Abstract
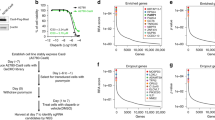
Ataxia telangiectasia mutated and RAD3 related (ATR) protein kinase plays critical roles in ensuring DNA replication, DNA repair, and cell cycle control in response to replication stress, making ATR inhibition a promising therapeutic strategy for cancer treatment. To identify genes whose loss makes tumor cells hypersensitive to ATR inhibition, we performed CRISPR/Cas9-based whole-genome screens in 3 independent cell lines treated with a highly selective ATR inhibitor, AZD6738. These screens uncovered a comprehensive genome-wide profile of ATR inhibitor sensitivity. From the candidate genes, we demonstrated that RNASEH2 deficiency is synthetic lethal with ATR inhibition both in vitro and in vivo. RNASEH2-deficient cells exhibited elevated levels of DNA damage and, when treated with AZD6738, underwent apoptosis (short-time treated) or senescence (long-time treated). Notably, RNASEH2 deficiency is frequently found in prostate adenocarcinoma; we found decreased RNASEH2B protein levels in prostate adenocarcinoma patient-derived xenograft (PDX) samples. Our findings suggest that ATR inhibition may be beneficial for cancer patients with reduced levels of RNASEH2 and that RNASEH2 merits further exploration as a potential biomarker for ATR inhibitor-based therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data not presented in the main figures or Supplementary Data are available from the authors.
References
Saldivar JC, Cortez D, Cimprich KA. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat Rev Mol Cell Biol. 2017;18:622–36.
Keszthelyi A, Minchell NE, Baxter J. The causes and consequences of topological stress during DNA replication. Genes (Basel). 2016;7:pii: E134.
Zeman MK, Cimprich KA. Causes and consequences of replication stress. Nat Cell Biol. 2014;16:2–9.
Techer H, Koundrioukoff S, Nicolas A, Debatisse M. The impact of replication stress on replication dynamics and DNA damage in vertebrate cells. Nat Rev Genet. 2017;18:535–50.
Saldivar JC, Cortez D, Cimprich KA. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat Rev Mol Cell Biol. 2017;18:622–36.
de Klein A, Muijtjens M, van Os R, Verhoeven Y, Smit B, Carr AM. et al. Targeted disruption of the cell-cycle checkpoint gene ATR leads to early embryonic lethality in mice. Curr Biol. 2000;10:479–82.
Brown EJ, Baltimore D. ATR disruption leads to chromosomal fragmentation and early embryonic lethality. Genes Dev. 2000;14:397–402.
Wright JA, et al. Protein kinase mutants of human ATR increase sensitivity to UV and ionizing radiation and abrogate cell cycle checkpoint control. Proc Natl Acad Sci USA. 1998;95:7445–50.
Rundle S, Bradbury A, Drew Y, Curtin NJ. Targeting the ATR-CHK1 axis in cancer therapy. Cancers (Basel). 2017;9:pii: E41.
Charrier JD. et al. Discovery of potent and selective inhibitors of Ataxia telangiectasia mutated and Rad3 related (ATR) protein kinase as potential anticancer agents. J Med Chem. 2011;54:2320–30.
Hall AB, et al. Potentiation of tumor responses to DNA damaging therapy by the selective ATR inhibitor VX-970. Oncotarget. 2014;5:5674–85.
Mohni KN, Kavanaugh GM, Cortez D. ATR pathway inhibition is synthetically lethal in cancer cells with ERCC1 deficiency. Cancer Res. 2014;74:2835–45.
Williamson CT. et al. ATR inhibitors as a synthetic lethal therapy for tumours deficient in ARID1A. Nat Commun. 2016;7:13837.
Kwok M, Davies N, Agathanggelou A. et al. ATR inhibition induces synthetic lethality and overcomes chemoresistance in TP53- or ATM-defective chronic lymphocytic leukemia cells. Blood. 2016;127:582–95.
Reaper PM, et al. Selective killing of ATM- or p53-deficient cancer cells through inhibition of ATR. Nat Chem Biol. 2011;7:428–30.
