Abstract
Hepatocellular carcinoma (HCC), the major form of liver cancer, has shown increasing incidence and poor prognosis. Adipose tissue is known to function in energy storage and metabolism regulation by the secretion of adipokines. Circular RNAs (circRNAs), a novel type of noncoding RNA, have recently been recognized as key factors in tumor development, but the role of exosome circRNAs derived from adipose tissues has not been defined yet. Here, adipose-secreted circRNAs were found to regulate deubiquitination in HCC, thus facilitating cell growth. It was observed that exosome circ-deubiquitination (circ-DB) is upregulated in HCC patients with higher body fat ratios. Moreover, in vitro and in vivo studies showed that exo-circ-DB promotes HCC growth and reduces DNA damage via the suppression of miR-34a and the activation of deubiquitination-related USP7. Finally, the results showed that the effects of adipose exosomes on HCC cells can be reversed by knockdown of circ-DB. These results indicate that exosome circRNAs secreted from adipocytes promote tumor growth and reduce DNA damage by suppressing miR-34a and activating the USP7/Cyclin A2 signaling pathway.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC), one of the most common pathological types of primary liver tumor, is ranked as the second leading cause of death associated with cancer due to its poor 5-year-survival rate [1]. Chronic hepatitis B virus (HBV) or hepatitis C virus (HCV) infection, excessive drinking, immune-related hepatitis, and obesity are generally regarded as major risk factors contributing to HCC [2, 3]. Obesity is highly prevalent worldwide and is regarded as one of the largest contributors to the occurrence of nonalcoholic fatty liver disease (NAFLD) [4, 5]. However, the relationship between obesity and HCC is still poorly understood.
Exosomes are a type of microvesicle (MV) with a diameter of 40–100 nm and are secreted by the majority of cell types [6]. An increasing number of studies have shown that exosomes originate from multivesicular endosomes (multivesicular bodies) by inverse budding to form multivesicular bodies and are released into the extracellular space when the multivesicular body fuses with cell membrane exosomes [7]. Exosomes act as message transmitters in intercellular communication because they contain a variety of proteins, lipids, circRNAs, and microRNAs [8, 9]. CircRNAs are classified as new endogenous noncoding RNAs, different from common linear RNAs, and are characterized by covalently closed loops without 5′ or 3′ polarities. Recent studies have demonstrated that circRNAs can absorb microRNAs by stable complementary binding [10, 11] and serve as microRNA sponges to regulate gene expression. Although a growing number of studies have proven that circRNAs are abundant and stable in exosomes [12, 13], the functions of exosome circRNAs are not clear. In HCC cells, miR-34a plays a crucial role in the regulation of β-catenin[14]. However, the interaction between miR-34a and circRNA remains unknown.
Ubiquitin-specific protease 7 (USP7) is a deubiquitinating enzyme that plays a crucial role in various cellular processes, including the cell cycle, proliferation, and DNA repair [15, 16]. A high level of USP7 is frequently found in HCC tissues, which is correlated with tumor growth and invasion, and leads to poor overall survival [17, 18]. The USP7 inhibitor P5091 induces chronic lymphocytic leukemia (CLL) growth arrest and apoptosis [19]. In the process of hepatocellular tumorigenesis, ubiquitin-specific protease drives cell cycle progression [20]. Recent studies have demonstrated that Cyclin A2 is an important downstream substrate of USP7 and that the active USP7/Cyclin A2 signaling pathway can promote the growth of breast cancer [21].
In this study, we first compared the expression patterns of exosome circRNAs between HCC patients with higher and lower body fat ratios (BFRs) and found that exosome circ-BD (circular RNA related to deubiquitylation, has_circ_0025129) as well as USP7 are positively related to the BFR. Moreover, circ-DB was highly expressed in adipocyte exosomes and promoted USP7 and Cyclin A2 expression by absorbing miR-34a. We also detected that adipose exosomes promoted HCC proliferation and decreased DNA damage. The data from in vivo studies illustrate that exosomes from adipose tissues obviously reduced the level of miR-34a and activated the USP7/Cyclin A2 signaling pathway. The effects described above were abolished by the knockdown of circ-DB in HCC cells. By measuring the tumor weight, mouse weight, body fat ratio, and liver metastases, we showed that circ-DB acted as a booster to increase tumor progression. Thus, adipose-derived exosomes mediate the delivery of circRNAs and promote the tumorigenesis of HCC by regulating the deubiquitination-related miR-34a/USP7 axis.
Materials and methods
Human tissues
All human HCC tissue samples were obtained from Tianjin Medical University Cancer Institute and Hospital. Informed consent was provided by all patients, and the Ethics Committee of Tianjin Medical University Cancer Institute and Hospital approved all aspects of this study.
Cell culture
HepG2 (human hepatocellular adenocarcinoma), Hepa 1-6 (mouse hepatocellular adenocarcinoma), and 3T3L1 cell lines were purchased from the cell bank of the Chinese Academy of Sciences (Shanghai, China). HepG2 and Hepa 1-6 cells were maintained in DMEM (Gibco, USA) supplemented with 10% FBS; 3T3L1 cells were maintained in DMEM/F12 at 37 °C in a humidified atmosphere of 5% CO2. To induct 3T3L1 cells, indomethacin (125 mM; Sigma), dexamethasone (5 mM; Sigma), insulin (0.5 mg/ml), isobutylmethylxanthine (0.5 mM; Sigma), rosiglitazone (1 mM; Sigma), and T3 (1 nM; Sigma) were added to DMEM/F12.
Isolation of exosomes from medium and plasma
Exosomes in medium, as well as in plasma, were isolated by gradient centrifugation according to recent reports [22]. Following initial centrifugation for 30 min at 3000 × g, cells and other debris were removed, and then, the supernatant was harvested and centrifuged at 10,000 × g for 30 min to remove MVs that were larger than exosomes. Finally, the supernatant was centrifuged at 110,000 × g for 70 min (the isolation process was performed at 4 °C), and exosomes were resuspended in PBS stored at -80 °C.
