Abstract
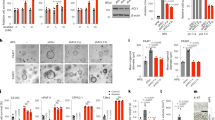
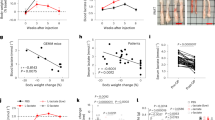
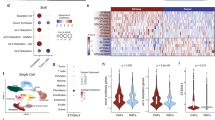
Interleukin-8 (IL-8) is a pro-inflammatory chemokine that is associated with induction of chemotaxis and degranulation of neutrophils. IL-8 is overexpressed in many tumors, including colon and lung cancer, and recent studies demonstrated essential roles for IL-8 in tumor progression within the tumor microenvironment. However, the molecular mechanism underlying the functions of IL-8 in tumor progression is unclear. In this study, we found that IL-8 is overexpressed in colon and lung cancer cells with cancer stem cell (CSC)-like characteristics and is required for CSC properties, including tumor-initiating abilities. These findings suggest that IL-8 plays an essential role in the development of CSCs. We also showed that IL-8 stimulation of colon and lung cancer cells-induced glucose uptake and expressions of glucose transporter 3 (GLUT3) and glucosamine fructose-6-phosphate aminotransferase (GFAT), a regulator of glucose flux to the hexosamine biosynthetic pathway, resulting in enhancement of protein O-GlcNAcylation. We demonstrated that these events are required for the generation and maintenance CSC-like characteristics of colon and lung cancer cells. Moreover, an O-GlcNAcylation inhibitor, OSMI1, reduced CSC number and tumor development in vivo. Together, these results reveal that IL-8-induced O-GlcNAcylation is required for generation and maintenance of CSCs of colon and lung cancer cells and suggests this regulatory pathway as a candidate therapeutic target of CSCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Ha H, Debnath B, Neamati N. Role of the CXCL8-CXCR1/2 axis in cancer and inflammatory diseases. Theranostics. 2017;7:1543–88.
Waugh DJ, Wilson C. The interleukin-8 pathway in cancer. Clin Cancer Res. 2008;14:6735–41.
Shahzad A, Knapp M, Lang I, Kohler G. Interleukin 8 (IL-8) - a universal biomarker? Int Arch Med. 2010;3:11.
Standiford TJ, Kunkel SL, Basha MA, Chensue SW, Lynch JP 3rd, Toews GB, et al. Interleukin-8 gene expression by a pulmonary epithelial cell line. A model for cytokine networks in the lung. J Clin Invest. 1990;86:1945–53.
Liu Q, Li A, Tian Y, Wu JD, Liu Y, Li T, et al. The CXCL8-CXCR1/2 pathways in cancer. Cytokine Growth Factor Rev. 2016;31:61–71.
Alfaro C, Sanmamed MF, Rodriguez-Ruiz ME, Teijeira A, Onate C, Gonzalez A, et al. Interleukin-8 in cancer pathogenesis, treatment and follow-up. Cancer Treat Rev. 2017;60:24–31.
David JM, Dominguez C, Hamilton DH, Palena C.The IL-8/IL-8R axis: a double agent in tumor immune resistance. Vaccines. 2016;4:22. https://doi.org/10.3390/vaccines4030022
Liu Q, Li A, Yu S, Qin S, Han N, Pestell RG, et al. DACH1 antagonizes CXCL8 to repress tumorigenesis of lung adenocarcinoma and improve prognosis. J Hematol Oncol. 2018;11:53.
Shi L, Wang L, Wang B, Cretoiu SM, Wang Q, Wang X, et al. Regulatory mechanisms of betacellulin in CXCL8 production from lung cancer cells. J Transl Med. 2014;12:70.
Wang J, Wang Y, Wang S, Cai J, Shi J, Sui X, et al. Bone marrow-derived mesenchymal stem cell-secreted IL-8 promotes the angiogenesis and growth of colorectal cancer. Oncotarget. 2015;6:42825–37.
Singh S, Wu S, Varney M, Singh AP, Singh RK. CXCR1 and CXCR2 silencing modulates CXCL8-dependent endothelial cell proliferation, migration and capillary-like structure formation. Microvasc Res. 2011;82:318–25.
Visciano C, Liotti F, Prevete N, Cali G, Franco R, Collina F, et al. Mast cells induce epithelial-to-mesenchymal transition and stem cell features in human thyroid cancer cells through an IL-8-Akt-Slug pathway. Oncogene. 2015;34:5175–86.
Hwang WL, Yang MH, Tsai ML, Lan HY, Su SH, Chang SC, et al. SNAIL regulates interleukin-8 expression, stem cell-like activity, and tumorigenicity of human colorectal carcinoma cells. Gastroenterology. 2011;141:279–91. 291 e271-275
Batlle E, Clevers H. Cancer stem cells revisited. Nat Med. 2017;23:1124–34.
Visvader JE. Cells of origin in cancer. Nature. 2011;469:314–22.
Arwert EN, Hoste E, Watt FM. Epithelial stem cells, wound healing and cancer. Nat Rev Cancer. 2012;12:170–80.
