Abstract
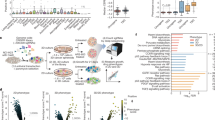
The RAS proteins are the most frequently mutated oncogenes in cancer, with highest frequency found in pancreatic, lung, and colon tumors. Moreover, the activity of RAS is required for the proliferation and/or survival of these tumor cells and thus represents a high-value target for therapeutic development. Direct targeting of RAS has proven challenging for multiple reasons stemming from the biology of the protein, the complexity of downstream effector pathways and upstream regulatory networks. Thus, significant efforts have been directed at identifying downstream targets on which RAS is dependent. These efforts have proven challenging, in part due to confounding factors such as reliance on two-dimensional adherent monolayer cell cultures that inadequately recapitulate the physiologic context to which cells are exposed in vivo. To overcome these issues, we implemented a high-throughput screening (HTS) approach using a spheroid-based 3-dimensional culture format, thought to more closely reflect conditions experienced by cells in vivo. Using isogenic cell pairs, differing in the status of KRAS, we identified Proscillaridin A as a selective inhibitor of cells harboring the oncogenic KRasG12V allele. Significantly, the identification of Proscillaridin A was facilitated by the 3D screening platform and would not have been discovered employing standard 2D culturing methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Pylayeva-Gupta Y, Grabocka E, Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer. 2011;11:761–74.
Downward J. Targeting RAS signalling pathways in cancer therapy. Nat Rev Cancer. 2003;3:11–22.
Prior IA, Lewis PD, Mattos C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 2012;72:2457–67.
Gysin S, Salt M, Young A, McCormick F. Therapeutic strategies for targeting ras proteins. Genes Cancer. 2011;2:359–72.
Singh A, Settleman J. Oncogenic K-ras “addiction” and synthetic lethality. Cell Cycle. 2009;8:2676–7.
Chan DA, Giaccia AJ. Harnessing synthetic lethal interactions in anticancer drug discovery. Nat Rev Drug Discov. 2011;10:351–64.
Scholl C, Frohling S, Dunn IF, Schinzel AC, Barbie DA, Kim SY, et al. Synthetic lethal interaction between oncogenic KRAS dependency and STK33 suppression in human cancer cells. Cell. 2009;137:821–34.
Barbie DA, Tamayo P, Boehm JS, Kim SY, Moody SE, Dunn IF, et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature. 2009;462:108–12.
Luo J, Emanuele MJ, Li D, Creighton CJ, Schlabach MR, Westbrook TF, et al. A genome-wide RNAi screen identifies multiple synthetic lethal interactions with the Ras oncogene. Cell. 2009;137:835–48.
Vicent S, Chen R, Sayles LC, Lin C, Walker RG, Gillespie AK, et al. Wilms tumor 1 (WT1) regulates KRAS-driven oncogenesis and senescence in mouse and human models. J Clin Invest. 2010;120:3940–52.
Wang Y, Ngo VN, Marani M, Yang Y, Wright G, Staudt LM, et al. Critical role for transcriptional repressor Snail2 in transformation by oncogenic RAS in colorectal carcinoma cells. Oncogene. 2010;29:4658–70.
Babij C, Zhang Y, Kurzeja RJ, Munzli A, Shehabeldin A, Fernando M, et al. STK33 kinase activity is nonessential in KRAS-dependent cancer cells. Cancer Res. 2011;71:5818–26.
Luo T, Masson K, Jaffe JD, Silkworth W, Ross NT, Scherer CA, et al. STK33 kinase inhibitor BRD-8899 has no effect on KRAS-dependent cancer cell viability. Proc Natl Acad Sci USA. 2012;109:2860–5.
Weiwer M, Spoonamore J, Wei J, Guichard B, Ross NT, Masson K, et al. A potent and selective quinoxalinone-based STK33 inhibitor does not show synthetic lethality in KRAS-dependent cells. ACS Med Chem Lett. 2012;3:1034–8.
Bittker JA, Weiwer M, Lewis T, Shimada K, Yang WS, MacPherson L, et al. Screen for RAS-selective lethal compounds and VDAC ligands—Probe 2. 2010, https://www.ncbi.nlm.nih.gov/pubmed/22834036.
Bittker JA, Weiwer M, Shimada K, Yang WS, MacPherson L, Dandapani S, et al. Screen for RAS-selective lethal compounds and VDAC ligands—Probe 1. 2010, https://www.ncbi.nlm.nih.gov/books/NBK55069/.
Torrance CJ, Agrawal V, Vogelstein B, Kinzler KW. Use of isogenic human cancer cells for high-throughput screening and drug discovery. Nat Biotechnol. 2001;19:940–5.