Transmission electron microscopy assay
The exosomes were fixed overnight in 2.5% glutaraldehyde at pH 7.2 at 4 °C. The prepared samples were washed in PBS 3 times (5 min, each time) and fixed in 1% osmium tetroxide for 60 min at room temperature. Then, the sample blocks were prepared as follows: the samples were embedded in 10% gelatin, fixed in glutaraldehyde at 4 °C and cut into several blocks (less than 1 mm). Dehydration of the sample blocks was performed for 10 min at each step in increasing concentrations of alcohol (30%, 50%, 70%, 90%, 95%, and 100% × 3, 10 min, each step). Using propylene oxide to exchange pure alcohol, specimens were infiltrated with increasing concentrations (25%, 50%, 75%, and 100%) of Quetol-812 epoxy resin mixed with propylene oxide, with at least 3 h per step. Samples were embedded in pure, fresh Quetol-812 epoxy resin and polymerized at 35 °C for 12 h, 45 °C for 12 h, and 60 °C for 24 h. Ultrathin sections (100 nm) were cut using a Leica UC6 ultramicrotome and then stained with uranyl acetate for 10 min and with lead citrate for 5 min at room temperature before observation in an FEI Tecnai T20 transmission electron microscope.
Nanoparticle tracking analysis
The numbers and sizes of exosomes were directly measured using the Nanosight NS 300 system (NanoSight Technology, Malvern, UK) [23, 24]. Exosomes were resuspended in PBS at a concentration of 5 μg/mL and further diluted 100- to 500-fold to achieve 20–100 objects per frame. Samples were manually injected into the sample chamber at ambient temperature. Each sample was configured with a 488 nm laser and a high-sensitivity sCMOS camera and was monitored in triplicate at a camera setting of 13 with an acquisition time of 30 s and a detection threshold setting of 7. At least 200 completed tracks were analyzed per video. Finally, the data were analyzed using NTA analytical software (version 2.3).
Agilent methods for ceRNA array
Total RNA was isolated using TRIzol reagent (Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s protocol and purified by using an RNeasy Mini Kit (Qiagen, GmBH, Germany). RNA samples of each group were then used to generate fluorescence-labeled cRNA targets for the SBC human ceRNA array V1.0 (4 × 180 K). The labeled cRNA targets were then hybridized with the slides. After hybridization, the slides were scanned on an Agilent Microarray Scanner (Agilent Technologies, Santa Clara, CA, USA). The data were extracted with Feature Extraction software 10.7 (Agilent Technologies, Santa Clara, CA, USA). The raw data were normalized by the Quantile algorithm, limma package in the R program. The microarray experiments were performed by following the protocol of Agilent Technologies Inc. at Shanghai Biotechnology Corporation. The ratios were calculated for the above-the-mean body fat ratio and below-the-mean body fat ratio groups in HCC. Genes with a fold change of at least 2 were selected for further analysis. The selected parent genes of circRNAs were grouped in functional categories based on the Gene Ontology database (GO: http://www.geneontology.org/), and the functional pathways (KEGG) were also analyzed by using online gene enrichment analysis system of Shanghai Biotechnology Corporation.
RNA isolation and quantitative RT-PCR
Quantification of miRNAs, circRNAs, and mRNAs was performed as described, with slight modifications [25, 26]. Total RNA was extracted from the cultured cells and tissues using TRIzol Reagent (Invitrogen) according to the manufacturer’s protocol. Reverse transcription and qPCR were performed on an Applied Biosystems AB7500 Real-Time PCR system. miRNA determination was performed using TaqMan microRNA probes (Applied Biosystems, Foster City, CA). All PCRs were run in triplicate, and the relative amount of gene normalized to control was calculated with the equation 2-ΔCT, in which ΔCT = CT gene − CT control. A comparative CT method was used to compare each condition to the control reactions. U6 snRNA was used as an internal control for miRNAs, and the circRNA and mRNA levels were normalized to GAPDH.
The primers of circ-DB were as follows:
5′-CTGCTCCTCCAGCTCTT-3′(circ-DB sense);
5′-AGTGATCTTGAACCCCAAAG-3′(circ-DB anti-sense).
Knockdown of circRNAs in exosomes
3T3L1 cells were transfected with siRNAs of circ-DB, and exosomes were isolated from medium at 48 h. The sequence of circ-DB siRNA was as follows: TGCTTTAATTTTCTCAGAATTCT.
Immuno-precipitation
Immuno-precipitation using anti-cyclin A2 antibody was performed at 48 h or 72 h after treatment. Cells were lysated by the lysis buffer containing 150 mM KCl, 25 mM Tris-HCl, pH7.4, 5 mM EDTA, 0.5% Triton X-100, 5 mM dithiothreitol (DTT), PMSF and cocktail. The supernatant was mixed with cyclin A2 antibody (Abcam) overnight at 4 °C, and then co-cultured with beads (santa cruz) for 2–4 h at RT. The beads were washed five times in lysis buffer followed by western blotting (WB) analysis.
Biotin-coupled miRNA capture
Biotin-labeled miRNAs were transfected into HEK293T cells at a final concentration of 20 nM. Cells were collected at 48 h following transfection and treated as described above.
miRNA target prediction
miRNA target prediction and analysis were performed with the algorithms from TargetScan (http://www.targetscan.org/), PicTar (http://pictar.mdc-berlin.de/), and miRanda (http://www.microrna.org/).
Western blotting
USP7 and Cyclin A2 expression was assessed by western blot analysis, and the samples were normalized to GAPDH. Total protein was prepared in RIPA buffer. Each sample was separated by 10% SDS-PAGE and transferred onto PVDF membranes (EMD Millipore, Billerica, MA, USA). Following blocking for 1 h, the membranes were incubated with primary antibodies, such as anti-USP7 (1:1000, Santa Cruz, sc-133204), anti-Cyclin A2 (1:1000, Abcam, ab181591), anti-CD63 (1:1000, Santa Cruz, sc-5275), anti-TSG101 (1:1000, Santa Cruz, sc-136111), anti-Alix (1: 1000, Santa Cruz, sc-53540), and anti-GAPDH (1:3000, Santa Cruz, sc-32233), overnight at 4 °C and then incubated with the corresponding secondary antibodies at 1:2000 dilution for 2 h at room temperature.