Schafer M, Werner S. Cancer as an overhealing wound: an old hypothesis revisited. Nat Rev Mol Cell Biol. 2008;9:628–38.
Zhou BB, Zhang H, Damelin M, Geles KG, Grindley JC, Dirks PB. Tumour-initiating cells: challenges and opportunities for anticancer drug discovery. Nat Rev Drug Discov. 2009;8:806–23.
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15:178–96.
Eramo A, Lotti F, Sette G, Pilozzi E, Biffoni M, Di Virgilio A, et al. Identification and expansion of the tumorigenic lung cancer stem cell population. Cell Death Differ. 2008;15:504–14.
Vermeulen L, Todaro M, de Sousa Mello F, Sprick MR, Kemper K, Perez Alea M, et al. Single-cell cloning of colon cancer stem cells reveals a multi-lineage differentiation capacity. Proc Natl Acad Sci USA. 2008;105:13427–32.
Corbet C. Stem cell metabolism in cancer and healthy tissues: pyruvate in the limelight. Front Pharmacol. 2017;8:958.
Kawauchi K, Araki K, Tobiume K, Tanaka N. p53 regulates glucose metabolism through an IKK-NF-kappaB pathway and inhibits cell transformation. Nat Cell Biol. 2008;10:611–8.
Kuang R, Jahangiri A, Mascharak S, Nguyen A, Chandra A, Flanigan PM, et al. GLUT3 upregulation promotes metabolic reprogramming associated with antiangiogenic therapy resistance. JCI Insight. 2017;2:e88815.
Mo L, Chen Q, Yang Y, Rui XP, Gu J. High expression of GLUT1 and GLUT3 correlate with neoadjuvant chemotherapy ineffectiveness breast cancer patients. Int J Clin Exp Pathol. 2016;9:9555–61.
Zhang Y, Wei C, Xi J, Tang Z, Liang C. Glucose transporter 3 performs a critical role in mTOR-mediated oncogenic glycolysis and tumorigenesis. Oncol Lett. 2015;9:2809–14.
Rodrigues-Ferreira C, da Silva AP, Galina A. Effect of the antitumoral alkylating agent 3-bromopyruvate on mitochondrial respiration: role of mitochondrially bound hexokinase. J Bioenerg Biomembr. 2012;44:39–49.
Nieman KM, Kenny HA, Penicka CV, Ladanyi A, Buell-Gutbrod R, Zillhardt MR, et al. Adipocytes promote ovarian cancer metastasis and provide energy for rapid tumor growth. Nat Med. 2011;17:1498–503.
Hardiville S, Hart GW. Nutrient regulation of signaling, transcription, and cell physiology by O-GlcNAcylation. Cell Metab. 2014;20:208–13.
Yang X, Qian K. Protein O-GlcNAcylation: emerging mechanisms and functions. Nat Rev Mol Cell Biol. 2017;18:452–65.
Oikari S, Makkonen K, Deen AJ, Tyni I, Karna R, Tammi RH, et al. Hexosamine biosynthesis in keratinocytes: roles of GFAT and GNPDA enzymes in the maintenance of UDP-GlcNAc content and hyaluronan synthesis. Glycobiology. 2016;26:710–22.
Dong T, Kang X, Liu Z, Zhao S, Ma W, Xuan Q, et al. Altered glycometabolism affects both clinical features and prognosis of triple-negative and neoadjuvant chemotherapy-treated breast cancer. Tumour Biol. 2016;37:8159–68.
Li L, Shao M, Peng P, Yang C, Song S, Duan F, et al. High expression of GFAT1 predicts unfavorable prognosis in patients with hepatocellular carcinoma. Oncotarget. 2017;8:19205–17.
King JB, West MB, Cook PF, Hanigan MH. A novel, species-specific class of uncompetitive inhibitors of gamma-glutamyl transpeptidase. J Biol Chem. 2009;284:9059–65.
Guo H, Zhang B, Nairn AV, Nagy T, Moremen KW, Buckhaults P, et al. O-linked N-acetylglucosamine (O-GlcNAc) expression levels epigenetically regulate colon cancer tumorigenesis by affecting the cancer stem cell compartment via modulating expression of transcriptional factor MYBL1. J Biol Chem. 2017;292:4123–37.
Mi W, Gu Y, Han C, Liu H, Fan Q, Zhang X, et al. O-GlcNAcylation is a novel regulator of lung and colon cancer malignancy. Biochim Biophys Acta. 2011;1812:514–9.
Singh JP, Zhang K, Wu J, Yang X. O-GlcNAc signaling in cancer metabolism and epigenetics. Cancer Lett. 2015;356:244–50.
Slawson C, Hart GW. O-GlcNAc signalling: implications for cancer cell biology. Nat Rev Cancer. 2011;11:678–84.
Beck B, Blanpain C. Unravelling cancer stem cell potential. Nat Rev Cancer. 2013;13:727–38.
Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331:1559–64.
Yuzwa SA, Macauley MS, Heinonen JE, Shan X, Dennis RJ, He Y, et al. A potent mechanism-inspired O-GlcNAcase inhibitor that blocks phosphorylation of tau in vivo. Nat Chem Biol. 2008;4:483–90.
