Abstract
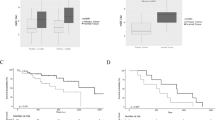
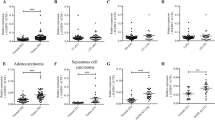
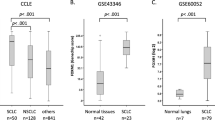
Lung cancer is one of the major causes of cancer death and clarification of its molecular pathology is highly prioritized. The physiological importance of mRNA degradation through the CCR4-NOT deadenylase has recently been highlighted. For example, mutation in CNOT3, a gene coding for CNOT3 subunit of the CCR4-NOT complex, is found to be associated with T-cell acute lymphoblastic leukemia, T-ALL, though its contribution to other cancers has not been reported. Here, we provide evidence suggesting that CNOT3 is required for the growth of non-small cell lung cancer. Depletion of CNOT3 suppresses proliferation of A549 human non-small cell lung cancer cells with enhanced mRNA stability and subsequent elevated expression of p21. In addition, we identified the mRNA for Krüppel-like factor 2 transcription factor, an inducer of p21, as a novel mRNA degradation target of CNOT3 in non-small cell lung cancer cells. Aberrant up-regulation of Krüppel-like factor 2 by CNOT3 depletion leads to impairment in the proliferation of A549 cells. Consistent with these findings, elevated mRNA expression of CNOT3 in non-small cell lung cancer in comparison with the paired normal lung epithelium was confirmed through scrutinization of the RNA-sequencing datasets from The Cancer Genome Atlas. Moreover, we found an inverse correlation between CNOT3 and CDKN1A (encoding p21) mRNA expression using the combined datasets of normal lung epithelium and non-small cell lung cancer. Thus, we propose that the up-regulation of CNOT3 facilitates the development of non-small cell lung cancer through down-regulation of Krüppel-like factor 2 and p21, contrary to tumor suppressive functions of CNOT3 in T-ALL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30.
Dela Cruz CS, Tanoue LT, Matthay RA. Lung cancer: epidemiology, etiology, and prevention. Clin Chest Med. 2011;32:605–44.
Petersen I, Warth A. Lung cancer: developments, concepts, and specific aspects of the new WHO classification. J Cancer Res Clin Oncol. 2016;142:895–904.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321–33.
Goldstrohm AC, Wickens M. Multifunctional deadenylase complexes diversify mRNA control. Nat Rev Mol Cell Biol. 2008;9:337–44.
Bartlam M, Yamamoto T. The structural basis for deadenylation by the CCR4-NOT complex. Protein Cell. 2010;1:443–52.
Castro D, Moreira M, Gouveia AM, Pozza DH, De Mello RA. MicroRNAs in lung cancer. Oncotarget. 2017;8:81679–85.
Lau NC, Kolkman A, van Schaik FM, Mulder KW, Pijnappel WW, Heck AJ, et al. Human Ccr4-Not complexes contain variable deadenylase subunits. Biochem J. 2009;422:443–53.
Ito K, Takahashi A, Morita M, Suzuki T, Yamamoto T. The role of the CNOT1 subunit of the CCR4-NOT complex in mRNA deadenylation and cell viability. Protein Cell. 2011;2:755–63.
Suzuki T, Kikuguchi C, Sharma S, Sasaki T, Tokumasu M, Adachi S, et al. CNOT3 suppression promotes necroptosis by stabilizing mRNAs for cell death-inducing proteins. Sci Rep. 2015;5:14779.
Morita M, Oike Y, Nagashima T, Kadomatsu T, Tabata M, Suzuki T, et al. Obesity resistance and increased hepatic expression of catabolism-related mRNAs in Cnot3+/− mice. EMBO J. 2011;30:4678–91.
Zheng X, Yang P, Lackford B, Bennett BD, Wang L, Li H, et al. CNOT3-dependent mRNA deadenylation safeguards the pluripotent state. Stem Cell Reports. 2016;7:897–910.
Watanabe C, Morita M, Hayata T, Nakamoto T, Kikuguchi C, Li X, et al. Stability of mRNA influences osteoporotic bone mass via CNOT3. Proc Natl Acad Sci USA. 2014;111:2692–7.
De Keersmaecker K, Atak ZK, Li N, Vicente C, Patchett S, Girardi T, et al. Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat Genet. 2013;45:186–90.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6:1–6.
Selamat SA, Chung BS, Girard L, Zhang W, Zhang Y, Campan M, et al. Genome-scale analysis of DNA methylation in lung adenocarcinoma and integration with mRNA expression. Genome Res. 2012;22:1197–211.
Hou J, Aerts J, den Hamer B, van Ijcken W, den Bakker M, Riegman P, et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLoS One. 2010;5:e10312.
Okayama H, Kohno T, Ishii Y, Shimada Y, Shiraishi K, Iwakawa R, et al. Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res. 2012;72:100–11.