Cell transfection
Cells were seeded in a specific plate and transfection was conducted after 24 h. Cell transfection with miR-34a mimics and inhibitors were conducted by using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions. miRNA mimics are double-strand RNAs that can form mature miRNAs in cells, while miRNA inhibitors are single-strand RNAs that interact with miRNAs and lead to the downregulation of miRNAs. For each well, equal doses (100 pmol) of miRNA mimics, inhibitors, siRNAs, or scrambled negative control RNA were used. The cells were harvested at 24 h after transfection for real-time quantitative PCR analysis and WB.
Oil red O staining
The cells were removed, washed twice with PBS, fixed with 4% formaldehyde at room temperature, and washed three times with PBS. To stain the lipids in adipocytes, the cells were treated with filtered Oil Red O solution for 1 h at room temperature and then washed twice with PBS. The resulting red-stained lipid droplets were observed under a microscope(21).
Luciferase assay
The reporter plasmid p-MIR-USP7 expressing the predicted miR-133 target-binding sites was produced by Genescript (Nanjing, China). Parts of the wild type and mutated 3′-UTR of USP7 were cloned into the firefly luciferase reporter. Similarly, the reporter plasmids p-MIR-circ-DB containing the miR-34a binding regions 1 or 2 were also produced by Genescript (Nanjing, China), and each plasmid expressed one single miR-34a binding region. Parts of the wild type and mutated 3′-UTR of miR-34a containing binding region 1 or 2 were cloned into the firefly luciferase reporter. Two milligrams of beta-galactosidase expression vector (Ambion) was chosen as a transfection control. For the subsequent luciferase reporter assays, 2 mg of firefly luciferase reporter plasmid with 2 mg of beta-galactosidase vector or equal doses (200 pmol) of mimics, inhibitors, and scrambled negative control RNA were transfected into the prepared cells. At 24 h after transfection, levels of luciferase were analyzed with a Dual Luciferase Assay Kit (Promega) according to the manufacturer’s instructions.
Immunohistochemistry
The tumors were fixed in 4% paraformaldehyde, embedded in paraffin, sectioned, and then stained with anti-USP7 antibodies (Abcam). Quantitative analysis was conducted by quantifying the fluorescence intensity from at least five sections.
Cell proliferation assay
Cells were incubated with 50 μm EdU (Ribo-Bio, Guangzhou, China) for 12 h and then fixed with 4% paraformaldehyde for 30 min at 25 °C. Next, the cells were washed in PBS (2 × 5 min, room temperature [RT]) and then permeabilized using PBS containing 0.3% Triton X-100 for 10 min. After extensive washing in PBS, the cells were incubated in Apollo staining solution (RiboBio) for 20 min, washed with NaCl/Pi (3 × 10 min, RT), and then incubated in Hoechst (1:2500; Roche Diagnostics, Mannheim, Germany) for 10 min at RT.
Immunofluorescence
Cells were cultured in 12-well plates with slides. At harvest, the cells were first fixed with 4% paraformaldehyde and then permeabilized with 0.2% Triton X-100 for 10 min, followed by confinement for 1 h. Then, the cells were incubated with Phospho-H2AFX-S139 antibody (1:100; ABclonal, AP0099) at 4 °C overnight and Alexa Fluor 488 goat anti-rabbit IgG (Invitrogen) for 1 h in the dark at RT. DAPI (Solarbio, China) was used for nuclear staining. For confocal microscopy, a Nikon C2 Plus confocal microscope was used.
Construction of circRNA overexpressed plasmid and lentivirus
The circ-DB overexpressing plasmid was constructed by GeneChem (Shanghai, China). Briefly, the GV485-circ-DB was constructed by PCR amplifying the circRNA locus by using circ-DB forward/reverse primers. The PCR fragment was inserted into KpnI/BamHI-digested GV485.
Establishment of tumors in mice
Male mice (C57 and ob/ob, 6–8 weeks old) were divided randomly into groups as described in Fig. 9a, and we were not blinded to the group allocation during the experiment. The lenti-viruses containing circ-DB overexpressing sequences or shRNA were purchased from GeneChem (Shanghai, China). Lenti-viruses were transfected into the Hepa 1-6 cells according to our protocol as shown in Fig. 8. The prepared Hepa 1-6 cells were injected subcutaneously into each C57 and ob/ob mouse (1 × 107 cells per mouse). The mice were sacrificed, and tumors were removed at day 35. All mice were kept in pathogen-free cages and killed 4 weeks after inoculation. All experimental procedures were approved by the Institutional Animal Care and Research Advisory Committee of Tianjin Medical University Cancer Institute and Hospital.
Statistics
All results were obtained from at least three separate experiments and are presented the mean ± standard deviation. Differences between samples are presented as the mean value ± standard deviation. The data were statistically analyzed using Student’s t-test in SPSS statistical software, with p < 0.05 considered statistically significant. * indicates p < 0.05, ** indicates p < 0.01, and *** indicates p < 0.001.
Results
The expression patterns of exo-circ-DB and USP7 in HCC
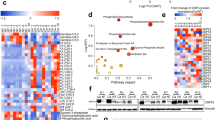
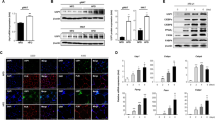
Exosomes were isolated from the plasma of both HCC patients and normal subjects (NC) as described, and transmission electron microscopy (TEM) showed that plasma exosomes exhibited a round-shaped morphology and a size of 50–100 nm. Moreover, the presence of the exosome markers CD63, Alix and tsg101 were confirmed by western blot (Fig. 1a-c). Forty HCC patients were divided into two groups based on the median body fat ratios, namely, the above-the-median group (n = 21) and below-the-median group (n = 19) (Fig. 1d). A panel of exosome circRNAs in plasma was dys-regulated, and circ-DB (hsa_circ_0025129|NM_001065|TNFRSF1A) in the above-the-median group showed a 3-fold increase compared with that in the below-the-median group (Fig. 1e). As expected, the expression level of USP7 was also clearly higher in the above-the-median group than in the below-the-median group (Fig. 1f, g). Levels of circ-DB in exosomes were determined using RT-qPCR, and USP7 protein was quantified by combination of WB and gray analysis. As is shown in Fig. 1j, circ-DB showed a positive correlation with the amounts of USP7 protein in HCC patients. We proposed the hypothesis that adipocyte exosomes carrying circ-DB were taken up by HCC cells, which reduced the level of miR-34a. As a result, the downregulation of miR-34a led to the overexpression of USP7 (Fig. 1h, i). These data suggested that the exosome circ-DB is upregulated in HCC patients with higher body fat ratio, and that circ-DB is positively related with USP7 expression.