Liu BQ, Meng X, Li C, Gao YY, Li N, Niu XF, et al. Glucosamine induces cell death via proteasome inhibition in human ALVA41 prostate cancer cell. Exp Mol Med. 2011;43:487–93.
Ortiz-Meoz RF, Jiang J, Lazarus MB, Orman M, Janetzko J, Fan C, et al. A small molecule that inhibits OGT activity in cells. ACS Chem Biol. 2015;10:1392–7.
Caldwell SA, Jackson SR, Shahriari KS, Lynch TP, Sethi G, Walker S, et al. Nutrient sensor O-GlcNAc transferase regulates breast cancer tumorigenesis through targeting of the oncogenic transcription factor FoxM1. Oncogene. 2010;29:2831–42.
Xiang R, Liao D, Cheng T, Zhou H, Shi Q, Chuang TS, et al. Downregulation of transcription factor SOX2 in cancer stem cells suppresses growth and metastasis of lung cancer. Br J Cancer. 2011;104:1410–7.
Zheng J, Xu L, Pan Y, Yu S, Wang H, Kennedy D, et al. Sox2 modulates motility and enhances progression of colorectal cancer via the Rho-ROCK signaling pathway. Oncotarget. 2017;8:98635–45.
Chu CS, Lo PW, Yeh YH, Hsu PH, Peng SH, Teng YC, et al. O-GlcNAcylation regulates EZH2 protein stability and function. Proc Natl Acad Sci USA. 2014;111:1355–60.
Jang H, Kim TW, Yoon S, Choi SY, Kang TW, Kim SY, et al. O-GlcNAc regulates pluripotency and reprogramming by directly acting on core components of the pluripotency network. Cell Stem Cell. 2012;11:62–74.
Allison DF, Wamsley JJ, Kumar M, Li D, Gray LG, Hart GW, et al. Modification of RelA by O-linked N-acetylglucosamine links glucose metabolism to NF-kappaB acetylation and transcription. Proc Natl Acad Sci USA. 2012;109:16888–93.
Ma Z, Vocadlo DJ, Vosseller K. Hyper-O-GlcNAcylation is anti-apoptotic and maintains constitutive NF-kappaB activity in pancreatic cancer cells. J Biol Chem. 2013;288:15121–30.
Gloster TM, Zandberg WF, Heinonen JE, Shen DL, Deng L, Vocadlo DJ. Hijacking a biosynthetic pathway yields a glycosyltransferase inhibitor within cells. Nat Chem Biol. 2011;7:174–81.
Bareiss PM, Paczulla A, Wang H, Schairer R, Wiehr S, Kohlhofer U, et al. SOX2 expression associates with stem cell state in human ovarian carcinoma. Cancer Res. 2013;73:5544–55.
Nakajima W, Sharma K, Lee JY, Maxim NT, Hicks MA, Vu TT, et al. DNA damaging agent-induced apoptosis is regulated by MCL-1 phosphorylation and degradation mediated by the Noxa/MCL-1/CDK2 complex. Oncotarget. 2016;7:36353–65.
Aguirre-Gamboa R, Gomez-Rueda H, Martinez-Ledesma E, Martinez-Torteya A, Chacolla-Huaringa R, Rodriguez-Barrientos A, et al. SurvExpress: an online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS ONE. 2013;8:e74250.
Acknowledgements
We thank Y. Abe. Y. Nakajima, A. Tanimura, and I. Uehara for discussion, and Y. Asano, M. Kawagoe and T. Takatera for technical support. This work was supported by Grants-in-Aid for scientific research from the Ministry of Education, Science, Sports, and Culture of Japan (MEXT/JSPS KAKEN Grant Number 17H04554). We thank Edanz Group (www.edanzediting.com/ac) for editing a draft of this manuscript.
Author contributions
M.S. and N.T. designed the study, analyzed the data, and wrote the manuscript. M.S. performed the experiments, and M.S. and N.T. analyzed the data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Shimizu, M., Tanaka, N. IL-8-induced O-GlcNAc modification via GLUT3 and GFAT regulates cancer stem cell-like properties in colon and lung cancer cells. Oncogene 38, 1520–1533 (2019). https://doi.org/10.1038/s41388-018-0533-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0533-4
This article is cited by
-
Current knowledge and potential intervention of hexosamine biosynthesis pathway in lung cancer
World Journal of Surgical Oncology (2023)
-
Loss of miR-637 promotes cancer cell stemness via WASH/IL-8 pathway and serves as a novel prognostic marker in esophageal squamous cell carcinoma
Biomarker Research (2022)
-
O-GlcNAcylation of MORC2 at threonine 556 by OGT couples TGF-β signaling to breast cancer progression
Cell Death & Differentiation (2022)
-
Enhanced O-GlcNAc modification induced by the RAS/MAPK/CDK1 pathway is required for SOX2 protein expression and generation of cancer stem cells
Scientific Reports (2022)
-
Deduction and exploration of the evolution and function of vertebrate GFPT family
Genes & Genomics (2022)