Abstract
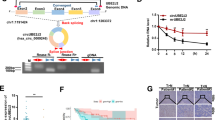
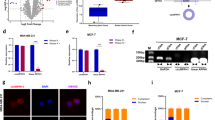
Circular RNAs are a large group of noncoding RNAs that are widely expressed in mammalian cells. Genome-wide analyses have revealed abundant and evolutionarily conserved circular RNAs across species, which suggest specific physiological roles of these species. Using a microarray approach, we detected increased expression of a circular RNA circ-Dnmt1 in eight breast cancer cell lines and in patients with breast carcinoma. Silencing circ-Dnmt1 inhibited cell proliferation and survival. Ectopic circ-Dnmt1 increased the proliferative and survival capacities of breast cancer cells by stimulating cellular autophagy. We found that circ-Dnmt1-mediated autophagy was essential in inhibiting cellular senescence and increasing tumor xenograft growth. We further found that ectopically expressed circ-Dnmt1 could interact with both p53 and AUF1, promoting the nuclear translocation of both proteins. Nuclear translocation of p53 induced cellular autophagy while AUF1 nuclear translocation reduced Dnmt1 mRNA instability, resulting in increased Dnmt1 translation. From here, functional Dnmt1 could then translocate into the nucleus, inhibiting p53 transcription. Computational algorithms revealed that both p53 and AUF1 could bind to different regions of circ-Dnmt1 RNA. Our results showed that the highly expressed circular RNA circ-Dnmt1 could bind to and regulate oncogenic proteins in breast cancer cells. Thus circ-Dnmt1 appears to be an oncogenic circular RNA with potential for further preclinical research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Enuka Y, Lauriola M, Feldman ME, Sas-Chen A, Ulitsky I, Yarden Y. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016;44:1370–83.
Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014;32:453–61.
Liu YC, Li JR, Sun CH, Andrews E, Chao RF, Lin FM, et al. CircNet: a database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016;44:D209–215.
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–8.
Zheng LL, Li JH, Wu J, Sun WJ, Liu S, Wang ZL, et al. deepBasev2.0: identification, expression, evolution and function of small RNAs, LncRNAs and circular RNAs from deep-sequencing data. Nucleic Acids Res. 2016;44:D196–202.
Zhang Y, Zhang XO, Chen T, Xiang JF, Yin QF, Xing YH, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51:792–806.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–64.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE. 2012;7:e30733.
Panda AC, De S, Grammatikakis I, Munk R, Yang X, Piao Y, et al. High-purity circular RNA isolation method (RPAD) reveals vast collection of intronic circRNAs. Nucleic Acids Res. 2017;45:e116.
Bai Y, Zhang Y, Han B, Yang L, Chen X, Huang R, et al. Circular RNA DLGAP4 ameliorates ischemic stroke outcomes by targeting miR-143 to regulate endothelial-mesenchymal transition associated with blood-brain barrier integrity. J Neurosci. 2017;38:32–50.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Huang R, Zhang Y, Han B, Bai Y, Zhou R, Gan G, et al. Circular RNA HIPK2 regulates astrocyte activation via cooperation of autophagy and ER stress by targeting MIR124-2HG. Autophagy. 2017;13:1722–41.
Hsiao KY, Lin YC, Gupta SK, Chang N, Yen L, Sun HS, et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017;77:2339–50.
Yang W, Du WW, Li X, Yee AJ, Yang BB. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene. 2016;35:3919–31.
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. J Natl Cancer Inst. 2018;110, https://doi.org/10.1093/jnci/djx166.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–30.
Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–58.
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–70.
Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, et al. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38:1402–12.
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010;6:e1001233.
Hansen TB, Kjems J, Damgaard CK. Circular RNA and miR-7 in cancer. Cancer Res. 2013;73:5609–12.
Yoshida Y, Izumi H, Torigoe T, Ishiguchi H, Yoshida T, Itoh H, et al. Binding of RNA to p53 regulates its oligomerization and DNA-binding activity. Oncogene. 2004;23:4371–9.
Nagata T, Kurihara Y, Matsuda G, Saeki J, Kohno T, Yanagida Y, et al. Structure and interactions with RNA of the N-terminal UUAG-specific RNA-binding domain of hnRNP D0. J Mol Biol. 1999;287:221–37.
Katahira M, Miyanoiri Y, Enokizono Y, Matsuda G, Nagata T, Ishikawa F, et al. Structure of the C-terminal RNA-binding domain of hnRNP D0 (AUF1), its interactions with RNA and DNA, and change in backbone dynamics upon complex formation with DNA. J Mol Biol. 2001;311:973–88.
Chen J, Sawyer N, Regan L. Protein-protein interactions: general trends in the relationship between binding affinity and interfacial buried surface area. Protein Sci. 2013;22:510–5.
Kastritis PL, Bonvin AM. On the binding affinity of macromolecular interactions: daring to ask why proteins interact. J R Soc Interface. 2013;10:20120835.
Yen RW, Vertino PM, Nelkin BD, Yu JJ, el-Deiry W, Cumaraswamy A, et al. Isolation and characterization of the cDNA encoding human DNA methyltransferase. Nucleic Acids Res. 1992;20:2287–91.