Prevo R, et al. The novel ATR inhibitor VE-821 increases sensitivity of pancreatic cancer cells to radiation and chemotherapy. Cancer Biol Ther. 2012;13:1072–81.
Fokas E. et al. Targeting ATR in vivo using the novel inhibitor VE-822 results in selective sensitization of pancreatic tumors to radiation. Cell Death Dis. 2012;3:e441.
Zhu S, Zhou Y, Wei W. Genome-wide CRISPR/Cas9 screening for high-throughput functional genomics in human cells. Methods Mol Biol. 2017;1656:175–81.
Steinhart Z, et al. Genome-wide CRISPR screens reveal a Wnt-FZD5 signaling circuit as a druggable vulnerability of RNF43-mutant pancreatic tumors. Nat Med. 2017;23:60–8.
Sidik SM, et al. A genome-wide CRISPR screen in toxoplasma identifies essential apicomplexan genes. Cell. 2016;166:1423–35 e1412.
Ruiz S, et al. A genome-wide CRISPR screen identifies CDC25A as a determinant of sensitivity to ATR inhibitors. Mol Cell. 2016;62:307–13.
Hart T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell. 2015;163:1515–26.
Hart T, Brown KR, Sircoulomb F, Rottapel R, Moffat J. Measuring error rates in genomic perturbation screens: gold standards for human functional genomics. Mol Syst Biol. 2014;10:733.
Hart T, Moffat J. BAGEL: a computational framework for identifying essential genes from pooled library screens. BMC Bioinformatics. 2016;17:164.
Checkley S. et al. Bridging the gap between in vitro and in vivo: Dose and schedule predictions for the ATR inhibitor AZD6738. Sci Rep. 2015;5:13545.
Guichard SM, Brown E, Odedra R, Hughes A, Heathcote D, Barnes J, et al. The pre-clinical in vitro and in vivo activity of AZD6738: a potent and selective inhibitor of ATR kinase. Cancer Res. 73, https://doi.org/10.1158/1538-7445.Am2013-3343 (2013).
Hiller B, et al. Mammalian RNase H2 removes ribonucleotides from DNA to maintain genome integrity. J Exp Med. 2012;209:1419–26.
Perrino FW, Harvey S, Shaban NM, Hollis T. RNaseH2 mutants that cause Aicardi-Goutieres syndrome are active nucleases. J Mol Med. 2009;87:25–30.
Pendergraft WF 3rd, Means TK. AGS, SLE, and RNASEH2 mutations: translating insights into therapeutic advances. J Clin Invest. 2015;125:102–4.
Crow YJ, et al. Mutations in genes encoding ribonuclease H2 subunits cause Aicardi-Goutieres syndrome and mimic congenital viral brain infection. Nat Genet. 2006;38:910–6.
Mackenzie KJ, et al. Ribonuclease H2 mutations induce a cGAS/STING-dependent innate immune response. EMBO J. 2016;35:831–44.
Pokatayev V, et al. RNase H2 catalytic core Aicardi-Goutieres syndrome-related mutant invokes cGAS-STING innate immune-sensing pathway in mice. J Exp Med. 2016;213:329–36.
Williams JS, Gehle DB, Kunkel TA. The role of RNase H2 in processing ribonucleotides incorporated during DNA replication. DNA Repair (Amst). 2017;53:52–8.
Pizzi S, et al. Reduction of hRNase H2 activity in Aicardi-Goutieres syndrome cells leads to replication stress and genome instability. Hum Mol Genet. 2015;24:649–58.
Chon H, et al. RNase H2 roles in genome integrity revealed by unlinking its activities. Nucleic Acids Res. 2013;41:3130–43.
McElhinny SAN, et al. Genome instability due to ribonucleotide incorporation into DNA. Nat Chem Biol. 2010;6:774–81.
Reijns MAM. et al. Enzymatic removal of ribonucleotides from DNA is essential for mammalian genome integrity and development. Cell. 2012;149:1008–22.
Zimmermann M, et al. CRISPR screens identify genomic ribonucleotides as a source of PARP-trapping lesions. Nature. 2018;559:285–9.
