Abstract
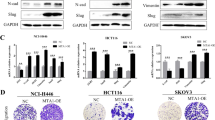
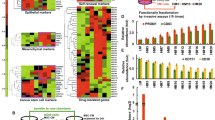
CDK8 is a transcription-regulating kinase that controls TGF-β/BMP-responsive SMAD transcriptional activation and turnover through YAP1 recruitment. However, how the CDK8/YAP1 pathway influences SMAD1 response in cancer remains unclear. Here we report that SMAD1-driven epithelial-to-mesenchymal transition (EMT) is critically dependent on matrix rigidity and YAP1 in a wide spectrum of cancer models. We find that both genetic and pharmacological inhibition of CDK8 and its homologous twin kinase CDK19 leads to abrogation of BMP-induced EMT. Notably, selectively blocking CDK8/19 specifically abrogates tumor cell invasion, changes in EMT-associated transcription factors, E-cadherin expression and YAP nuclear localization both in vitro and in vivo in a murine syngeneic EMT model. Furthermore, RNA-seq meta-analysis reveals a direct correlation between CDK8 and EMT-associated transcription factors in patients. Our findings demonstrate that CDK8, an emerging therapeutic target, coordinates growth factor and mechanical cues during EMT and invasion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hogan BL. Bone morphogenetic proteins: multifunctional regulators of vertebrate development. Genes Dev. 1996;10:1580–1594.
Richter A, Valdimarsdottir L, Hrafnkelsdottir HE, Runarsson JF, Omarsdottir AR, Ward-van Oostwaard D, et al. BMP4 promotes EMT and mesodermal commitment in human embryonic stem cells via SLUG and MSX2. Stem Cells. 2014;32:636–648.
Padgett RW, Wozney JM, Gelbart WM. Human BMP sequences can confer normal dorsal-ventral patterning in the Drosophila embryo. Proc Natl Acad Sci USA. 1993;90:2905–2909.
Chen M, Liang J, Ji H, Yang Z, Altilia S, Hu B, et al. CDK8/19 Mediator kinases potentiate induction of transcription by NFkappaB. Proc Natl Acad Sci USA. 2017;114:10208–10213.
Gordon KJ, Kirkbride KC, How T, Blobe GC. Bone morphogenetic proteins induce pancreatic cancer cell invasiveness through a Smad1-dependent mechanism that involves matrix metalloproteinase-2. Carcinogenesis. 2009;30:238–248.
Pickup MW, Owens P, Moses HL. TGF-beta, Bone Morphogenetic Protein, and Activin Signaling and the Tumor Microenvironment. Cold Spring Harbor perspectives in biology 2017; 9: a022285.
Theriault BL, Shepherd TG, Mujoomdar ML, Nachtigal MW. BMP4 induces EMT and Rho GTPase activation in human ovarian cancer cells. Carcinogenesis. 2007;28:1153–1162.
Yu PB, Hong CC, Sachidanandan C, Babitt JL, Deng DY, Hoyng SA, et al. Dorsomorphin inhibits BMP signals required for embryogenesis and iron metabolism. Nat Chem Biol. 2008;4:33–41.
Ampuja M, Alarmo EL, Owens P, Havunen R, Gorska AE, Moses HL, et al. The impact of bone morphogenetic protein 4 (BMP4) on breast cancer metastasis in a mouse xenograft model. Cancer Lett. 2016;375:238–244.
Hover LD, Owens P, Munden AL, Wang J, Chambless LB, Hopkins CR, et al. Bone morphogenetic protein signaling promotes tumorigenesis in a murine model of high-grade glioma. Neuro Oncol. 2016;18:928–938.
Owens P, Pickup MW, Novitskiy SV, Giltnane JM, Gorska AE, Hopkins CR, et al. Inhibition of BMP signaling suppresses metastasis in mammary cancer. Oncogene. 2015;34:2437–2449.
Le Page C, Puiffe ML, Meunier L, Zietarska M, de Ladurantaye M, Tonin PN, et al. BMP-2 signaling in ovarian cancer and its association with poor prognosis. J Ovarian Res. 2009;2:4.
