Abstract
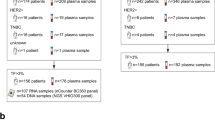
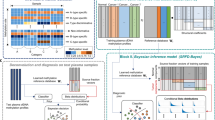
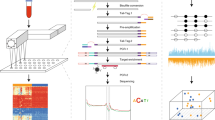
Blood circulating cell-free DNA (ccfDNA) is a suggested biosource of valuable clinical information for cancer, meeting the need for a minimally-invasive advancement in the route of precision medicine. In this paper, we evaluated the prognostic and predictive potential of ccfDNA parameters in early and advanced breast cancer. Groups consisted of 150 and 16 breast cancer patients under adjuvant and neoadjuvant therapy respectively, 34 patients with metastatic disease and 35 healthy volunteers. Direct quantification of ccfDNA in plasma revealed elevated concentrations correlated to the incidence of death, shorter PFS, and non-response to pharmacotherapy in the metastatic but not in the other groups. The methylation status of a panel of cancer-related genes chosen based on previous expression and epigenetic data (KLK10, SOX17, WNT5A, MSH2, GATA3) was assessed by quantitative methylation-specific PCR. All but the GATA3 gene was more frequently methylated in all the patient groups than in healthy individuals (all p < 0.05). The methylation of WNT5A was statistically significantly correlated to greater tumor size and poor prognosis characteristics and in advanced stage disease with shorter OS. In the metastatic group, also SOX17 methylation was significantly correlated to the incidence of death, shorter PFS, and OS. KLK10 methylation was significantly correlated to unfavorable clinicopathological characteristics and relapse, whereas in the adjuvant group to shorter DFI. Methylation of at least 3 or 4 genes was significantly correlated to shorter OS and no pharmacotherapy response, respectively. Classification analysis by a fully automated, machine learning software produced a single-parametric linear model using ccfDNA plasma concentration values, with great discriminating power to predict response to chemotherapy (AUC 0.803, 95% CI [0.606, 1.000]) in the metastatic group. Two more multi-parametric signatures were produced for the metastatic group, predicting survival and disease outcome. Finally, a multiple logistic regression model was constructed, discriminating between patient groups and healthy individuals. Overall, ccfDNA emerged as a highly potent predictive classifier in metastatic breast cancer. Upon prospective clinical evaluation, all the signatures produced could aid accurate prognosis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2018;68:394–424.
Pantel K, Alix-Panabieres C. Real-time liquid biopsy in cancer patients: fact or fiction? Cancer Res. 2013;73:6384–8.
Alix-Panabieres C, Pantel K. Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov. 2016;6:479–91.
Lamb YN, Dhillon S. Epi proColon® 2.0 CE: a blood-based screening test for colorectal cancer. Mol Diagn Ther. 2017;21:225–32.
Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer. 2011;11:426–37.
Elshimali YI, Khaddour H, Sarkissyan M, Wu Y, Vadgama JV. The clinical utilization of circulating cell free DNA (CCFDNA) in blood of cancer patients. Int J Mol Sci. 2013;14:18925–58.
Lu JL, Liang ZY. Circulating free DNA in the era of precision oncology: pre- and post-analytical concerns. Chronic Dis Transl Med. 2016;2:223–30.
Matthaios D, Balgkouranidou I, Karayiannakis A, Bolanaki H, Xenidis N, Amarantidis K, et al. Methylation status of the APC and RASSF1A promoter in cell-free circulating DNA and its prognostic role in patients with colorectal cancer. Oncol Lett. 2016;12:748–56.
Balgkouranidou I, Chimonidou M, Milaki G, Tsaroucha E, Kakolyris S, Georgoulias V, et al. SOX17 promoter methylation in plasma circulating tumor DNA of patients with non-small cell lung cancer. Clin Chem Lab Med. 2016;54:1385–93.
Mastoraki S, Strati A, Tzanikou E, Chimonidou M, Politaki E, Voutsina A, et al. ESR1 methylation: a liquid biopsy-based epigenetic assay for the follow-up of patients with metastatic breast cancer receiving endocrine treatment. Clin Cancer Res. 2018;24:1500–10.
Jiang P, Chan CW, Chan KC, Cheng SH, Wong J, Wong VW, et al. Lengthening and shortening of plasma DNA in hepatocellular carcinoma patients. Proc Natl Acad Sci USA. 2015;112:E1317–25.
Mouliere F, Rosenfeld N. Circulating tumor-derived DNA is shorter than somatic DNA in plasma. Proc Natl Acad Sci USA. 2015;112:3178–9.
Leon SA, Shapiro B, Sklaroff DM, Yaros MJ. Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res. 1977;37:646–50.
Chimonidou M, Strati A, Malamos N, Georgoulias V, Lianidou ES. SOX17 promoter methylation in circulating tumor cells and matched cell-free DNA isolated from plasma of patients with breast cancer. Clin Chem. 2013;59:270–9.