Klein CJ, Botuyan MV, Wu Y, Ward CJ, Nicholson GA, Hammans S, et al. Mutations in DNMT1 cause hereditary sensory neuropathy with dementia and hearing loss. Nat Genet. 2011;43:595–600.
Georgia S, Kanji M, Bhushan A. DNMT1 represses p53 to maintain progenitor cell survival during pancreatic organogenesis. Genes Dev. 2013;27:372–7.
Tasdemir E, Chiara Maiuri M, Morselli E, Criollo A, D’Amelio M, Djavaheri-Mergny M, et al. A dual role of p53 in the control of autophagy. Autophagy. 2008;4:810–4.
Tasdemir E, Maiuri MC, Galluzzi L, Vitale I, Djavaheri-Mergny M, D’Amelio M, et al. Regulation of autophagy by cytoplasmic p53. Nat Cell Biol. 2008;10:676–87.
Miller SJ, Suthiphongchai T, Zambetti GP, Ewen ME. p53 binds selectively to the 5′ untranslated region of cdk4, an RNA element necessary and sufficient for transforming growth factor beta- and p53-mediated translational inhibition of cdk4. Mol Cell Biol. 2000;20:8420–31.
Dempsey LA, Li MJ, DePace A, Bray-Ward P, Maizels N. The human HNRPD locus maps to 4q21 and encodes a highly conserved protein. Genomics. 1998;49:378–84.
Torrisani J, Unterberger A, Tendulkar SR, Shikimi K, Szyf M. AUF1 cell cycle variations define genomic DNA methylation by regulation of DNMT1 mRNA stability. Mol Cell Biol. 2007;27:395–410.
Gratacos FM, Brewer G. The role of AUF1 in regulated mRNA decay. Wiley Interdiscip Rev RNA. 2010;1:457–73.
Washietl S, Will S, Hendrix DA, Goff LA, Rinn JL, Berger B, et al. Computational analysis of noncoding RNAs. Wiley Interdiscip Rev RNA. 2012;3:759–78.
Tuszynska I, Magnus M, Jonak K, Dawson W, Bujnicki JM. NPDock: a web server for protein-nucleic acid docking. Nucleic Acids Res. 2015;43:W425–430.
Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, et al. Correlation of circular RNA abundance with proliferation--exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015;5:8057.
Yang Q, Du WW, Wu N, Yang W, Awan FM, Fang L, et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 2017;24:1609–20.
Rutnam ZJ, Du WW, Yang W, Yang X, Yang BB. The pseudogene TUSC2P promotes TUSC2 function by binding multiple microRNAs. Nat Commun. 2014;5:2914.
Yang X, Du WW, Li H, Liu F, Khorshidi A, Rutnam ZJ, et al. Both mature miR-17-5p and passenger strand miR-17-3p target TIMP3 and induce prostate tumor growth and invasion. Nucleic Acids Res. 2013;41:9688–704.
Shan SW, Lee DY, Deng Z, Shatseva T, Jeyapalan Z, Du WW, et al. MicroRNA MiR-17 retards tissue growth and represses fibronectin expression. Nat Cell Biol. 2009;11:1031–8.
Pietal MJ, Szostak N, Rother KM, Bujnicki JM. RNAmap2D—calculation, visualization and analysis of contact and distance maps for RNA and protein-RNA complex structures. BMC Bioinforma. 2012;13:333.
Du WW, Liu F, Shan SW, Ma XC, Gupta S, Jin T, et al. Inhibition of dexamethasone-induced fatty liver development by reducing miR-17-5p levels. Mol Ther. 2015;23:1222–33.
Yang BL, Cao L, Kiani C, Lee V, Zhang Y, Adams ME, et al. Tandem repeats are involved in G1 domain inhibition of versican expression and secretion and the G3 domain enhances glycosaminoglycan modification and product secretion via the complement-binding protein-like motif. J Biol Chem. 2000;275:21255–61.
Zeng Y, Du WW, Wu Y, Yang Z, Awan FM, Li X, et al. A circular RNA binds to and activates AKT phosphorylation and nuclear localization reducing apoptosis and enhancing cardiac repair. Theranostics. 2017;7:3842–55.
Shan SW, Fang L, Shatseva T, Rutnam ZJ, Yang X, Du W, et al. Mature miR-17-5p and passenger miR-17-3p induce hepatocellular carcinoma by targeting PTEN, GalNT7 and vimentin in different signal pathways. J Cell Sci. 2013;126:1517–30.
Acknowledgements
This work was supported by grants from Canadian Institutes of Health Research (PJT-153105), National Natural Science Foundation of Guangdong (2017A030310088), and Innovation and Entrepreneurship Leading Talent of Guangzhou Development Zone (2017-L181).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
These authors contributed equally: William W. Du, Weining Yang, Xiangmin Li, Faryal Mehwish Awan, Zhenguo Yang.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Du, W.W., Yang, W., Li, X. et al. A circular RNA circ-DNMT1 enhances breast cancer progression by activating autophagy. Oncogene 37, 5829–5842 (2018). https://doi.org/10.1038/s41388-018-0369-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0369-y