Abstract
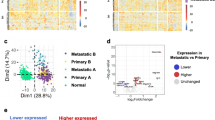
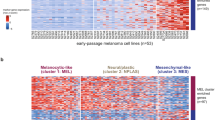
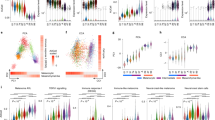
Recent studies revealed trajectories of mutational events in early melanomagenesis, but the accompanying changes in gene expression are far less understood. Therefore, we performed a comprehensive RNA-seq analysis of laser-microdissected melanocytic nevi (n = 23) and primary melanoma samples (n = 57) and characterized the molecular mechanisms of early melanoma development. Using self-organizing maps, unsupervised clustering, and analysis of pseudotime (PT) dynamics to identify evolutionary trajectories, we describe here two transcriptomic types of melanocytic nevi (N1 and N2) and primary melanomas (M1 and M2). N1/M1 lesions are characterized by pigmentation-type and MITF gene signatures, and a high prevalence of NRAS mutations in M1 melanomas. N2/M2 lesions are characterized by inflammatory-type and AXL gene signatures with an equal distribution of wild-type and mutated BRAF and low prevalence of NRAS mutations in M2 melanomas. Interestingly, N1 nevi and M1 melanomas and N2 nevi and M2 melanomas, respectively, cluster together, but there is no clustering in a stage-dependent manner. Transcriptional signatures of M1 melanomas harbor signatures of BRAF/MEK inhibitor resistance and M2 melanomas harbor signatures of anti-PD-1 antibody treatment resistance. Pseudotime dynamics of nevus and melanoma samples are suggestive for a switch-like immune-escape mechanism in melanoma development with downregulation of immune genes paralleled by an increasing expression of a cell cycle signature in late-stage melanomas. Taken together, the transcriptome analysis identifies gene signatures and mechanisms underlying development of melanoma in early and late stages with relevance for diagnostics and therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell. 2015;161:1681–96.
Solus JF, Kraft S. Ras, Raf, and MAP kinase in melanoma. Adv Anat Pathol. 2013;20:217–26.
Schadendorf D, Fisher DE, Garbe C, Gershenwald JE, Grob JJ, Halpern A, et al. Melanoma. Nat Rev Dis Prim. 2015;1:1–20.
Sullivan RJ, Flaherty KT. New strategies in melanoma: entering the era of combinatorial therapy. Clin Cancer Res. 2015;21:2424–35.
Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A, et al. The genetic evolution of melanoma from precursor lesions. N Engl J Med. 2015;373:1926–36.
Winnepenninckx V, Lazar V, Michiels S, Dessen P, Stas M, Alonso SR, et al. Gene expression profiling of primary cutaneous melanoma and clinical outcome. J Natl Cancer Inst. 2006;98:472–82.
Jönsson G, Busch C, Knappskog S, Geisler J, Miletic H, Ringnér M, et al. Gene expression profiling-based identification of molecular subtypes in stage IV melanomas with different clinical outcome. Clin Cancer Res. 2010;16:3356–67.
Harbst K, Staaf J, Lauss M, Karlsson A, Måsbäck A, Johansson I, et al. Molecular profiling reveals low- and high-grade forms of primary melanoma. Clin Cancer Res. 2012;18:4026–36.
Cirenajwis H, Ekedahl H, Lauss M, Harbst K, Carneiro A, Enoksson J, et al. Molecular stratification of metastatic melanoma using gene expression profiling: prediction of survival outcome and benefit from molecular targeted therapy. Oncotarget. 2015;6:12297–309.
Bradner JE, Hnisz D, Young RA. Transcriptional addiction in cancer. Cell. 2017;168:629–43.
Konieczkowski DJ, Johannessen CM, Abudayyeh O, Kim JW, Cooper ZA, Piris A, et al. A melanoma cell state distinction influences sensitivity to MAPK pathway inhibitors. Cancer Discov. 2014;4:816–27.
Müller J, Krijgsman O, Tsoi J, Robert L, Hugo W, Song C, et al. Low MITF/AXL ratio predicts early resistance to multiple targeted drugs in melanoma. Nat Commun. 2014;5:5712.
Breslow A. Thickness, cross-sectional areas and depth of invasion in the prognosis of cutaneous melanoma. Ann Surg. 1970;172:902–8.
Kauffmann A, Rosselli F, Lazar V, Winnepenninckx V, Mansuet-Lupo A, Dessen P, et al. High expression of DNA repair pathways is associated with metastasis in melanoma patients. Oncogene. 2008;27:565–73.
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database Hallmark gene set collection. Cell Syst. 2015;1:417–25.