The correlation between exo-circ-DB and USP7 in HCC. a TEM image of exosomes isolated from human plasma (scale bar, 100 nm). b Plasma exosomes were isolated from both normal subjects (NC) and HCC patients. Western blotting of three representative exosome-specific markers: CD63, Alix, and Tsg101. c The size range of the plasma exosomes checked by NAT analysis. d HCC patients were divided into the above-the-median group (n = 21) and below-the-median group (n = 19) according to the median body fat ratio of the HCC patients. e Preliminary circ-RNA screening of HCC by an Agilent Microarray Scanner (n = 21 and n = 19, respectively). f, g WB analysis of USP7 in the above-the-median group and below-the-median group (n = 21 and n = 19, respectively). h Representative images of the potential interaction between circ-DB and USP7. I. The predicting binding region of miR-34a in USP7 mRNA. J. The clinical relevance between plasma exo-circ-DB and USP7 protein (n = 21). **p < 0.01
The clinical relevance of exo-circ-DB, miR-34a, and USP7
We subsequently checked USP7 expression in para-carcinoma tissues (P) and tumor tissues (T) of both groups via immunohistochemistry (IHC). It was shown that USP7 was significantly higher in the above-the-median group than in the below-the-median group (Fig. 2a). In addition, the higher expression of USP7 coincided with a lower probability of survival (Fig. 2b). Exosomes contain various types of circ-RNA that play biological roles when taken up by target cells. RT-PCR indicated that the expression level of circ-DB carried by plasma exosomes (exo-circ-DB) was increased 10-fold in the above-the-median group compared with that in the below-the-median group (Fig. 2c). In HCC tumors, miR-34a showed decreased expression in the above-the-median group (Fig. 2d), while the expression of USP7 mRNA was higher in the below-the-median group (Fig. 2e). In addition, exo-circ-DB was negatively linked with the amount of miR-34a, which was negatively linked with USP7 (Fig. 2f, g). These data indicated that plasma exosome circ-DB is upregulated in the above-the-median group and is closely related to the expression of miR-34a and USP7 in HCC.
Clinical relevance among exo-circ-DB, miR-34a and USP7 in HCC. a The expression of USP7 in para-carcinoma tissues (P) and tumor tissues (T) was detected by immunohistochemistry (IHC). Scale bar, 100 μm. b The HCC patients were divided into two groups according to the mean value of USP7 mRNA. The expression of USP7 was closely linked with the survival in HCC (n = 1595 in high group, and n = 2356 in low group). c–e The levels of exo-circ-DB, miR34a of tumors and USP7 of tumors in the above-the-median group and below-the-median group (n = 21 and n = 19, respectively). f, g The negative relationship between exo-circ-DB and miR-34a (n = 21, R = 0.735), as well as between miR-34a and USP7 (n = 21, R = 0.762). **p < 0.01
The direct interaction between miR-34a and USP7
Recently, miR-34a was reported to be downregulated in some hematological tumors and was considered as a regulator of cancer-related genes linked to the cell cycle and to cell proliferation and survival [27, 28]. However, the functional contributions of miR-34a to HCC have not been experimentally established. To evaluate the possible effects of miR-34a on the growth of HC, USP7 was found to be among the predicted high confidence targets by all three methods. The 3′-UTR of human USP7 harbors the predicted binding site of has-miR-34a (Fig. 3a). miRNA mimics were used to overexpress miR-34a, while miRNA inhibitors were used to suppress miR-34a expression (Fig. 3b). Immuno-precipitation using biotin-labeled miR-34a was conducted to determine whether miR-34a directly interacts with USP7. The USP7 mRNA captured by biotin-miR-34a showed significant increase compared to the control group (Fig. 3c).Subsequently, the luciferase reporter assay was performed to further demonstrate that miR-34a regulates USP7 reporter. A luciferase reporter plasmid containing the wt miR-34a-binding region was constructed. The plasmids containing the reverse sequences of the miR-133-binding site (mut) was used as negative controls. The overexpression of miR-34a significantly suppressed luciferase activity, while in the mut group, this inhibition was almost abolished (Fig. 3d). The expression of both USP7 and Cyclin A2 significantly decreased after treatment with miR-34a. However, the expression of Cyclin A2 was suppressed with the knockdown of miR-34a (Fig. 3e–g). Human miRNAs negatively regulate gene expression at the post-transcriptional level, and do not lead to mRNA degradation; and USP7 stabilize cyclin A2 protein by inhibiting ubiquitination and has no effects on the levels of cyclin A2 mRNA. As is expected, the miR-34a mimics and inhibitors regulates USP7 and cyclin A2 expression without causing the variation of mRNA (Fig. 3h). In addition, the expression of Cyclin A2 was suppressed when USP7 was knocked down by siRNA (Fig. 3i, j). These results demonstrated that miR-34a directly suppresses USP7 and that Cyclin A2 is an important downstream substrate of USP7.
