Abstract
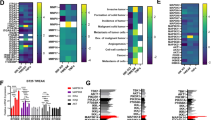
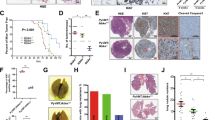
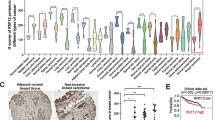
HER2/ErbB2 activation turns on transcriptional processes that induce local invasion and lead to systemic metastasis. The early transcriptional changes needed for ErbB2-induced invasion are poorly understood. Here, we link ErbB2 activation to invasion via ErbB2-induced, SUMO-directed phosphorylation of a single serine residue, S27, of the transcription factor myeloid zinc finger-1 (MZF1). Utilizing an antibody against MZF1-pS27, we show that the phosphorylation of S27 correlates significantly (p < 0.0001) with high-level expression of ErbB2 in primary invasive breast tumors. Phosphorylation of MZF1-S27 is an early response to ErbB2 activation and results in increased transcriptional activity of MZF1. It is needed for the ErbB2-induced expression of MZF1 target genes CTSB and PRKCA, and invasion of single-cells from ErbB2-expressing breast cancer spheroids. The phosphorylation of MZF1-S27 is preceded by poly-SUMOylation of K23, which can make S27 accessible to efficient phosphorylation by PAK4. Based on our results, we suggest for an activation mechanism where phosphorylation of MZF1-S27 triggers MZF1 dissociation from its transcriptional repressors such as the CCCTC-binding factor (CTCF). Our findings increase understanding of the regulation of invasive signaling in breast cancer by uncovering a detailed biological mechanism of how ErbB2 activation can rapidly lead to its invasion-promoting target gene expression and invasion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The raw data from the mass spectrometry analysis and TMA staining are available upon request. The scripts and the files of the modeling and molecular dynamics simulation are available as a Github repository at https://github.com/ELELAB/MZF1_SUMO.
References
Arteaga CL, Engelman JA. ERBB receptors: from oncogene discovery to basic science to mechanism-based cancer therapeutics. Cancer Cell. 2014;25:282–303.
Slamon DJ, Clark GM, Wong SG, Levin WJ, Ullrich A, McGuire WL. Human breast cancer: correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science. 1987;235:177–182.
Venur VA and Leone JP. Targeted Therapies for Brain Metastases from Breast Cancer. Int J Mol Sci, 2016;17–27.
Duchnowska R, Sperinde J, Chenna A, Huang W, Weidler JM, Winslow J. et al. Quantitative HER2 and p95HER2 levels in primary breast cancers and matched brain metastases. Neuro Oncol. 2015;17:1241–9.
Dittmer J. Mechanisms governing metastatic dormancy in breast cancer. Semin Cancer Biol. 2017;44:72–82.
Segatto O, King CR, Pierce JH, Di Fiore PP, Aaronson SA. Different structural alterations upregulate in vitro tyrosine kinase activity and transforming potency of the erbB-2 gene. Mol Cell Biol. 1988;8:5570–4.
Saez R, Molina MA, Ramsey EE, Rojo F, Keenan EJ, Albanell J. et al. p95HER-2 predicts worse outcome inpatients with HER-2-positive breast cancer. Clin Cancer Res. 2006;12:424–31.
Rafn B, Nielsen CF, Andersen SH, Szyniarowski P, Corcelle-Termeau E, Valo E. ErbB2-driven breast cancer cell invasion depends on a complex signaling network activating myeloid zinc finger-1-dependent cathepsin B expression. Mol Cell. 2012;45:764–76.
Scaltriti M, Rojo F, Ocana A, Anido J, Guzman M, Cortes J. et al. Expression of p95HER2, a truncated form of the HER2 receptor, and response to anti-HER2 therapies in breast cancer. J Natl Cancer Inst. 2007;99:628–38.
Tural D, Akar E, Mutlu H, Kilickap S. P95 HER2 fragments and breast cancer outcome. Expert Rev Anticancer Ther. 2014;14:1089–96.
Brix DM, Rafn B, Bundgaard Clemmensen K, Andersen SH, Ambartsumian N, Jaattela M. et al. Screening and identification of small molecule inhibitors of ErbB2-induced invasion. Mol Oncol. 2014;8:1703–18.
Hamalisto S, Jaattela M. Lysosomes in cancer-living on the edge (of the cell). Curr Opin Cell Biol. 2016;39:69–76.
Sevenich L, Joyce JA. Pericellular proteolysis in cancer. Genes Dev. 2014;28:2331–47.
Kallunki T, Olsen OD, Jaattela M. Cancer-associated lysosomal changes: friends or foes?. Oncogene. 2013;32:1995–2004.
Fonovic M, Turk B. Cysteine cathepsins and extracellular matrix degradation. Biochim Biophys Acta. 2014;1840:2560–70.