Jonsson M, Dejmek J, Bendahl PO, Andersson T. Loss of Wnt-5a protein is associated with early relapse in invasive ductal breast carcinomas. Cancer Res. 2002;62:409–16.
Yousef GM, Yacoub GM, Polymeris ME, Popalis C, Soosaipillai A, Diamandis EP. Kallikrein gene downregulation in breast cancer. Br J Cancer. 2004;90:167–72.
Kappil MA, Liao Y, Terry MB, Santella RM. DNA repair gene expression levels as indicators of breast cancer in the Breast Cancer Family Registry. Anticancer Res. 2016;36:4039–44.
McCleskey BC, Penedo TL, Zhang K, Hameed O, Siegal GP, Wei S. GATA3 expression in advanced breast cancer: prognostic value and organ-specific relapse. Am J Clin Pathol. 2015;144:756–63.
Tsamardinos I, Greasidou E, Borboudakis G. Bootstrapping the out-of-sample predictions for efficient and accurate cross-validation. Mach Learn. 2018;107:1895–922.
Catarino R, Ferreira MM, Rodrigues H, Coelho A, Nogal A, Sousa A, et al. Quantification of free circulating tumor DNA as a diagnostic marker for breast cancer. DNA Cell Biol. 2008;27:415–21.
Salvi S, Gurioli G, De Giorgi U, Conteduca V, Tedaldi G, Calistri D, et al. Cell-free DNA as a diagnostic marker for cancer: current insights. Onco Targets Ther. 2016;9:6549–59.
Tangvarasittichai O, Jaiwang W, Tangvarasittichai S. The plasma DNA concentration as a potential breast cancer screening marker. Indian J Clin Biochem. 2015;30:55–8.
Agassi R, Czeiger D, Shaked G, Avriel A, Sheynin J, Lavrenkov K, et al. Measurement of circulating cell-free DNA levels by a simple fluorescent test in patients with breast cancer. Am J Clin Pathol. 2015;143:18–24.
Olsson E, Winter C, George A, Chen Y, Howlin J, Tang MH, et al. Serial monitoring of circulating tumor DNA in patients with primary breast cancer for detection of occult metastatic disease. EMBO Mol Med. 2015;7:1034–47.
Tan G, Chu C, Gui X, Li J, Chen Q. The prognostic value of circulating cell-free DNA in breast cancer: a meta-analysis. Medicine. 2018;97:e0197.
Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013;368:1199–209.
Anker P, Stroun M, Maurice PA. Spontaneous release of DNA by human blood lymphocytes as shown in an in vitro system. Cancer Res. 1975;35:2375–82.
Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313:139–42.
Laktionov PP, Tamkovich SN, Rykova EY, Bryzgunova OE, Starikov AV, Kuznetsova NP, et al. Extracellular circulating nucleic acids in human plasma in health and disease. Nucleosides Nucleotides Nucleic Acids. 2004;23:879–83.
Madhavan D, Wallwiener M, Bents K, Zucknick M, Nees J, Schott S, et al. Plasma DNA integrity as a biomarker for primary and metastatic breast cancer and potential marker for early diagnosis. Breast Cancer Res Treat. 2014;146:163–74.
Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN. Fragment length of circulating tumor. DNA. 2016;12:e1006162.
Umetani N, Giuliano AE, Hiramatsu SH, Amersi F, Nakagawa T, Martino S, et al. Prediction of breast tumor progression by integrity of free circulating DNA in serum. J Clin Oncol. 2006;24:4270–6.
Leris AC, Roberts TR, Jiang WG, Newbold RF, Mokbel K. WNT5A expression in human breast cancer. Anticancer Res. 2005;25:731–4.
Trifa F, Karray-Chouayekh S, Jmal E, Jmaa ZB, Khabir A, Sellami-Boudawara T, et al. Loss of WIF-1 and Wnt5a expression is related to aggressiveness of sporadic breast cancer in Tunisian patients. Tumour Biol. 2013;34:1625–33.
Fu DY, Wang ZM, Li C, Wang BL, Shen ZZ, Huang W, et al. Sox17, the canonical Wnt antagonist, is epigenetically inactivated by promoter methylation in human breast cancer. Breast Cancer Res Treat. 2010;119:601–12.
Fu D, Ren C, Tan H, Wei J, Zhu Y, He C, et al. Sox17 promoter methylation in plasma DNA is associated with poor survival and can be used as a prognostic factor in breast cancer. Medicine. 2015;94:e637.
Kioulafa M, Kaklamanis L, Stathopoulos E, Mavroudis D, Georgoulias V, Lianidou ES. Kallikrein 10 (KLK10) methylation as a novel prognostic biomarker in early breast cancer. Ann Oncol. 2009;20:1020–5.