Validation of USP7 as a direct target of miR-34a in HCC cells as well as targeting of Cyclin A2. a The putative binding sites for miR-34a and USP7. b RT-PCR analysis of the level of miR-34a, which we constructed (n = 3). c RT-qPCR analysis of USP7 mRNA captured by biotin-miR-34a (n = 3). d Direct recognition of USP7 by miR-34a (n = 3). Firefly luciferase reporters containing either wild-type (WT) or mutant (Mut) USP7, miR-34a mimics and the associated normal control were co-transfected into 293 T cells. The relative luciferase levels were detected after transfection (n = 3). e–g WB revealed the expression levels of USP7 and cyclin A2 in HCC cells treated with miR-34a mimics or miR-34a inhibitors (n = 3). h Corresponding quantification of USP7 and cyclin A2 expression as demonstrated by RT-PCR (n = 3). i, j The expression level of Cyclin A2 when USP7 was overexpressed or inhibited by siRNA (n = 3). **p < 0.01
The effects of adipocyte exosomes on miR-34a, USP7, and Cyclin A2 in HCC cells
The exosomes secreted from 3T3L1 had a typical round morphology, a size of 100 nm, and high expression of CD63, Alix, and tsg10 (Fig. 4a, b). As is well known, adipocytes at different stages of differentiation show different biological properties. The oil red stain assay showed that lipid droplets existed in mature adipocytes but not in pre-adipocytes (Fig. 4c). Compared with pre-adipocytes, the relative levels of circ-DB were remarkably higher in mature adipocytes, and the same results were observed for exosomes from pre-adipocytes and mature adipocytes, respectively (Fig. 4d, e). The HCC cells were treated with exosomes secreted from mature adipocytes (Fig. 4f), and the levels of miR-34a, USP7 and cyclin A2 were detected. To remove circ-DB from exosomes, siRNA of circ-DB was transfected into 3T3L1 cells and exosomes were collected at 48 h post-transfection. The expression of miR-34a was suppressed but that expression of USP7 and cyclin A2 was enhanced; while these effects were blocked when circ-DB was removed from 3T3L1 exosomes (Fig. 4g–i). In summary, adipocyte exosomes acted as carriers of exo-circ-DB to upregulate USP7 and cyclin A2.
The expression of miR-34a, USP7, and Cyclin A2 is regulated by adipocyte exosomes. a Transmission electron microscopy image of 3T3L1 exosomes (scale bar, 100 nm). b CD63, Alix, and Tsg101 expression as detected by WB. c Oil red O staining of mature adipocytes and pre-adipocytes. d, e The difference in circ-DB expression in mature and pre-adipocytes (d), as well as in exosomes from 3T3L1 cells (e) (n = 3). f Graphical representation of HCC cells treated with mature adipocyte exosomes. g–i Quantification of miR-34a and the expression levels of USP7 and cyclin A2 in HCC cells treated with mature adipocyte exosomes (n = 3). **p < 0.01
The correlation between circ-DB and miR-34a
The sequences of the two binding regions for circ-DB and miR-34a are shown in Fig. 5a, b. Subsequently, we performed the luciferase reporter assay to further reveal the direct contact of circ-DB and miR-34a (Fig. 5c, d). A luciferase reporter plasmid containing miR-34a-binding region 1 or 2 was constructed; each plasmid contained one single miR-34a-binding region. The plasmids containing the reverse sequences of the miR-34a-binding sites (mut) were used as negative controls. Overexpressed miR-34a significantly suppressed luciferase activity, but this inhibitory effect was abolished in the mut group (Fig. 5c, d). The expression of miR-34a was enhanced when circ-DB was knocked down by siRNA (Fig. 5e, f). To further evaluate the effects of circ-DB on USP7 or Cyclin A2, we used WB to detect the expression levels of USP7 and Cyclin A2. The expression of both USP7 and Cyclin A2 was significantly enhanced by overexpressed circ-DB but was inhibited when circ-DB siRNA was used instead (Fig. 5g, h). USP7 mRNA showed little change under both conditions (Fig. 5i). These results demonstrated that circ-DB acts as a sponge of miR-34a.
circ-DB regulates the miR-34a/USP7/Cyclin A2 pathway. a, b Two of the predicted binding regions of miR-34a in circ-DB. c, d Verification of the interaction between miR-34a and circ-DB by using luciferase assay (n = 3). e RT-PCR quantification of circ-DB that we constructed or the inhibition effect of circ-DB siRNA. f The level of miR-34a when cells were treated with circ-DB-OE plasmid and circ-DB siRNA. g, h The expression level of USP7 and Cyclin A2 in mature adipocytes treated with circ-DB-OE plasmid and circ-DB (n = 3). i Quantification of USP7 in 3T3L1 cells treated with circ-DB-OE plasmid and circ-DB siRNA (f) (n = 3) **p < 0.01, and ***p < 0.001, respectively
Adipose exo-circ-DB promotes growth of and inhibits DNA damage in HCC cells
Subsequently, we checked the effects of adipose-derived exosomes on the biological behavior of HCC cells. It was clearly shown that exosomes of 3T3L1 cells promote the proliferation and migration of HCC cells, but this promotion was eliminated when circ-DB was knocked down in exosomes (Fig. 6a–d). As expected, adipose exosomes and circ-DB also reduced DNA damage in HepG2 cells (Fig. 6e, f).
Exosome circ-DB promotes tumor growth and reduces DNA damage. HCC cells were pretreated with 3T3 exo or 3T3 exo from which circ-DB was deleted by circ-DB siRNA. a, b Determination of cell proliferation by Edu assay (n = 3). c, d Cell migration was determined by transwell assay (n = 3). e, f Evaluation of DNA damage (n = 3). **p < 0.01
HepG2 cells were transfected with circ-DB overexpression plasmids and circ-DB siRNA to further clarify the biological role of circ-DB in HCC. It was shown that the proliferation of HCC cells was significantly enhanced by treatment with circ-DB-OE (Fig. 7a, b) or USP7-OE (Fig. 7e, f), but was clearly inhibited when treated with siRNA-circ-DB or siRNA-USP7. Furthermore, DNA damage in HCC cells was reduced by treatment with circ-DB-OE (Fig. 7c, d) or USP7-OE (Fig. 7g, h) and was relatively increased when the cells were treated with siRNA-circ-DB or siRNA-USP7.
Role of circ-DB and USP7 in regulating proliferation and DNA damage in HCC cells. HCC cells were pretreated with circ-DB overexpressing plasmid (OE.circ-DB) and circ-DB siRNA deleted by circ-DB siRNA (si.circ-DB). a, b The cells were stained with DAPI and EdU, and then, the nuclei were imaged to reveal the proliferation of HCC cells (n = 3). c, d The cells were stained with DAPI and H2AFX, and then, the nuclei were imaged (n = 3). HCC cells were pretreated with USP7-overexpressing plasmid (OE.USP7) and USP7 siRNA (si.USP7). e, f The cells were stained with DAPI and EdU, and then, the nuclei were imaged to reveal the proliferation of HCC cells (n = 3). g, h The cells were stained with DAPI and H2AFX, and then, the nuclei were imaged (n = 3). **p < 0.01
These data illustrated that adipose exosomes promote the proliferation and migration of and reduced DNA damage in HCC cells by delivering circ-DB.