Tirosh I, Izar B, Prakadan SM, Wadsworth MH 2nd, Treacy D, Trombetta JJ, et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science. 2016;352:189–96.
Hugo W, Shi H, Sun L, Piva M, Song C, Kong X, et al. Non-genomic and immune evolution of melanoma acquiring MAPKi resistance. Cell. 2015;162:1271–85.
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell. 2016;165:35–44.
Wirth H, Löffler M, von Bergen M, Binder H. Expression cartography of human tissues using self organizing maps. BMC Bioinforma. 2011;12:306.
Binder H, Wirth H. Analysis of large-scale OMIC data using self organizing maps. In: Khosrow-Pour M (ed). Encyclopedia of Information Science and Technology, 3 rd edn. IGI Global: Hershey, PA, USA, 2014, pp 1642–1654.
Camp JG, Sekine K, Gerber T, Loeffler-Wirth H, Binder H, Gac M, et al. Multilineage communication regulates human liver bud development from pluripotency. Nature. 2017;546:533–8.
Haqq C, Nosrati M, Sudilovsky D, Crothers J, Khodabakhsh D, Pulliam BL, et al. The gene expression signatures of melanoma progression. Proc Natl Acad Sci USA. 2005;102:6092–7.
Roadmap Epigenomics Consortium, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30.
Keenen B, Qi H, Saladi SV, Yeung M, de la Serna IL. Heterogeneous SWI/SNF chromatin remodeling complexes promote expression of microphthalmia-associated transcription factor target genes in melanoma. Oncogene. 2010;29:81–92.
Lin H, Wong RP, Martinka M, Li G. BRG1 expression is increased in human cutaneous melanoma. Br J Dermatol. 2010;163:502–10.
Clarke LE, Warf BM, Flake DD 2nd, Hartman AR, Tahan S, Shea CR, et al. Clinical validation of a gene expression signature that differentiates benign nevi from malignant melanoma. J Cutan Pathol. 2015;42:244–52.
Clarke LE, Flake DD 2nd, Busam K, Cockerell C, Helm K, McNiff J, et al. An independent validation of a gene expression signature to differentiate malignant melanoma from benign melanocytic nevi. Cancer. 2017;123:617–28.
Hulstaert E, Brochez L, Volders PJ, Vandesompele J, Mestdagh P. Long non-coding RNAs in cutaneous melanoma: clinical perspectives. Oncotarget. 2017;8:43470–80.
Bendall SC, Davis KL, Amir el-AD, Tadmor MD, Simonds EF, Chen TJ, et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell. 2014;157:714–25.
Bindea G, Mlecnik B, Tosolini M, Kirilovsky A, Waldner M, Obenauf AC, et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity. 2013;39:782–95.
Angelova M, Charoentong P, Hackl H, Fischer ML, Snajder R, Krogsdam AM, et al. Characterization of the immunophenotypes and antigenomes of colorectal cancers reveals distinct tumor escape mechanisms and novel targets for immunotherapy. Genome Biol. 2015;16:64.
Gerber T, Willscher E, Loeffler-Wirth H, Hopp L, Schadendorf D, Schartl M, et al. Mapping heterogeneity in patient-derived melanoma cultures by single-cell RNA-seq. Oncotarget. 2017;8:846–62.
Riesenberg S, Groetchen A, Siddaway R, Bald T, Reinhardt J, Smorra D, et al. MITF and c-Jun antagonism interconnects melanoma dedifferentiation with pro-inflammatory cytokine responsiveness and myeloid cell recruitment. Nat Commun. 2015;6:8755.
Maurus K, Hufnagel A, Geiger F, Graf S, Berking C, Heinemann A, et al. The AP-1 transcription factor FOSL1 causes melanocyte reprogramming and transformation. Oncogene. 2017;36:5110–21.
Riaz N, Havel JJ, Makarov V, Desrichard A, Urba WJ, Sims JS, et al. Tumor and Microenvironment Evolution during Immunotherapy with Nivolumab. Cell. 2017;171:934–49.
Chen PL, Roh W, Reuben A, Cooper ZA, Spencer CN, Prieto PA, et al. Analysis of immune signatures in longitudinal tumor samples yields insight into biomarkers of response and mechanisms of resistance to immune checkpoint blockade. Cancer Discov. 2016;6:827–37.
Haymaker C, Wu R, Bernatchez C, Radvanyi L. PD-1 and BTLA and CD8(+) T-cell "exhaustion" in cancer: "exercising" an alternative viewpoint. Oncoimmunology. 2012;1:735–8.
Saladi SV, Keenen B, Marathe HG, Qi H, Chin KV, de la Serna IL. Modulation of extracellular matrix/adhesion molecule expression by BRG1 is associated with increased melanoma invasiveness. Mol Cancer. 2010;9:280.