Landi MT, Dracheva T, Rotunno M, Figueroa JD, Liu H, Dasgupta A, et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One. 2008;3:e1651.
Su LJ, Chang CW, Wu YC, Chen KC, Lin CJ, Liang SC, et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics. 2007;8:140.
Kanamori-Katayama M, Itoh M, Kawaji H, Lassmann T, Katayama S, Kojima M, et al. Unamplified cap analysis of gene expression on a single-molecule sequencer. Genome Res. 2011;21:1150–9.
Forrest AR, Kawaji H, Rehli M, Baillie JK, de Hoon MJ, Haberle V, et al. A promoter-level mammalian expression atlas. Nature. 2014;507:462–70.
Zheng X, Dumitru R, Lackford BL, Freudenberg JM, Singh AP, Archer TK, et al. Cnot1, Cnot2, and Cnot3 maintain mouse and human ESC identity and inhibit extraembryonic differentiation. Stem Cells. 2012;30:910–22.
Neely GG, Kuba K, Cammarato A, Isobe K, Amann S, Zhang L, et al. A global in vivo Drosophila RNAi screen identifies NOT3 as a conserved regulator of heart function. Cell. 2010;141:142–53.
Cejas P, Cavazza A, Yandava CN, Moreno V, Horst D, Moreno-Rubio J, et al. Transcriptional regulator CNOT3 defines an aggressive colorectal cancer subtype. Cancer Res. 2017;77:766–79.
Takahashi A, Kikuguchi C, Morita M, Shimodaira T, Tokai-Nishizumi N, Yokoyama K, et al. Involvement of CNOT3 in mitotic progression through inhibition of MAD1 expression. Biochem Biophys Res Commun. 2012;419:268–73.
Nabel EG. CDKs and CKIs: molecular targets for tissue remodelling. Nat Rev Drug Discov. 2002;1:587–98.
Kawabe S, Roth JA, Wilson DR, Meyn RE. Adenovirus-mediated p16INK4a gene expression radiosensitizes non-small cell lung cancer cells in a p53-dependent manner. Oncogene. 2000;19:5359–66.
Schuster K, Venkateswaran N, Rabellino A, Girard L, Pena-Llopis S, Scaglioni PP. Nullifying the CDKN2AB locus promotes mutant K-ras lung tumorigenesis. Mol Cancer Res. 2014;12:912–23.
Wu J, Lingrel JB. KLF2 inhibits Jurkat T leukemia cell growth via upregulation of cyclin-dependent kinase inhibitor p21WAF1/CIP1. Oncogene. 2004;23:8088–96.
Wani MA, Wert SE, Lingrel JB. Lung Kruppel-like factor, a zinc finger transcription factor, is essential for normal lung development. J Biol Chem. 1999;274:21180–5.
Winkelmann R, Sandrock L, Kirberg J, Jack HM, Schuh W. KLF2—a negative regulator of pre-B cell clonal expansion and B cell activation. PLoS One. 2014;9:e97953.
Tetreault MP, Yang Y, Katz JP. Kruppel-like factors in cancer. Nat Rev Cancer. 2013;13:701–13.
Yin L, Wang JP, Xu TP, Chen WM, Huang MD, Xia R, et al. Downregulation of Kruppel-like factor 2 is associated with poor prognosis for nonsmall-cell lung cancer. Tumour Biol. 2015;36:3075–84.
Harris T, Pan Q, Sironi J, Lutz D, Tian J, Sapkar J, et al. Both gene amplification and allelic loss occur at 14q13.3 in lung cancer. Clin Cancer Res. 2011;17:690–9.
Sordella R, Bell DW, Haber DA, Settleman J. Gefitinib-sensitizing EGFR mutations in lung cancer activate anti-apoptotic pathways. Science. 2004;305:1163–7.
Shirai YT, Suzuki T, Morita M, Takahashi A, Yamamoto T. Multifunctional roles of the mammalian CCR4-NOT complex in physiological phenomena. Front Genet. 2014;5:286.
Faraji F, Hu Y, Wu G, Goldberger NE, Walker RC, Zhang J, et al. An integrated systems genetics screen reveals the transcriptional structure of inherited predisposition to metastatic disease. Genome Res. 2014;24:227–40.
Faraji F, Hu Y, Yang HH, Lee MP, Winkler GS, Hafner M, et al. Post-transcriptional control of tumor cell autonomous metastatic potential by CCR4-NOT deadenylase CNOT7. PLoS Genet. 2016;12:e1005820.
Mittal S, Aslam A, Doidge R, Medica R, Winkler GS. The Ccr4a (CNOT6) and Ccr4b (CNOT6L) deadenylase subunits of the human Ccr4-Not complex contribute to the prevention of cell death and senescence. Mol Biol Cell. 2011;22:748–58.
Aslam A, Mittal S, Koch F, Andrau JC, Winkler GS. The Ccr4-NOT deadenylase subunits CNOT7 and CNOT8 have overlapping roles and modulate cell proliferation. Mol Biol Cell. 2009;20:3840–50.