USP7 mediates the deubiquitination of cyclin A2 in HCC
Cyclin A2 is a member of the cyclin family, promotes DNA synthesis during S phase and facilitates cell cycle transition from G2 to M phase [29]. Dysfunction of cyclin A2 is associated with erroneous cell proliferation and chromosomal instability [30]. Here, we examined the role of USP7 in mediating the deubiquitination of cyclin A2 by a combination of cyclin A2 immune-precipitation and UB WB. The expression of cyclin A2 was determined by loading total protein of HCC cells, and the ubiquitination of cyclin A2 was analyzed by using the immunoprecipitate pulled down by cyclin A2 antibody. As expected, exosomes of 3T3L1 cells, overexpressed USP7 and overexpressed circ-DB promoted cyclin A2 expression but significantly suppressed the ubiquitination of cyclin A2 (Fig. 8a–c). The inhibition of both USP7 and circ-DB increased the relative levels of ubiquitin-cyclin A2 (Fig. 8a–c). Meanwhile, miR-34 played an opposite role in regulating cyclin A2 ubiquitination (Fig. 8d). Therefore, USP7 acts as an oncogene by decreasing the ubiquitination of a number of proteins, including cyclin A2.
Exo-circ-DB and USP7 regulates Deubiquitination of cyclin A2. HCC cells were treated as described above, and immune-precipitation was used to analyze the level of ubiquitination. a WB analysis of Cyclin A2 ubiquitination in HCC cells treated with 3T3 exosomes (n = 3). b WB analysis of Cyclin A2 ubiquitination in HCC cells treated with USP7 plasmid or USP7 siRNA (n = 3). c WB analysis of Cyclin A2 ubiquitination in HCC cells treated with circ-DB plasmid or circ-DB siRNA (n = 3). d WB analysis of Cyclin A2 ubiquitination in HCC cells treated with miR-34a mimics or miR-34a inhibitors (n = 3). **p < 0.01
circ-DB enhances tumor growth and body fat ratio in vivo
The effects of exosome circ-RNA secreted from adipocytes on tumor growth in vivo were accessed by using tumor-implanted mice. Mouse-derived hepatocellular cancer cells (Hepa1-6 cells) were transfected with lentivirus with a circ-DB overexpressing (circ-DB-OE) sequence or circ-DB shRNA (circ-DB-KD). The subcutaneous implantation of hepatocellular tumor cells was performed in C57 mice and ob/ob mice, and PBS was used as a negative control. The mice were sacrificed at day 28, and the data were analyzed. The diagram of the animal experimental design is shown in Fig. 9a. For statistical significance, each group includes 6 mice. We subsequently isolated exosomes from mouse serum and detected Tsg101, CD63, and Alix using WB analysis (Fig. 9b, c). The level of circ-DB in serum exosomes from ob/ob mice was nearly 5-fold that in serum exosomes from C57 mice (Fig. 9d). The C57 mice implanted with circ-DB-OE cells also displayed obvious upregulation of circ-DB in both serum exosomes and tumor tissues (Fig. 9e, g). The level of circ-DB in serum exosomes showed little change in ob/ob mice (Fig. 9f). Circ-DB in tumor tissues was also clearly increased in the ob/ob group compared with that in the C57 group (Fig. 9g), and the transfection of circ-DB-OE or circ-DB-KD lentivirus led to a clear increase or reduction in circ-DB, respectively (Fig. 9g). This finding suggested that adipose-derived exosomes can deliver circ-DB into HCC tumors in vivo.
a Effects of exosome circ-DB on hepatocellular carcinoma in murine model. Schematic description of the experimental design used to establish the animal model. b TEM photograph of exosomes isolated from mouse plasma. c WB analysis of the exosome markers. d The quantification of circ-DB in serum exosomes isolated from untreated C57 and ob/ob mice (n = 6). e f Relative levels of circ-DB in serum exosomes obtained from C57 mice (e) or ob/ob mice (f) (n = 6). g Relative levels of circ-DB in tumor tissues (n = 6). h Photos of tumors obtained from six groups (n = 6). i–l Tumor weights, mouse weights, body fat ratios, and numbers of liver metastases in each group (n = 6). **p < 0.01
Photos of tumors obtained from each group are shown in Fig. 9h. The weight of tumor tissues (Fig. 9i) and the number of metastases (Fig. 9j) were increased in the circ-DB-OE group of C57 mice but decreased in the circ-DB-KO group of ob/ob mice with tumors. The weight of the ob/ob mice ranged from 30 g to 35 g, and the weight of C57 mice was ~22 g (Fig. 9j). The BFR in ob/ob mice was double that in C57 mice (Fig. 9k). Moreover, the number of liver metastases also doubled in the tumor-implanted ob/ob mice, but the knockdown of circ-DB dramatically suppressed liver metastasis (Fig. 9l). These data illustrated that adipose-secreted circ-DB promoted tumor growth and metastasis.
The expression pattern of the circ-DB/miR-34a/USP7 axis in tumor tissues of mice
The miR-34a levels in tumor tissues showed sharp downregulation in ob/ob mice compared with those in C57 mice (Fig. 8a). The overexpression of circ-DB led to a clear decrease in miR-34a in the tumor tissues of C57 mice, while the absence of circ-DB significantly promoted miR-34a expression in the tumor tissues of ob/ob mice (Fig. 10a). However, USP7 mRNA only showed slight changes in the tumor tissues of each group (Fig. 10b). The expression of both USP7 and Cyclin A2 in C57 mouse tumors was upregulated in the circ-DB-OE group compared with that in the control, while the expression of both USP7 and Cyclin A2 in ob/ob mice was suppressed by the removal of circ-DB (Fig. 10c–f). These data further illustrated that circ-DB works as a miR-34a sponge in HCC tumors and that the downregulation of miR-34a activates USP7/cyclin A2, resulting in enhanced tumor growth and metastasis.
Discussion
Obesity is an independent risk factor that accelerates HCC development [31]. It is well known that obesity is a global epidemic frequently linked with severe diseases and that high body mass index (BMI) in late adolescent men increases the risk of future severe liver disease, including HCC [32]. However, the underlying mechanisms still require further characterization.