Blanco Calvo M, Bolos Fernandez V, Medina Villaamil V, Aparicio Gallego G, Diaz Prado S, Grande Pulido E. Biology of BMP signalling and cancer. Clin Transl Oncol. 2009;11:126–137.
Yokoyama Y, Watanabe T, Tamura Y, Hashizume Y, Miyazono K, Ehata S. Autocrine BMP-4 signaling is a therapeutic target in colorectal cancer. Cancer Res. 2017;77:4026–4038.
Yamashita H, Ten Dijke P, Heldin CH, Miyazono K. Bone morphogenetic protein receptors. Bone. 1996;19:569–574.
Massague J, Wotton D. Transcriptional control by the TGF-beta/Smad signaling system. EMBO J. 2000;19:1745–1754.
Wrana JL, Attisano L, Carcamo J, Zentella A, Doody J, Laiho M, et al. TGF beta signals through a heteromeric protein kinase receptor complex. Cell. 1992;71:1003–1014.
Wrana JL, Attisano L, Wieser R, Ventura F, Massague J. Mechanism of activation of the TGF-beta receptor. Nature. 1994;370:341–347.
Alarcon C, Zaromytidou AI, Xi Q, Gao S, Yu J, Fujisawa S, et al. Nuclear CDKs drive Smad transcriptional activation and turnover in BMP and TGF-beta pathways. Cell. 2009;139:757–769.
Donner AJ, Ebmeier CC, Taatjes DJ, Espinosa JM. CDK8 is a positive regulator of transcriptional elongation within the serum response network. Nat Struct Mol Biol. 2010;17:194–201.
Galbraith MD, Donner AJ, Espinosa JM. CDK8: a positive regulator of transcription. Transcription. 2010;1:4–12.
Galbraith MD, Allen MA, Bensard CL, Wang X, Schwinn MK, Qin B, et al. HIF1A employs CDK8-mediator to stimulate RNAPII elongation in response to hypoxia. Cell. 2013;153:1327–1339.
Dupont S, Morsut L, Aragona M, Enzo E, Giulitti S, Cordenonsi M, et al. Role of YAP/TAZ in mechanotransduction. Nature. 2011;474:179–183.
Wrighton KH. Mechanotransduction: YAP and TAZ feel the force. Nat Rev Mol Cell Biol. 2011;12:404.
Osborne LD, Li GZ, How T, O’Brien ET, 3rd, Blobe GC, Superfine R et al. TGF-beta regulates LARG and GEF-H1 during EMT to impact stiffening response to force and cell invasion. Mol Biol Cell 2014;25:3528–3540.
Discher DE, Janmey P, Wang YL. Tissue cells feel and respond to the stiffness of their substrate. Science. 2005;310:1139–1143.
Paszek MJ, Zahir N, Johnson KR, Lakins JN, Rozenberg GI, Gefen A, et al. Tensional homeostasis and the malignant phenotype. Cancer Cell. 2005;8:241–254.
Broude EV, Gyorffy B, Chumanevich AA, Chen M, McDermott MS, Shtutman M, et al. Expression of CDK8 and CDK8-interacting genes as potential biomarkers in breast cancer. Curr Cancer Drug Targets. 2015;15:739–749.
Crown J. CDK8: a new breast cancer target. Oncotarget. 2017;8:14269–14270.
Firestein R, Bass AJ, Kim SY, Dunn IF, Silver SJ, Guney I, et al. CDK8 is a colorectal cancer oncogene that regulates beta-catenin activity. Nature. 2008;455:547–551.
McDermott MS, Chumanevich AA, Lim CU, Liang J, Chen M, Altilia S, et al. Inhibition of CDK8 mediator kinase suppresses estrogen dependent transcription and the growth of estrogen receptor positive breast cancer. Oncotarget. 2017;8:12558–12575.