Mason SD, Joyce JA. Proteolytic networks in cancer. Trends Cell Biol. 2011;21:228–37.
Mudduluru G, Vajkoczy P, Allgayer H. Myeloid zinc finger 1 induces migration, invasion, and in vivo metastasis through Axl gene expression in solid cancer. Mol Cancer Res. 2010;8:159–69.
Yue CH, Chiu YW, Tung JN, Tzang BS, Shiu JJ, Huang WH. et al. Expression of protein kinase C alpha and the MZF-1 and Elk-1 transcription factors in human breast cancer cells. Chin J Physiol. 2012;55:31–6.
Tsai LH, Wu JY, Cheng YW, Chen CY, Sheu GT, Wu TC. et al. The MZF1/c-MYC axis mediates lung adenocarcinoma progression caused by wild-type lkb1 loss. Oncogene. 2015;34:1641–9.
Deng Y, Wang J, Wang G, Jin Y, Luo X, Xia X. et al. p55PIK transcriptionally activated by MZF1 promotes colorectal cancer cell proliferation. Biomed Res Int. 2013;2013:868131
Eguchi T, Prince T, Wegiel B, Calderwood SK. Role and Regulation of Myeloid Zinc Finger Protein 1 in Cancer. J Cell Biochem. 2015;116:2146–2154.
Weber CE, Kothari AN, Wai PY, Li NY, Driver J, Zapf MA. et al. Osteopontin mediates an MZF1-TGF-beta1-dependent transformation of mesenchymal stem cells into cancer-associated fibroblasts in breast cancer. Oncogene. 2015;34:4821–4833.
Egeblad M, Mortensen OH, Jaattela M. Truncated ErbB2 receptor enhances ErbB1 signaling and induces reversible, ERK-independent loss of epithelial morphology. Int J Cancer. 2001;94:185–91.
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell. 2006;10:515–27.
Nagashima T, Shimodaira H, Ide K, Nakakuki T, Tani Y, Takahashi K. et al. Quantitative transcriptional control of ErbB receptor signaling undergoes graded to biphasic response for cell differentiation. J Biol Chem. 2007;282:4045–56.
Sander TL, Haas AL, Peterson MJ, Morris JF. Identification of a novel SCAN box-related protein that interacts with MZF1B. The leucine-rich SCAN box mediates hetero- and homoprotein associations. J Biol Chem. 2000;275:12857–67.
Ogawa H, Murayama A, Nagata S, Fukunaga R. Regulation of myeloid zinc finger protein 2A transactivation activity through phosphorylation by mitogen-activated protein kinases. J Biol Chem. 2003;278:2921–7.
Yue CH, Huang CY, Tsai JH, Hsu CW, Hsieh YH, Lin H. et al. MZF-1/Elk-1 Complex Binds to Protein Kinase Calpha Promoter and Is Involved in Hepatocellular Carcinoma. PLoS One. 2015;10:e0127420
Tvingsholm SA, Hansen MB, Clemmensen KKB, Brix DM, Rafn B, Frankel LB. et al. Let-7 microRNA controls invasion-promoting lysosomal changes via the oncogenic transcription factor myeloid zinc finger-1. Oncogenesis. 2018;7:14
Slamon DJ, Godolphin W, Jones LA, Holt JA, Wong SG, Keith DE. et al. Studies of the HER-2/neu protooncogenein human breast and ovarian cancer. Science. 1989;244:707–12.
Baker AF, Dragovich T, Ihle NT, Williams R, Fenoglio-Preiser C, Powis G. Stability of phosphoprotein as a biological marker of tumor signaling. Clin Cancer Res. 2005;11:4338–40.
Bawa-Khalfe T, Yeh ET. SUMO losing balance: SUMO proteases disrupt SUMO homeostasis to facilitate cancer development and progression. Genes Cancer. 2010;1:748–52.
Kim KI, Baek SH. SUMOylation code in cancer development and metastasis. Mol Cells. 2006;22:247–53.
Hietakangas V, Anckar J, Blomster HA, Fujimoto M, Palvimo JJ, Nakai A. et al. PDSM, a motif for phosphorylation-dependent SUMO modification. Proc Natl Acad Sci USA. 2006;103:45–50.
Varjosalo M, Sacco R, Stukalov A, van Drogen A, Planyavsky M, Hauri S. et al. Interlaboratory reproducibility of large-scale human protein-complex analysis by standardized AP-MS. Nat Methods. 2013;10:307–14.
Mathelier A, Zhao X, Zhang AW, Parcy F, Worsley-Hunt R, Arenillas DJ. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 2014;42:D142–7.
Marshall AD, Bailey CG, Rasko JE. CTCF and BORIS in genome regulation and cancer. Curr Opin Genet Dev. 2014;24:8–15.
Seeler JS, Dejean A. SUMO and the robustness of cancer. Nat Rev Cancer. 2017;17:184–97.