It has been reported that exosomes are closely related to tumor development, including angiogenesis, metastasis, drug resistance, and immune escape [33, 34]. The use of exosomes as biological delivery vehicles is of considerable interest for cancer due to a stable environment for a variety of RNAs and chemotherapeutics [35, 36], but the role of adipocyte-derived exosomes in HCC has not been clarified yet.
Noncoding RNAs are abundant in exosomes and are involved in each stage of cancer [37]. A recent study illustrated that adipose-derived exosomes containing various circulating exosome miRNAs regulate gene expression in distant tissues, implying that the exosome is an important carrier in long-distance signal transduction between adipocytes and other cells [8]. The biological role of other components in adipose-derived exosomes has not been reported yet, and here, we showed that adipocyte-secreted circRNAs regulate the deubiquitination-related pathway in HCC cells. CircRNAs, as well as long noncoding RNAs (lncRNAs), have been recognized as miRNA sponges in various types of cells, leading to the suppression of miRNA function. However, the role of secreted circRNAs in targeted cells is far from fully understood.
In this study, the in vivo experiments showed that overexpressed circ-DB in HCC observably downregulated miR-34a, leading to the activation of USP7/Cyclin A2 and tumor progression. Has_circ_0025129 is circRNA that is derived from TNFRSF1A (TNFR1) and is located in the chr12 [38]. It has been reported that deubiquitination of NF-kappaB by USP7 is essential for TLR- and TNFR-induced gene expression [39], thus USP7 may has a positive feedback in regulating TNFR1 and also circ-DB formation.
The upregulation of USP7 mediated by adipose-derived exosomes plays a key role in reducing the ubiquitination of a number of oncogenes. Our result indicates that HCC patients with higher BFR may have rapid tumor progression. Considering that exosomes can also be delivered in a paracrine manner, fatty liver can be a high risk that lead to poor prognosis in HCC.
In conclusion, the present study demonstrated that exosome circRNAs secreted from adipocytes play an important role to promote the tumor growth of HCC through the absorption of miR-34a and activation of the USP7/Cyclin A2 pathway. These findings provide insight into understanding the association between adipose tissues and hepatocellular cancer.
Change history
24 July 2023
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1038/s41388-023-02786-4
References
Ziogas IA, Tsoulfas G. Advances and challenges in laparoscopic surgery in the management of hepatocellular carcinoma. World J Gastrointest Surg. 2017;9:233–45.
Axley P, Ahmed Z, Ravi S, Singal AK. Hepatitis C virus and hepatocellular carcinoma: a narrative review. J Clin Transl Hepatol. 2018;6:79–84.
Petruzziello A. Epidemiology of hepatitis B virus (HBV) and hepatitis C virus (HCV) related hepatocellular carcinoma. Open Virol J. 2018;12:26–32.
Aspichueta P. Lipid-rich environment: a key role promoting carcinogenesis in obesity-related non-alcoholic fatty liver disease. Gut. 2018;67:1376–7.
Gan L, Liu Z, Sun C. Obesity linking to hepatocellular carcinoma: a global view. Biochim Biophys Acta. 2018;1869:97–102.
Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2:569–79.
Yu B, Shao H, Su C, Jiang Y, Chen X, Bai L, et al. Exosomes derived from MSCs ameliorate retinal laser injury partially by inhibition of MCP-1. Sci Rep. 2016;6:34562.
Sun L, Xu R, Sun X, Duan Y, Han Y, Zhao Y, et al. Safety evaluation of exosomes derived from human umbilical cord mesenchymal stromal cell. Cytotherapy2016;18:413–22.
Cooks T, Pateras IS, Jenkins LM, Patel KM, Robles AI, Morris J, et al. Mutant p53 cancers reprogram macrophages to tumor supporting macrophages via exosomal miR-1246. Nat Commun. 2018;9:771.
Wilusz JE, Sharp PA. Molecular biology. A circuitous route to noncoding RNA. Science. 2013;340:440–1.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu JZ. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Li SS, Yao JP, Xie MJ, Liu YN, Zheng M. Exosomal miRNAs in hepatocellular carcinoma development and clinical responses. J Hematology Oncology 2018; 11: 54.
Cao CH, Sun JY, Zhang DY, Guo XJ, Xie LW, Li X, et al. The long intergenic noncoding RNA UFC1, a target of MicroRNA 34a, interacts with the mRNA stabilizing protein HuR to increase levels of beta-catenin in HCC cells. Gastroenterology. 2015;148:415–U249.
Coombs N, Sompallae R, Olbermann P, Gastaldello S, Goppel D, Masucci MG, et al. Helicobacter pylori affects the cellular deubiquitinase USP7 and ubiquitin-regulated components TRAF6 and the tumour suppressor p53. Int J Med Microbiol. 2011;301:213–24.
Zhang C, Lu J, Zhang QW, Zhao W, Guo JH, Liu SL, et al. USP7 promotes cell proliferation through the stabilization of Ki-67 protein in non-small cell lung cancer cells. Int J Biochem Cell Biol. 2016;79:209–21.
Novellasdemunt L, Foglizzo V, Cuadrado L, Antas P, Kucharska A, Encheva V, et al. USP7 is a tumor-specific WNT activator for APC-mutated colorectal cancer by mediating beta-catenin deubiquitination. Cell Rep. 2017;21:612–27.
Morra F, Merolla F, Napolitano V, Ilardi G, Miro C, Paladino S, et al. The combined effect of USP7 inhibitors and PARP inhibitors in hormone-sensitive and castration-resistant prostate cancer cells. Oncotarget. 2017;8:31815–29.
Carra G, Panuzzo C, Torti D, Parvis G, Crivellaro S, Familiari U, et al. Therapeutic inhibition of USP7-PTEN network in chronic lymphocytic leukemia: a strategy to overcome TP53 mutated/deleted clones. Oncotarget. 2017;8:35508–22.
Dong L, Yu L, Bai C, Liu L, Long H, Shi L, et al. USP27-mediated Cyclin E stabilization drives cell cycle progression and hepatocellular tumorigenesis. Oncogene. 2018;37:2702–13.