Jiang W, Xu X, Deng S, Luo J, Xu H, Wang C, et al. Methylation of Kruppel-like factor 2 (KLF2) associates with its expression and non-small cell lung cancer progression. Am J Transl Res. 2017;9:2024–37.
Wehrkamp CJ, Natarajan SK, Mohr AM, Phillippi MA, Mott JL. miR-106b-responsive gene landscape identifies regulation of Kruppel-like factor family. RNA Biol. 2017;15:391–403.
Li W, Sun M, Zang C, Ma P, He J, Zhang M, et al. Upregulated long non-coding RNA AGAP2-AS1 represses LATS2 and KLF2 expression through interacting with EZH2 and LSD1 in non-small-cell lung cancer cells. Cell Death Dis. 2016;7:e2225.
Inoue T, Morita M, Hijikata A, Fukuda-Yuzawa Y, Adachi S, Isono K, et al. CNOT3 contributes to early B cell development by controlling Igh rearrangement and p53 mRNA stability. J Exp Med. 2015;212:1465–79.
Anaya J. OncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Comput Sci. 2016;2:e67.
Campbell JD, Alexandrov A, Kim J, Wala J, Berger AH, Pedamallu CS, et al. Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet. 2016;48:607–16.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2:401–4.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E, et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell. 2012;150:1107–20.
Horie M, Yamaguchi Y, Saito A, Nagase T, Lizio M, Itoh M, et al. Transcriptome analysis of periodontitis-associated fibroblasts by CAGE sequencing identified DLX5 and RUNX2 long variant as novel regulators involved in periodontitis. Sci Rep. 2016;6:33666.
Andersson R, Gebhard C, Miguel-Escalada I, Hoof I, Bornholdt J, Boyd M, et al. An atlas of active enhancers across human cell types and tissues. Nature. 2014;507:455–61.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Horie M, Kaczkowski B, Ohshima M, Matsuzaki H, Noguchi S, Mikami Y, et al. Integrative CAGE and DNA methylation profiling identify epigenetically regulated genes in NSCLC. Mol Cancer Res. 2017;15:1354–65.
Hoshino Y, Katsuno Y, Ehata S, Miyazono K. Autocrine TGF-beta protects breast cancer cells from apoptosis through reduction of BH3-only protein, Bim. J Biochem. 2011;149:55–65.
Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–10.
Shirai YT, Ehata S, Yashiro M, Yanagihara K, Hirakawa K, Miyazono K. Bone morphogenetic protein-2 and -4 play tumor suppressive roles in human diffuse-type gastric carcinoma. Am J Pathol. 2011;179:2920–30.
Hoshina N, Tanimura A, Yamasaki M, Inoue T, Fukabori R, Kuroda T, et al. Protocadherin 17 regulates presynaptic assembly in topographic corticobasal Ganglia circuits. Neuron. 2013;78:839–54.
Morita M, Suzuki T, Nakamura T, Yokoyama K, Miyasaka T, Yamamoto T. Depletion of mammalian CCR4b deadenylase triggers elevation of the p27Kip1 mRNA level and impairs cell growth. Mol Cell Biol. 2007;27:4980–90.
Acknowledgements
The authors would like to thank Dr. Hiroyuki Miyoshi (RIKEN) for lentivirus vector system. The authors also thank all the members of Cell Signal Unit at the Okinawa Institute of Science and Technology Graduate University (OIST) for discussion and advice. This research was supported by subsidiary of OIST and KAKENHI Grants-in-Aid for scientific research (S) (21229006 to TY), for Scientific Research on Innovative Areas (25121734 to TY), for Young Scientists (B) (26830066 to AM and 16K18437 to MH), and for Research Activity Start-up (24890289 to Y-TS and 26893050 to MH) from the Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Supplementary information
Rights and permissions
About this article
Cite this article
Shirai, YT., Mizutani, A., Nishijima, S. et al. CNOT3 targets negative cell cycle regulators in non-small cell lung cancer development. Oncogene 38, 2580–2594 (2019). https://doi.org/10.1038/s41388-018-0603-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0603-7
This article is cited by
-
Targeting the up-regulated CNOT3 reverses therapeutic resistance and metastatic progression of EGFR-mutant non-small cell lung cancer
Cell Death Discovery (2023)
-
Identification of a miRNA multi-targeting therapeutic strategy in glioblastoma
Cell Death & Disease (2023)
-
Translation factor eIF5a is essential for IFNγ production and cell cycle regulation in primary CD8+ T lymphocytes
Nature Communications (2022)
-
RHBDD1 silencing inhibited cell growth and invasion of non-small cell lung cancer by mediating ZEB1/PI3K/AKT signaling pathway
Journal of Molecular Histology (2021)
-
DNA methylation predicts age and provides insight into exceptional longevity of bats
Nature Communications (2021)