Noll L, Peterson FC, Hayes PL, Volkman BF, Sander T. Heterodimer formation of the myeloid zinc finger 1 SCAN domain and association with promyelocytic leukemia nuclear bodies. Leuk Res. 2008;32:1582–92.
Xiao Y, Pollack D, Nieves E, Winchell A, Callaway M, Vigodner M. Can your protein be sumoylated? A quick summary and important tips to study SUMO-modified proteins. Anal Biochem. 2015;477:95–7.
Subramonian D, Raghunayakula S, Olsen JV, Beningo KA, Paschen W, Zhang XD. Analysis of changes in SUMO-2/3 modification during breast cancer progression and metastasis. J Proteome Res. 2014;13:3905–18.
Minden A. The pak4 protein kinase in breast cancer. ISRN Oncol. 2012;2012:694201.
Abdel-Magid AF. P21-activated kinase 4 (PAK4) inhibitors as potential cancer therapy. ACS Med Chem Lett. 2015;6:17–8.
Ko H, Kim S, Yang K, Kim K. Phosphorylation-dependent stabilization of MZF1 upregulates N-cadherin expression during protein kinase CK2-mediated epithelial-mesenchymal transition. Oncogenesis. 2018;7:27
Ha BH, Morse EM, Turk BE, Boggon TJ. Signaling, regulation, and specificity of the type II p21-activated kinases. J Biol Chem. 2015;290:12975–83.
Hendriks IA, Lyon D, Young C, Jensen LJ, Vertegaal AC, Nielsen ML. Site-specific mapping of the human SUMO proteome reveals co-modification with phosphorylation. Nat Struct Mol Biol. 2017;24:325–36.
Tan M, Li P, Sun M, Yin G, Yu D. Upregulation and activation of PKC alpha by ErbB2 through Src promotes breast cancer cell invasion that can be blocked by combined treatment with PKC alpha and Src inhibitors. Oncogene. 2006;25:3286–95.
Bailey TA, Luan H, Tom E, Bielecki TA, Mohapatra B, Ahmad G. et al. A kinase inhibitor screen reveals protein kinase C-dependent endocytic recycling of ErbB2 in breast cancer cells. J Biol Chem. 2014;289:30443–58.
Mertins P, Mani DR, Ruggles KV, Gillette MA, Clauser KR, Wang P. et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature. 2016;534:55–62.
Kim S, Yu NK, Kaang BK. CTCF as a multifunctional protein in genome regulation and gene expression. Exp Mol Med. 2015;47:e166.
Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:e45.
Ivascu A, Kubbies M. Rapid generation of single-tumor spheroids for high-throughput cell function and toxicity analysis. J Biomol Screen. 2006;11:922–32.
Acknowledgements
We acknowledge the excellent technical assistance of Anni Hadesten and Louise Vandervox and give special thanks to Professor Audrey Minden for PAK4, Professor Marin Barisic for the ΔT-3xHA-DEST CW plasmids, and Dr. Vincent Collins for help with the language editing.
Funding
Funding was provided by the Novo Nordisk Foundation (NNF15OC0017324) (TK), the Danish Medical Research Council (0602-02386B) (TK), the Danish Cancer Society Scientific Committee (KBVU) (R124-A7854-15-S2 and R56-A3108-12-S2) (TK), the Danish National Research Foundation (DNRF125) (MJ), the European Research Council (AdG 340751) (MJ), EU-PRACE DECI13th grant for computational resources (EP), and the DeiC Pilot Grant 2015-2016 for the Danish Supercomputing Center Computerome (EP).
Author information
Authors and Affiliations
Contributions
DMB, SAT, MBH, KBC, TO, VS, ML, PP, EP, and JM performed the experiments. KH, IG, PJ, and MV provided materials and/or facilities, TK planned the project, and wrote the manuscript, and TK, DMB, and MJ edited it.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Brix, D.M., Tvingsholm, S.A., Hansen, M.B. et al. Release of transcriptional repression via ErbB2-induced, SUMO-directed phosphorylation of myeloid zinc finger-1 serine 27 activates lysosome redistribution and invasion. Oncogene 38, 3170–3184 (2019). https://doi.org/10.1038/s41388-018-0653-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0653-x
This article is cited by
-
Activation of invasion by oncogenic reprogramming of cholesterol metabolism via increased NPC1 expression and macropinocytosis
Oncogene (2023)
-
MZF1 mediates oncogene-induced senescence by promoting the transcription of p16INK4A
Oncogene (2022)
-
P2x4 receptor promotes mammary cancer progression by sustaining autophagy and associated mesenchymal transition
Oncogene (2022)
-
Identification of lysosome‐targeting drugs with anti‐inflammatory activity as potential invasion inhibitors of treatment resistant HER2 positive cancers
Cellular Oncology (2021)
-
Coordinated dysregulation of cancer progression by the HER family and p21-activated kinases
Cancer and Metastasis Reviews (2020)