Wang Q, Ma S, Song N, Li X, Liu L, Yang S, et al. Stabilization of histone demethylase PHF8 by USP7 promotes breast carcinogenesis. J Clin Invest. 2016;126:2205–20.
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9:654–9.
Felicetti F, De Feo A, Coscia C, Puglisi R, Pedini F, Pasquini L, et al. Exosome-mediated transfer of miR-222 is sufficient to increase tumor malignancy in melanoma. J Transl Med. 2016;14:56.
Wei Y, Li MZ, Cui SF, Wang D, Zhang CY, Zen K et al. Shikonin inhibits the proliferation of human breast cancer cells by reducing tumor-derived exosomes. Molecules 2016; 21, 777.
Schmittgen TD, Lee EJ, Jiang JM, Sarkar A, Yang LQ, Elton TS, et al. Real-time PCR quantification of precursor and mature microRNA. Methods. 2008;44:31–38.
Chen CF, Ridzon DA, Broomer AJ, Zhou ZH, Lee DH, Nguyen JT, et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005;33:e179.
Deng X, Zheng H, Li D, Xue Y, Wang Q, Yan S, et al. MicroRNA-34a regulates proliferation and apoptosis of gastric cancer cells by targeting silent information regulator 1. Exp Ther Med. 2018;15:3705–14.
Lv T, Song K, Zhang L, Li W, Chen Y, Diao Y, et al. MiRNA-34a decreases ovarian cancer cell proliferation and chemoresistance by targeting HDAC1. Biochem Cell Biol. 2018;96:663–71.
Malumbres M, Barbacid M. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer. 2009;9:153–66.
Gopinathan L, Tan SL, Padmakumar VC, Coppola V, Tessarollo L, Kaldis P. Loss of Cdk2 and cyclin A2 impairs cell proliferation and tumorigenesis. Cancer Res. 2014;74:3870–9.
Palmer C, Corpuz T, Guirguis M, O’Toole S, Yan K, Bu Y, et al. The effect of obesity on intrahepatic cytokine and chemokine expression in chronic hepatitis C infection. Gut. 2010;59:397–404.
Hagstrom H, Tynelius P, Rasmussen F. High BMI in late adolescence predicts future severe liver disease and hepatocellular carcinoma: a national, population-based cohort study in 1.2 million men. Gut. 2017;67:1536–42.
Yin Y, Cai X, Chen X, Liang HW, Zhang YJ, Li J, et al. Tumor-secreted miR-214 induces regulatory T cells: a major link between immune evasion and tumor growth. Cell Res. 2014;24:1164–80.
Hannafon BN, Carpenter KJ, Berry WL, Janknecht R, Dooley WC, Ding WQ. Exosome-mediated microRNA signaling from breast cancer cells is altered by the anti-angiogenesis agent docosahexaenoic acid (DHA). Mol Cancer. 2015;14:133.
Zhang HY, Bai M, Deng T, Liu R, Wang X, Qu YJ, et al. Cell-derived microvesicles mediate the delivery of miR-29a/c to suppress angiogenesis in gastric carcinoma. Cancer Lett. 2016;375:331–9.
Farooqi AA, Desai NN, Qureshi MZ, Librelotto DRN, Gasparri ML, Bishayee A, et al. Exosome biogenesis, bioactivities and functions as new delivery systems of natural compounds. Biotechnol Adv. 2018;36:328–34.
Ji Q, Zhang CC, Sun XT, Li Q. Circular RNAs function as competing endogenous RNAs in multiple types of cancer. Oncol Lett. 2018;15:23–30.
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO. Cell-type specific features of circular RNA expression. PLoS Genet. 2013;9:e1003777.
Colleran A, Collins PE, O’Carroll C, Ahmed A, Mao XC, McManus B, et al. Deubiquitination of NF-kappa B by Ubiquitin-Specific Protease-7 promotes transcription. Proc Natl Acad Sci USA. 2013;110:618–23.
Acknowledgements
We thank Lei Zhu of Tianjin Medical University Cancer Institute and Hospital for kindly providing information of patients for this study.
Funding
This work was supported by National Natural Science Foundation of China (Nos. 81772629, 81772843, 81702431, 81702275, 81702437, 81602158, and 81602156), Individualized Medical Platform of National Clinical Research Center for Cancer (13ZCZCSY20300) and Demonstrative research platform of clinical evaluation technology for new anticancer drugs (No. 2018ZX09206004). This study was also supported by CAMS Innovation Fund for Medical Sciences (CIFMS, 2016-I2M-1-017 and 2017-I2M-B&R-13), Nature Science Foundation of Tianjin City (16PTSYJC00170) and the Science&Technology Development fund of Tianjin Education Commission for Higher Education (2018KJ046). The funders had no role in study design, collection, analysis, or interpretation of the data, in the writing of the report, or in the decision to submit this article for publication.
Author contributions
HZ and TD performed most of the experiments, analyzed the data, and wrote the manuscript. SG, YL, and MB reviewed and edited the manuscript. KZ and QF performed some of the experiments. YB and GY designed the experiments and edited the manuscript. YB is the guarantor of this work and, having had full access to all data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, H., Deng, T., Ge, S. et al. RETRACTED ARTICLE: Exosome circRNA secreted from adipocytes promotes the growth of hepatocellular carcinoma by targeting deubiquitination-related USP7. Oncogene 38, 2844–2859 (2019). https://doi.org/10.1038/s41388-018-0619-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0619-z
This article is cited by
-
A comprehensive database of exosome molecular biomarkers and disease-gene associations
Scientific Data (2024)
-
White adipocyte-derived exosomal miR-23b inhibits thermogenesis by targeting Elf4 to regulate GLP-1R transcription
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Establishment and Evaluation of Exosomes-Related Gene Risk Model in Hepatocellular Carcinoma
Biochemical Genetics (2024)
-
Exosomal circ_0026611 contributes to lymphangiogenesis by reducing PROX1 acetylation and ubiquitination in human lymphatic endothelial cells (HLECs)
Cellular & Molecular Biology Letters (2023)
-
Exosomal circRNA: emerging insights into cancer progression and clinical application potential
Journal of Hematology & Oncology (2023)