Hart T, et al. Evaluation and Design of Genome-Wide CRISPR/SpCas9 Knockout Screens. G3-Genes Genom Genet. 2017;7:2719–27.
Kim H, George E, Brown E, Zhang RG, Krepler C, Tanyi J, et al. Targeting the ATR/CHK1 axis in BRCA1/2 mutant ovarian cancer using an orthotopic patient-derived xenograft (PDX) model. Clin Cancer Res. 2016;22 (2 Suppl).
Kim H, et al. Targeting the ATR/CHK1 axis with PARP inhibition results in tumor regression in BRCA-mutant ovarian cancer models. Clin Cancer Res. 2017;23:3097–108.
Huang S, et al. MED12 controls the response to multiple cancer drugs through regulation of TGF-beta receptor signaling. Cell. 2012;151:937–50.
He T, et al. Methylation of SLFN11 is a marker of poor prognosis and cisplatin resistance in colorectal cancer. Epigenomics. 2017;9:849–62.
Mu Y, et al. SLFN11 inhibits checkpoint maintenance and homologous recombination repair. EMBO Rep. 2016;17:94–109.
Bartsch K, et al. Absence of RNase H2 triggers generation of immunogenic micronuclei removed by autophagy. Hum Mol Genet. 2017;26:3960–72.
Rice GI, et al. Synonymous mutations in RNASEH2A create cryptic splice sites impairing RNase H2 enzyme function in Aicardi-Goutieres syndrome. Hum Mutat. 2013;34:1066–70.
Grasso CS, et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature. 2012;487:239–43.
Robinson D, et al. Integrative clinical genomics of advanced prostate. Cancer Cell. 2015;162:454.
Kumar A, et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat Med. 2016;22:369.
Abeshouse A, et al. The molecular taxonomy of primary prostate. Cancer Cell. 2015;163:1011–25.
Taylor BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Uhlen M, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Gao J, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Li T, Chen ZJ. The cGAS-cGAMP-STING pathway connects DNA damage to inflammation, senescence, and cancer. J Exp Med. 2018;215:1287–99.
Yang H, Wang H, Ren J, Chen Q, Chen ZJ. cGAS is essential for cellular senescence. Proc Natl Acad Sci USA. 2017;114:E4612–20.
Hart T, et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell. 2015;163:1515–26.
Acknowledgements
We thank the members of Dr. Junjie Chen’s lab for their kind help and Dr. Lei Li for his suggestions regarding the experimental design. We also thank Amy Ninetto from the Department of Scientific Publications at MD Anderson for editing the manuscript. This work was supported in part by CPRIT (RP160667) and NIH grants (CA157448, CA193124, CA210929, CA216911, and CA216437) to JC and MD Anderson’s NIH Cancer Center Support Grant (CA016672).
Author contributions
CW and JC conceived the project. CW, JZ, XF, MT, ZC, and MS performed the experiments. MEM provided technical support for the screen work. GW and GTH analyzed the deep-sequencing results. PS and NMN provided and analyzed the PDX samples. CW and JC wrote the manuscript with input from all authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, C., Wang, G., Feng, X. et al. Genome-wide CRISPR screens reveal synthetic lethality of RNASEH2 deficiency and ATR inhibition. Oncogene 38, 2451–2463 (2019). https://doi.org/10.1038/s41388-018-0606-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0606-4
This article is cited by
-
Targeting ATR in patients with cancer
Nature Reviews Clinical Oncology (2024)
-
From pre-clinical to translational brain metastasis research: current challenges and emerging opportunities
Clinical & Experimental Metastasis (2024)
-
Genome-wide CRISPR/Cas9 library screening identified ATM signaling network genes as critical drivers for resistance to ATR inhibition in soft-tissue sarcomas: synthetic lethality and therapeutic implications
Experimental Hematology & Oncology (2023)
-
Revolutionizing DNA repair research and cancer therapy with CRISPR–Cas screens
Nature Reviews Molecular Cell Biology (2023)
-
ATR kinase supports normal proliferation in the early S phase by preventing replication resource exhaustion
Nature Communications (2023)