Campbell KR, Yau C. Order under uncertainty: robust differential expression analysis using probabilistic models for pseudotime inference. PLoS Comput Biol. 2016;12:e1005212.
Reid JE, Wernisch L. Pseudotime estimation: deconfounding single cell time series. Bioinformatics. 2016;32:2973–80.
Landsberg J, Kohlmeyer J, Renn M, Bald T, Rogava M, Cron M, et al. Melanomas resist T-cell therapy through inflammation-induced reversible dedifferentiation. Nature. 2012;490:412–6.
Jaeger J, Koczan D, Thiesen HJ, Ibrahim SM, Gross G, Spang R, et al. Gene expression signatures for tumor progression, tumor subtype, and tumor thickness in laser-microdissected melanoma tissues. Clin Cancer Res. 2007;13:806–15.
Wu TD, Nacu S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics. 2010;26:873–81.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Wirth H, von Bergen M, Binder H. Mining SOM expression portraits: feature selection and integrating concepts of molecular function. BioData Min. 2012;5:18.
Löffler-Wirth H, Kalcher M, Binder H. oposSOM: R-package for high-dimensional portraying of genome-wide expression landscapes on bioconductor. Bioinformatics. 2015;31:3225–7.
Törönen P, Ojala P, Marttinen P, Holm L. Robust extraction of functional signals from gene set analysis using a generalized threshold free scoring function. BMC Bioinforma. 2009;10:307.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Hopp L, Wirth H, Fasold M, Binder H. Portraying the expression landscapes of cancer subtypes: a glioblastoma multiforme and prostate cancer case study. Syst Biomed. 2013;1:99–121.
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet. 2008;40:499–507.
Shen H, Powers N, Saini N, Comstock CE, Sharma A, Weaver K, et al. The SWI/SNF ATPase Brm is a gatekeeper of proliferative control in prostate cancer. Cancer Res. 2008;68:10154–62.
Acknowledgements
MK, MD, AB, SM, and MS were supported by funding of the Deutsche Krebshilfe, Melanomverbund, grant number 109716.
Author contributions
MK, HL-W, MD, EW, GD, JK, TK, BN, JL, PZ, CM, HJS, LH, HB, SM, AB, and MS conceived and designed the work that led to the submission, acquired data, and played an important role in interpreting the results and approved the final version. SH, TT, JU, MZ, SK, MH, and AR supported the interpretation of the results and approved the final version.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
MK has received honoraria from the Speakers Bureau of Roche Pharma AG and travel support from Novartis Pharma GmbH and Bristol-Myers Squibb GmbH (BMS). MZ has received speakers honoraria and honoraria for Medical Advisory Boards from BMS, Roche Pharma AG, MSD Sharp & Dohme GmbH (MSD), Amgen GmbH, Novartis Pharma GmbH, Pfizer Pharma GmbH, and Merck Serono GmbH as well as financial support for travel support from BMS and Amgen GmbH. TT has received honoraria from the Speakers Bureau of Roche Pharma AG and travel support from Novartis Pharma GmbH and BMS. CM has received travel support from MSD and Pfizer GmbH and Roche Pharma AG and received honoraria from Novartis Pharma GmbH. JU is on the advisory board or has received honoraria and travel support from Amgen, BMS, GlaxoSmithKline GmbH &nCo, LeoPharma, MSD, Novartis, Roche Pharma AG. AR received travel grants and honoraria from Roche Pharma AG, TEVA, MMS, MSD, Amgen, and Novartis Pharma GmbH and a research grant from Novartis Pharma GmbH. JL, HL-W, MD, EW, GD, JK, LH, SH, PZ, HJS, MH, SK, SM, AB, HB and MS declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kunz, M., Löffler-Wirth, H., Dannemann, M. et al. RNA-seq analysis identifies different transcriptomic types and developmental trajectories of primary melanomas. Oncogene 37, 6136–6151 (2018). https://doi.org/10.1038/s41388-018-0385-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0385-y
This article is cited by
-
NRN1 interacts with Notch to increase oncogenic STAT3 signaling in melanoma
Cell Communication and Signaling (2024)
-
Unsupervised analysis of whole transcriptome data from human pluripotent stem cells cardiac differentiation
Scientific Reports (2024)
-
Functional analysis of recurrent CDC20 promoter variants in human melanoma
Communications Biology (2023)
-
Scalable transcriptomics analysis with Dask: applications in data science and machine learning
BMC Bioinformatics (2022)
-
Transcriptome profile of the sinoatrial ring reveals conserved and novel genetic programs of the zebrafish pacemaker
BMC Genomics (2021)