Rzymski T, Mikula M, Wiklik K, Brzozka K. CDK8 kinase--An emerging target in targeted cancer therapy. Biochim Biophys Acta. 2015;1854:1617–1629.
Xu W, Wang Z, Zhang W, Qian K, Li H, Kong D, et al. Mutated K-ras activates CDK8 to stimulate the epithelial-to-mesenchymal transition in pancreatic cancer in part via the Wnt/beta-catenin signaling pathway. Cancer Lett. 2015;356:613–627.
Waldmeier L, Meyer-Schaller N, Diepenbruck M, Christofori G. Py2T murine breast cancer cells, a versatile model of TGFbeta-induced EMT in vitro and in vivo. PLoS ONE. 2012;7:e48651.
Inman GJ, Nicolas FJ, Callahan JF, Harling JD, Gaster LM, Reith AD, et al. SB-431542 is a potent and specific inhibitor of transforming growth factor-beta superfamily type I activin receptor-like kinase (ALK) receptors ALK4, ALK5, and ALK7. Mol Pharmacol. 2002;62:65–74.
Holtzhausen A, Golzio C, How T, Lee YH, Schiemann WP, Katsanis N, et al. Novel bone morphogenetic protein signaling through Smad2 and Smad3 to regulate cancer progression and development. FASEB J. 2013;28:1248–1267.
Petridou S, Maltseva O, Spanakis S, Masur SK. TGF-beta receptor expression and smad2 localization are cell density dependent in fibroblasts. Invest Ophthalmol Vis Sci. 2000;41:89–95.
Nallet-Staub F, Yin X, Gilbert C, Marsaud V, Ben Mimoun S, Javelaud D, et al. Cell density sensing alters TGF-beta signaling in a cell-type-specific manner, independent from Hippo pathway activation. Dev Cell. 2015;32:640–651.
Roninson IB, Porter DC, Wentland MP. CDK8-CDK19 selective inhibitors and their use in anti-metastatic and chemopreventative methods for cancer. US patent 9,321,737 (2016).
Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell. 2014;158:929–944.
Acloque H, Thiery JP, Nieto MA. The physiology and pathology of the EMT. Meeting on the epithelial-mesenchymal transition. EMBO Rep. 2008;9:322–326.
Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139:871–890.
Tiwari N, Tiwari VK, Waldmeier L, Balwierz PJ, Arnold P, Pachkov M, et al. Sox4 is a master regulator of epithelial-mesenchymal transition by controlling Ezh2 expression and epigenetic reprogramming. Cancer Cell. 2013;23:768–783.
Leight JL, Wozniak MA, Chen S, Lynch ML, Chen CS. Matrix rigidity regulates a switch between TGF-beta1-induced apoptosis and epithelial-mesenchymal transition. Mol Biol Cell. 2012;23:781–791.
Liu Y, He K, Hu Y, Guo X, Wang D, Shi W, et al. YAP modulates TGF-beta1-induced simultaneous apoptosis and EMT through upregulation of the EGF receptor. Sci Rep. 2017;7:45523.
Huang Z, Hu J, Pan J, Wang Y, Hu G, Zhou J, et al. YAP stabilizes SMAD1 and promotes BMP2-induced neocortical astrocytic differentiation. Development. 2016;143:2398–2409.
Inman GJ, Nicolas FJ, Hill CS. Nucleocytoplasmic shuttling of Smads 2, 3, and 4 permits sensing of TGF-beta receptor activity. Mol Cell. 2002;10:283–294.
Schmierer B, Hill CS. Kinetic analysis of Smad nucleocytoplasmic shuttling reveals a mechanism for transforming growth factor beta-dependent nuclear accumulation of Smads. Mol Cell Biol. 2005;25:9845–9858.
Chen HB, Rud JG, Lin K, Xu L. Nuclear targeting of transforming growth factor-beta-activated Smad complexes. J Biol Chem. 2005;280:21329–21336.
Varelas X, Sakuma R, Samavarchi-Tehrani P, Peerani R, Rao BM, Dembowy J, et al. TAZ controls Smad nucleocytoplasmic shuttling and regulates human embryonic stem-cell self-renewal. Nat Cell Biol. 2008;10:837–848.
