Abstract
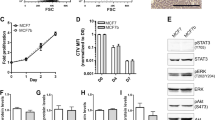
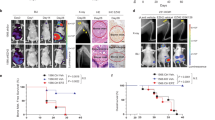
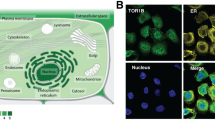
Bone is the most common metastatic site for breast cancer. Estrogen-related-receptor alpha (ERRα) has been implicated in cancer cell invasiveness. Here, we established that ERRα promotes spontaneous metastatic dissemination of breast cancer cells from primary mammary tumors to the skeleton. We carried out cohort studies, pharmacological inhibition, gain-of-function analyses in vivo and cellular and molecular studies in vitro to identify new biomarkers in breast cancer metastases. Meta-analysis of human primary breast tumors revealed that high ERRα expression levels were associated with bone but not lung metastases. ERRα expression was also detected in circulating tumor cells from metastatic breast cancer patients. ERRα overexpression in murine 4T1 breast cancer cells promoted spontaneous bone micro-metastases formation when tumor cells were inoculated orthotopically, whereas lung metastases occurred irrespective of ERRα expression level. In vivo, Rank was identified as a target for ERRα. That was confirmed in vitro in Rankl stimulated tumor cell invasion, in mTOR/pS6K phosphorylation, by transactivation assay, ChIP and bioinformatics analyses. Moreover, pharmacological inhibition of ERRα reduced primary tumor growth, bone micro-metastases formation and Rank expression in vitro and in vivo. Transcriptomic studies and meta-analysis confirmed a positive association between metastases and ERRα/RANK in breast cancer patients and also revealed a positive correlation between ERRα and BRCA1mut carriers. Taken together, our results reveal a novel ERRα/RANK axis by which ERRα in primary breast cancer promotes early dissemination of cancer cells to bone. These findings suggest that ERRα may be a useful therapeutic target to prevent bone metastases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lambert A, Pattabiraman D, Weinberg R. Emerging biological principles of metastasis. Cell. 2017;168:670–91.
Kan C, Vargas G, Pape F, Clézardin P. Cancer cell colonisation in the bone microenvironment. Int J Mol Sci. 2016;17:E1674.
Brodowicz T, Hadji P, Niepel D, Diel I. Early identification and intervention matters: a comprehensive review of current evidence and recommendations for the monitoring of bone health in patients with cancer. Cancer Treat Rev. 2017;61:23–34.
D’Oronzo S, Brown J, Coleman R. The role of biomarkers in the management of bone-homing malignancies. JBone Oncol. 2017;9:1–9.
Lacey D, Boyle W, Simonet W, Kostenuik P, Dougall W, Sullivan J, et al. Bench to bedside: elucidation of the OPG-RANK-RANKL pathway and the development of denosumab. Nat Rev Drug Discov. 2012;11:401–19.
Benoit G, Cooney A, Giguere V, Ingraham H, Lazar M, Muscat G, et al. International Union of Pharmacology. LXVI. Orphan nuclear receptors. Pharmacol Rev. 2006;58:798–836.
Giguere V, Yang N, Segui P, Evans RM. Identification of a new class of steroid hormone receptors. Nature. 1988;331:91–4.
Willy PJ, Murray IR, Qian J, Busch BB, Stevens WC Jr, Martin R, et al. Regulation of PPARgamma coactivator 1alpha (PGC-1alpha) signaling by an estrogen-related receptor alpha (ERRalpha) ligand. Proc Natl Acad Sci USA. 2004;101:8912–7.
Patch RJ, Searle LL, Kim AJ, De D, Zhu X, Askari HB, et al. Identification of diaryl ether-based ligands for estrogen-related receptor alpha as potential antidiabetic agents. J Med Chem. 2011;54:788–808.
Audet-Walsh E, Giguére V. The multiple universes of estrogen-related receptor α and γ in metabolic control and related diseases. Acta Pharmacol Sin. 2015;36:51–61.
Ariazi EA, Clark GM, Mertz JE. Estrogen-related receptor alpha and estrogen-related receptor gamma associate with unfavorable and favorable biomarkers, respectively, in human breast cancer. Cancer Res. 2002;62:6510–8.
