, et al. CHD1L promotes lineage reversion of hepatocellular carcinoma through opening chromatin for key developmental transcription factors. Hepatology. 2016;63:1544–59.
Cook JJ, Wildsmith KR, Gilberto DB, Holahan MA, Kinney GG, Mathers PD, et al. Acute gamma-secretase inhibition of nonhuman primate CNS shifts amyloid precursor protein (APP) metabolism from amyloid-beta production to alternative APP fragments without amyloid-beta rebound. J Neurosci. 2010;30:6743–50.
Kim K, Lu Z, Hay ED. Direct evidence for a role of beta-catenin/LEF-1 signaling pathway in induction of EMT. Cell Biol Int. 2002;26:463–76.
Reya T, Clevers H. Wnt signalling in stem cells and cancer. Nature. 2005;434:843–50.
Zhang Y, Yu J, Shi C, Huang Y, Wang Y, Yang T, et al. Lef1 contributes to the differentiation of bulge stem cells by nuclear translocation and cross-talk with the Notch signaling pathway. Int J Med Sci. 2013;10:738–46.
Waterman ML. Lymphoid enhancer factor/T cell factor expression in colorectal cancer. Cancer Metastasis Rev. 2004;23:41–52.
Galceran J, Sustmann C, Hsu SC, Folberth S, Grosschedl R. LEF1-mediated regulation of Delta-like1 links Wnt and Notch signaling in somitogenesis. Genes Dev. 2004;18:2718–23.
Collu GM, Hidalgo-Sastre A, Brennan K. Wnt-Notch signalling crosstalk in development and disease. Cell Mol Life Sci. 2014;71:3553–67.
Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science. 1999;284:770–6.
Ranganathan P, Weaver KL, Capobianco AJ. Notch signalling in solid tumours: a little bit of everything but not all the time. Nat Rev Cancer. 2011;11:338–51.
Ji J, Wang XW. Clinical implications of cancer stem cell biology in hepatocellular carcinoma. Semin Oncol. 2012;39:461–72.
Ma NF, Hu L, Fung JM, Xie D, Zheng BJ, Chen L, et al. Isolation and characterization of a novel oncogene, amplified in liver cancer 1, within a commonly amplified region at 1q21 in hepatocellular carcinoma. Hepatology. 2008;47:503–10.
Chen L, Hu L, Chan TH, Tsao GS, Xie D, Huo KK, et al. Chromodomain helicase/adenosine triphosphatase DNA binding protein 1-like (CHD1l) gene suppresses the nucleus-to-mitochondria translocation of nur77 to sustain hepatocellular carcinoma cell survival. Hepatology. 2009;50:122–9.
Chen L, Chan TH, Yuan YF, Hu L, Huang J, Ma S, et al. CHD1L promotes hepatocellular carcinoma progression and metastasis in mice and is associated with these processes in human patients. J Clin Invest. 2010;120:1178–91.
Chen L, Yuan YF, Li Y, Chan TH, Zheng BJ, Huang J, et al. Clinical significance of CHD1L in hepatocellular carcinoma and therapeutic potentials of virus-mediated CHD1L depletion. Gut. 2011;60:534–43.
Ma S, Lee TK, Zheng BJ, Chan KW, Guan XY. CD133 + HCC cancer stem cells confer chemoresistance by preferential expression of the Akt/PKB survival pathway. Oncogene. 2008;27:1749–58.
Fu L, Qin YR, Xie D, Hu L, Kwong DL, Srivastava G, et al. Characterization of a novel tumor-suppressor gene PLC delta 1 at 3p22 in esophageal squamous cell carcinoma. Cancer Res. 2007;67:10720–6.
Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2017;46:D956–D96.
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19:649–58.
Funding
This work was supported by grants from the Hong Kong Research Grant Council (RGC), including GRF (17143716 and 767313), Collaborative Research Funds (C7038-14G and C7027-14G), Theme-based Research Scheme (T12-704/16-R), and National Natural Science Foundation of China (81772554, 81272416, and 81372583). This work was also supported by the Shenzhen Peacock Team Project (KQTD 2015033117210153). Professor XY Guan is a Sophie YM Chan Professor in Cancer Research.
Author information
Authors and Affiliations
Contributions
S.F. and X.Y.G. initiated and designed the study. S.F. wrote the manuscript with inputs from X.Y.G. and S.F. and M.L. designed the experiments and interpreted the results. S.F. performed all the experiments with assistance from M.L., L.L., F.F.Z., Y.L., Q.Y. and Y.Z.C., HCC clinical samples and the relevant clinical information were provided by Y.H.Z. and Y.F.Y. and X.Y.G. supervised the project.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Fang, S., Liu, M., Li, L. et al. Lymphoid enhancer-binding factor-1 promotes stemness and poor differentiation of hepatocellular carcinoma by directly activating the NOTCH pathway. Oncogene 38, 4061–4074 (2019). https://doi.org/10.1038/s41388-019-0704-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0704-y
This article is cited by
-
ZMIZ2 facilitates hepatocellular carcinoma progression via LEF1 mediated activation of Wnt/β-catenin pathway
Experimental Hematology & Oncology (2024)
-
SOX9 is a key component of RUNX2-regulated transcriptional circuitry in osteosarcoma
Cell & Bioscience (2023)
-
Notch signaling regulates a metabolic switch through inhibiting PGC-1α and mitochondrial biogenesis in dedifferentiated liposarcoma
Oncogene (2023)
-
The cancer-testis lncRNA LINC01977 promotes HCC progression by interacting with RBM39 to prevent Notch2 ubiquitination
Cell Death Discovery (2023)
-
Clinical Prognostic Implications of Wnt Hub Genes Expression in Medulloblastoma
Cellular and Molecular Neurobiology (2023)