Markaki M, Tsamardinos I, Langhammer A, Lagani V, Hveem K, Roe OD. A validated clinical risk prediction model for lung cancer in smokers of all ages and exposure types: a HUNT study. EBioMedicine. 2018;31:36–46.
Orfanoudaki G, Markaki M, Chatzi K, Tsamardinos I, Economou A. MatureP: prediction of secreted proteins with exclusive information from their mature regions. Sci Rep. 2017;7:3263.
Li Y, Melnikov AA, Levenson V, Guerra E, Simeone P, Alberti S, et al. A seven-gene CpG-island methylation panel predicts breast cancer progression. BMC Cancer. 2015;15:417.
List M, Hauschild AC, Tan Q, Kruse TA, Mollenhauer J, Baumbach J, et al. Classification of breast cancer subtypes by combining gene expression and DNA methylation data. J Integr Bioinformatics. 2014;11:236.
van der Meide WF, Snellenberg S, Meijer CJ, Baalbergen A, Helmerhorst TJ, van der Sluis WB, et al. Promoter methylation analysis of WNT/beta-catenin signaling pathway regulators to detect adenocarcinoma or its precursor lesion of the cervix. Gynecol Oncol. 2011;123:116–22.
Li B, Goyal J, Dhar S, Dimri G, Evron E, Sukumar S, et al. CpG methylation as a basis for breast tumor-specific loss of NES1/kallikrein 10 expression. Cancer Res. 2001;61:8014–21.
Moura Lima E, Ferreira Leal M, Cardoso Smith Mde A, Rodriguez Burbano R, Pimentel de Assumpcao P, Bello MJ, et al. DNA mismatch repair gene methylation in gastric cancer in individuals from northern Brazil. Biocell. 2008;32:237–43.
Cooper SJ, Zou H, Legrand SN, Marlow LA, von Roemeling CA, Radisky DC, et al. Loss of type III transforming growth factor-beta receptor expression is due to methylation silencing of the transcription factor GATA3 in renal cell carcinoma. Oncogene. 2010;29:2905–15.
Hattermann K, Mehdorn HM, Mentlein R, Schultka S, Held-Feindt J. A methylation-specific and SYBR-green-based quantitative polymerase chain reaction technique for O6-methylguanine DNA methyltransferase promoter methylation analysis. Anal Biochem. 2008;377:62–71.
Miao F, Chen Z, Genuth S, Paterson A, Zhang L, Wu X, et al. Evaluating the role of epigenetic histone modifications in the metabolic memory of type 1 diabetes. Diabetes. 2014;63:1748–62.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45:228–47.
Boser BGI, Vapnik V. A training algorithm for optimal margin classifiers. ACM Digital Library. 1992:144–52.
Hoerl AE, Kennard RW. Ridge regression: biased estimation for nonorthogonal problems. Technometrics. 1970;12:55–67.
Breiman L. Random forests. Mach Learn. 2001;45:5–32.
Breiman LFH, Olshen RA, Stone CJ. Classification and regression trees. Wadsworth International Group; Belmont, California, 1984.
Acknowledgements
We would like to thank Dr. Vassilis Vassilakakis and Dr. Maria Markaki for preliminary data processing.
Funding
Ms Maria Panagopoulou received a scholarship for the implementation of her PhD Thesis, co-funded through the Act: “PROGRAM FOR THE GRANTING OF SCHOLARSHIPS FOR POSTGRADUATE STUDIES OF SECOND CYCLE STUDIES”. State Scholarship Foundation in Greece (IKY) (Operational Program “Human Resources Development—Education and Lifelong Learning”, Partnership Agreement PA 2014-2020).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The study was approved by the Scientific Board of the University General Hospital of Evros (PGNE), following assessment by Ethics Committee (decision 14/895/28.11.11), and was conducted according to the ethical principles of the 1964 Declaration of Helsinki and its later amendments.
Informed consent
All patients participated after signing a voluntary informed consent.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Panagopoulou, M., Karaglani, M., Balgkouranidou, I. et al. Circulating cell-free DNA in breast cancer: size profiling, levels, and methylation patterns lead to prognostic and predictive classifiers. Oncogene 38, 3387–3401 (2019). https://doi.org/10.1038/s41388-018-0660-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0660-y
This article is cited by
-
Extensive review on breast cancer its etiology, progression, prognostic markers, and treatment
Medical Oncology (2023)
-
The double agents in liquid biopsy: promoter and informant biomarkers of early metastases in breast cancer
Molecular Cancer (2022)
-
Just Add Data: automated predictive modeling for knowledge discovery and feature selection
npj Precision Oncology (2022)
-
Functional regulations between genetic alteration-driven genes and drug target genes acting as prognostic biomarkers in breast cancer
Scientific Reports (2022)
-
Synergy between the Levels of Methylation of microRNA Gene Sets in Primary Tumors and Metastases of Ovarian Cancer Patients
Bulletin of Experimental Biology and Medicine (2022)