Wu D, Cheung A, Wang Y, Yu S, Chan F. The emerging roles of orphan nuclear receptors in prostate cancer. Biochim Biophys Acta. 2016;1866:23–36.
Tam I, Giguère G. There and back again: the journey of the estrogen-related receptors in the cancer realm. J Steroid Biochem Mol Biol. 2016;157:13–9.
Deblois GGV. Oestrogen-related receptors in breast cancer: control of cellular metabolism and beyond. Nat Rev Cancer. 2013;13:27–36.
Dwyer MA, Joseph JD, Wade HE, Eaton ML, Kunder RS, Kazmin D, et al. WNT11 expression is induced by estrogen-related receptor alpha and beta-catenin and acts in an autocrine manner to increase cancer cell migration. Cancer Res. 2010;70:9298–308.
Bonnelye E, Aubin JE. An energetic orphan in an endocrine tissue: a revised perspective of the function of estrogen receptor-related receptor alpha in bone and cartilage. J Bone Miner Res. 2013;28:225–33.
Bae S, Lee M, Mun S, Giannopoulou E, Yong-Gonzalez V, Cross J, et al. MYC-dependent oxidative metabolism regulates osteoclastogenesis via nuclear receptor ERRα. J Clin Invest. 2017;127:2555–68.
Delhon I, Gutzwiller S, Morvan F, Rangwala S, Wyder L, Evans G, et al. Absence of estrogen receptor-related-alpha increases osteoblastic differentiation and cancellous bone mineral density. Endocrinology. 2009;150:4463–72.
Wei WWX, Yang M, Smith LC, Dechow PC, Sonoda J, Evans RM, et al. PGC1beta mediates PPARgamma activation of osteoclastogenesis and rosiglitazone-induced bone loss. Cell Metab. 2010;11:503–16.
Fradet A, Sorel H, Bouazza L, Goehrig D, Depalle B, Bellahcene A, et al. Dual function of ERRalpha in breast cancer and bone metastasis formation: implication of VEGF and osteoprotegerin. Cancer Res. 2011;71:5728–38.
Fradet A, Bouchet M, Delliaux C, Gervais G, Kan C, Benetollo C, et al. Estrogen related receptor alpha in castration-resistant prostate cancer cells promotes tumor progression in bone. Oncotarget. 2016;7:77071–86.
Schmid M, Jakesz R, Samonigg H, Kubista E, Gnant M, Menzel C, et al. Randomized trial of tamoxifen versus tamoxifen plus aminoglutethimide as adjuvant treatment in postmenopausal breast cancer patients with hormone receptor-positive disease: Austrian breast and colorectal cancer study group trial 6. J Clin Oncol. 2003;21:984–90.
van de Vijver M, He Y, van’t Veer L, Dai H, Hart A, Voskuil D, et al. A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med. 2002;19:1999–2009.
Souza Garcia C, Rios de Araujo M, Paz Lopes M, Ferreira M, Dantas Cassali G. Morphological and immunophenotipical characterization of murine mammary carcinoma 4t1. Braz J Vet Pathol. 2014;7:158–65.
Aslakson CJ, Miller FR. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 1992;52:1399–405.
Stein RA, Gaillard S, McDonnell DP. Estrogen-related receptor alpha induces the expression of vascular endothelial growth factor in breast cancer cells. J Steroid Biochem Mol Biol. 2009;114:106–12.
Boyle WJ, Simonet WS, Lacey DL. Osteoclast differentiation and activation. Nature. 2003;423:337–42.
Santini D, Schiavon G, Vincenzi B, Gaeta L, Pantano F, Russo A, et al. Receptor activator of NF-kB (RANK) expression in primary tumors associates with bone metastasis occurrence in breast cancer patients. PLoS One. 2011;6:e19234.
Jones DH, Nakashima T, Sanchez OH, Kozieradzki I, Komarova SV, Sarosi I, et al. Regulation of cancer cell migration and bone metastasis by RANKL. Nature. 2006;440:692–6.
Hong E, Levasseur M, Dufour C, Perry M, Giguère V. Loss of estrogen-related receptor α promotes hepatocarcinogenesis development via metabolic and inflammatory disturbances. Proc Natl Acad Sci USA. 2013;110:17975–80.