Martina SJ, McDermott AAC, Chang-uk Lim, Jiaxin Liang, Mengqian Chen, Serena Altilia, David Oliver, James M. Rae, Michael Shtutman, Hippokratis Kiaris, Balázs Győrffy, Igor B. Roninson, Eugenia V. Broude. Inhibition of CDK8 Mediator kinase suppresses estrogen dependent transcription and the growth of estrogen receptor positive breast cancer. Oncotarget. 2017;8:12558–12575.
Varadaraj A, Patel P, Serrao A, Bandyopadhay T, Lee NY, Jazaeri AA, et al. Epigenetic regulation of GDF2 suppresses anoikis in ovarian and breast epithelia. Neoplasia. 2015;17:826–838.
Pannu J, Nakerakanti S, Smith E, ten Dijke P, Trojanowska M. Transforming growth factor-beta receptor type I-dependent fibrogenic gene program is mediated via activation of Smad1 and ERK1/2 pathways. J Biol Chem. 2007;282:10405–10413.
Swaminathan V, Mythreye K, O’Brien ET, Berchuck A, Blobe GC, Superfine R. Mechanical stiffness grades metastatic potential in patient tumor cells and in cancer cell lines. Cancer Res. 2011;71:5075–5080.
Chumanevich AA, Chaparala A, Witalison EE, Tashkandi H, Hofseth AB, Lane C, et al. Looking for the best anti-colitis medicine: A comparative analysis of current and prospective compounds. Oncotarget. 2017;8:228–237.
Acknowledgements
We thank the COBRE Functional Genomics Core (USC Center for Targeted Therapeutics) for lentiviral preparations, COBRE Microscopy and Flow cytometry Core (USC Center for Targeted Therapeutics) for assistance with imaging, Qi Zhang, Zach Mack, Nick Lenze, Priyanka Singh, and Pratik Patel for technical assistance. Grant Support: NIH P20GM109091 (E.V.B., K.M., I.B.R.), Marsha Rivkin Foundation (K.M.), Ovarian Cancer Research Fund (K.M.), University of South Carolina startup funds (K.M.), University of South Carolina SPARC Graduate Student Research, and Predoctoral NIH GM122379-01 (L.M.J.).
Author Contributions
A.S., L.M.J., A.C., and J.L. B.H conducted experiments, acquired and analyzed data. M.G. conducted the bioinformatics analyses. N.Y.L. provided reagents and analyzed data. E.V.B., I.B.R., and K.M. designed experiments, analyzed data, and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
I.B.R. is the founder and president and J.L., M.C., H.K., K.M, and E.V.B. are consultants of Senex Biotechnology, Inc. Other authors declare no conflict of interest.
Additional information
Equal first co-authorship: Anne Serrao and Laura M. Jenkins.
These authors contributed equally: Alexander A. Chumanevich, Ben Horst, Jiaxin Liang.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Serrao, A., Jenkins, L.M., Chumanevich, A.A. et al. Mediator kinase CDK8/CDK19 drives YAP1-dependent BMP4-induced EMT in cancer. Oncogene 37, 4792–4808 (2018). https://doi.org/10.1038/s41388-018-0316-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0316-y
This article is cited by
-
Discovery of small molecule degraders for modulating cell cycle
Frontiers of Medicine (2023)
-
PET imaging of new target CDK19 in prostate cancer
European Journal of Nuclear Medicine and Molecular Imaging (2023)
-
Exosomal circPABPC1 promotes colorectal cancer liver metastases by regulating HMGA2 in the nucleus and BMP4/ADAM19 in the cytoplasm
Cell Death Discovery (2022)
-
Hotair promotes the migration and proliferation in ovarian cancer by miR-222-3p/CDK19 axis
Cellular and Molecular Life Sciences (2022)
-
CDK8 maintains stemness and tumorigenicity of glioma stem cells by regulating the c-MYC pathway
Oncogene (2021)