Bartkowiak K, Kwiatkowski M, Buck F, Gorges T, Nilse L, Assmann V, et al. Disseminated tumor cells persist in the bone marrow of breast cancer patients through sustained activation of the unfolded protein response. Cancer Res. 2015;75:5367–77.
Carnesecchi J, Forcet C, Zhang L, Tribollet V, Barenton B, Boudra R, et al. ERRα induces H3K9 demethylation by LSD1 to promote cell invasion. Proc Natl Acad Sci USA. 2017;114:3909–14.
Deblois G, Smith H, Tam I, Gravel S, Caron M, Savage P, et al. ERRα mediates metabolic adaptations driving lapatinib resistance in breast cancer. Nat Commun. 2016;12:12156.
Stark C, Breitkreutz B, Reguly T, Boucher L, Breitkreutz A, Tyers M. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 2006;34:D535–9.
Zhang X, Wang Q, Gerald W, Hudis C, Norton L, Smid M, et al. Latent bone metastasis in breast cancer tied to Src-dependent survival signals. Cancer Cell. 2009;16:67–78.
Bieche I, Parfait B, Le Doussal V, Olivi M, Rio M, Lidereau R, et al. Identification of CGA as a novel estrogen receptor-responsive gene in breast cancer: an outstanding candidate marker to predict the response to endocrine therapy. Cancer Res. 2001;61:1652–8.
Sigl V, Owusu-Boaitey K, Joshi P, Kavirayani A, Wirnsberger G, Novatchkova M, et al. RANKL/RANK control Brca1 mutation-driven mammary tumors. Cell Res. 2016;26:761–74.
Castet A, Herledan A, Bonnet S, Jalaguier S, Vanacker JM, Cavailles V. Receptor-interacting protein 140 differentially regulates estrogen receptor-related receptor transactivation depending on target genes. Mol Endocrinol. 2006;20:1035–47.
Sumi D, Ignarro L. Sp1 transcription factor expression is regulated by estrogen-related receptor alpha1. Biochem Biophys Res Commun. 2005;328:165–72.
Zhang L, Teng Y, Zhang Y, Liu J, Xu L, Qu J, et al. Receptor activator for nuclear factor κ B expression predicts poor prognosis in breast cancer patients with bone metastasis but not in patients with visceral metastasis. J Clin Pathol. 2012;65:36–40.
Campbell J, Karolak M, Ma Y, Perrien D, Masood-Campbell S, Penner N et al. Stimulation of host bone marrow stromal cells by sympathetic nerves promotes breast cancer bone metastasis in mice. PLoS Biol [Internet]. 2012. https://doi.org/10.1371/journal.pbio.1001363
Blake M, Tometsko M, Miller R, Jones J, Dougall W. RANK expression on breast cancer cells promotes skeletal metastasis. Clin Exp Metastas. 2014;31:233–45.
Zeng R, Faccio R, Novack D. Alternative NF-κB regulates RANKL-induced osteoclast differentiation and mitochondrial biogenesis via independent mechanisms. J Bone Miner Res. 2015;30:2287–99.
Dirckx N, Tower R, Mercken E, Vangoitsenhoven R, Moreau-Triby C, Breugelmans T, et al. Vhl deletion in osteoblasts boosts cellular glycolysis and improves global glucose metabolism. J Clin Invest. 2018;128:1087–105.
Takeno A, Kanazawa I, Notsu M, Tanaka K, Sugimoto T. Glucose uptake inhibition decreases expressions of receptor activator of nuclear factor-kappa B ligand (RANKL) and osteocalcin in osteocytic MLO-Y4-A2 cells. Am J Physiol Endocrinol Metab. 2018;314:E115–23.
Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin A, Kim S, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Salatino S, Kupr B, Baresic M, van Nimwegen E, Handschin C. The genomic context and corecruitment of SP1 affect ERRα coactivation by PGC-1α in muscle cells. Mol Endocrinol. 2016;30:809–25.
Nolan E, Vaillant F, Branstetter D, Pal B, Giner G, Whitehead L, et al. RANK ligand as a potential target for breast cancer prevention in BRCA1-mutation carriers. Nat Med. 2016;22:933–9.
Ishibashi H, Suzuki T, Suzuki S, Moriya T, Kaneko C, Takizawa T, et al. Sex steroid hormone receptors in human thymoma. J Clin Endocrinol Metab. 2003;88:2309–17.
Rakha E, El-Sayed M, Lee A, Elston C, Grainge M, Hodi Z, et al. Prognostic significance of nottingham histologic grade in invasive breast carcinoma. J Clin Oncol. 2008;26:3153–8.
Curtis C, Shah S, Chin S, Turashvili G, Rueda O, Dunning M, et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346–52.
Bos P, Zhang X, Nadal C, Shu W, Gomis R, Nguyen D, et al. Genes that mediate breast cancer metastasis to the brain. Nature. 2009;459:1005–9.
Wang Y, Klijn J, Zhang Y, Sieuwerts A, Look M, Yang F, et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet. 2005;365:671–9.
Minn A, Gupta G, Siegel P, Bos P, Shu W, Giri D, et al. Genes that mediate breast cancer metastasis to lung. Nature. 2005;436:518–24.
Pantel K, Dickmanns A, Zippelius A, Klein C, Shi J, Hoechtlen-Vollmar W, et al. Establishment of micrometastatic carcinoma cell lines: a novel source of tumor cell vaccines. J Natl Cancer Inst. 1995;87:1162–8.
Bonnelye E, Saltel F, Chabadel A, Zirngibl RA, Aubin JE, Jurdic P. Involvement of the orphan nuclear estrogen receptor-related receptor alpha in osteoclast adhesion and transmigration. J Mol Endocrinol. 2010;45:365–77.
Chaveroux CEL, Dufour CR, Shatnawi A, Khoutorsky A, Bourque G, Sonenberg N, et al. Molecular and genetic crosstalks between mTOR and ERRα are key determinants of rapamycin-induced nonalcoholic fatty liver. Cell Metab. 2013;17:586–98.
David M, Ribeiro J, Descotes F, Serre CM, Barbier M, Murone M, et al. Targeting lysophosphatidic acid receptor type 1 with Debio 0719 inhibits spontaneous metastasis dissemination of breast cancer cells independently of cell proliferation and angiogenesis. Int J Oncol. 2012;40:1133–41.
Shannon P, Markiel A, Ozier O, Baliga N, Wang J, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Yevshin I, Sharipov L, Valeev T, Kel A, Kolpakov F. GTRD: a database of transcription factor binding sites identified by ChIP-seq experiments. Nucleic Acids Res. 2017;45:D61–7.
Delliaux C, Tian T, Bouchet M, Fradet A, Vanpouille N, Flourens A, et al. TMPRSS2:ERG gene fusion expression regulates bone markers and enhances the osteoblastic phenotype of prostate cancer bone metastases. Cancer Lett. 2018;438:32–43.
Acknowledgements
The authors thank Anne Flourens, Cyprien Tilmant, Tina Louadj, and both CeCIL and ALECS platforms (Faculté de Médecine Laennec, Lyon) for their assistance.
Funding
This work was supported by the National Center for Scientific Research (CNRS) to EB, the National Institute of Health and Medical Research (INSERM), the University of Lyon1, La Ligue Nationale (Drôme), Inserm-Transfert (EB). GV was supported by the Labex DEVweCAN, MG, MB by the French National Cancer Institute (INCa), CK by the Marie-Curie-Individual-Fellowship (655777-miROMeS). CAP, LC, and MM by CANCER-ID (FP7/2007-2013) and EFPIA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Vargas, G., Bouchet, M., Bouazza, L. et al. ERRα promotes breast cancer cell dissemination to bone by increasing RANK expression in primary breast tumors. Oncogene 38, 950–964 (2019). https://doi.org/10.1038/s41388-018-0579-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0579-3
This article is cited by
-
ERRα: unraveling its role as a key player in cell migration
Oncogene (2024)
-
The future of clinical trials—goals, ideas, and discussion
memo - Magazine of European Medical Oncology (2024)
-
Estrogen-related receptor alpha in select host functions and cancer: new frontiers
Molecular and Cellular Biochemistry (2022)
-
ERRα expression in ovarian cancer and promotes ovarian cancer cells migration in vitro
Archives of Gynecology and Obstetrics (2022)
-
Dual targeting of SREBP2 and ERRα by carnosic acid suppresses RANKL-mediated osteoclastogenesis and prevents ovariectomy-induced bone loss
Cell Death & Differentiation (2020)