Guo W, Wu S, Liu J, Fang B. Identification of a small molecule with synthetic lethality for K-ras and protein kinase C iota. Cancer Res. 2008;68:7403–8.
Shaw AT, Winslow MM, Magendantz M, Ouyang C, Dowdle J, Subramanian A, et al. Selective killing of K-ras mutant cancer cells by small molecule inducers of oxidative stress. Proc Natl Acad Sci USA. 2011;108:8773–8.
Fennema E, Rivron N, Rouwkema J, van Blitterswijk C, de Boer J. Spheroid culture as a tool for creating 3D complex tissues. Trends Biotechnol. 2013;31:108–15.
Cukierman E, Pankov R, Stevens DR, Yamada KM. Taking cell-matrix adhesions to the third dimension. Science. 2001;294:1708–12.
Pampaloni F, Reynaud EG, Stelzer EH. The third dimension bridges the gap between cell culture and live tissue. Nat Rev Mol Cell Biol. 2007;8:839–45.
Antoni D, Burckel H, Josset E, Noel G. Three-dimensional cell culture: a breakthrough in vivo. Int J Mol Sci. 2015;16:5517–27.
Zanoni M, Piccinini F, Arienti C, Zamagni A, Santi S, Polico R, et al. 3D tumor spheroid models for in vitro therapeutic screening: a systematic approach to enhance the biological relevance of data obtained. Sci Rep. 2016;6:19103.
Yamada KM, Cukierman E. Modeling tissue morphogenesis and cancer in 3D. Cell. 2007;130:601–10.
Powell K. Adding depth to cell culture. Science . 2017;356:96–8.
Clevers H. Modeling development and disease with organoids. Cell. 2016;165:1586–97.
Madoux F, Tanner A, Vessels M, Willetts L, Hou S, Scampavia L, et al. A 1536-well 3D viability assay to assess the cytotoxic effect of drugs on spheroids. SLAS Discov. 2017;22:516–24.
Zhang J-H, Chung TDY, Oldenburg KR. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J Biomol Screen. 1999;4:67–73.
Longati P, Jia X, Eimer J, Wagman A, Witt M-R, Rehnmark S, et al. 3D pancreatic carcinoma spheroids induce a matrix-rich, chemoresistant phenotype offering a better model for drug testing. BMC Cancer. 2013;13:95.
Verjans ET, Doijen J, Luyten W, Landuyt B, Schoofs L. Three-dimensional cell culture models for anticancer drug screening: worth the effort? J Cell Physiol. 2017;233:2993–3003.
Tung YC, Hsiao AY, Allen SG, Torisawa YS, Ho M, Takayama S. High-throughput 3D spheroid culture and drug testing using a 384 hanging drop array. Analyst. 2011;136:473–8.
Ramaiahgari SC, den Braver MW, Herpers B, Terpstra V, Commandeur JN, van de Water B, et al. A 3D in vitro model of differentiated HepG2 cell spheroids with improved liver-like properties for repeated dose high-throughput toxicity studies. Arch Toxicol. 2014;88:1083–95.
Lingrel JB, Kuntzweiler T. Na +, K(+)-ATPase. J Biol Chem. 1994;269:19659–62.
Alevizopoulos K, Calogeropoulou T, Lang F, Stournaras C. Na +/K + ATPase inhibitors in cancer. Curr Drug Targets. 2014;15:988–1000.
Delebinski CI, Georgi S, Kleinsimon S, Twardziok M, Kopp B, Melzig MF, et al. Analysis of proliferation and apoptotic induction by 20 steroid glycosides in 143B osteosarcoma cells in vitro. Cell Prolif. 2015;48:600–10.
Denicolai E, Baeza-Kallee N, Tchoghandjian A, Carre M, Colin C, Jiglaire CJ, et al. Proscillaridin A is cytotoxic for glioblastoma cell lines and controls tumor xenograft growth in vivo. Oncotarget . 2014;5:10934–48.
Felth J, Rickardson L, Rosen J, Wickstrom M, Fryknas M, Lindskog M, et al. Cytotoxic effects of cardiac glycosides in colon cancer cells, alone and in combination with standard chemotherapeutic drugs. J Nat Prod. 2009;72:1969–74.
Kim N, Yim HY, He N, Lee CJ, Kim JH, Choi JS, et al. Cardiac glycosides display selective efficacy for STK11 mutant lung cancer. Sci Rep. 2016;6:29721.
Foster SA, Whalen DM, Ozen A, Wongchenko MJ, Yin J, Yen I, et al. Activation mechanism of oncogenic deletion mutations in BRAF, EGFR, and HER2. Cancer Cell. 2016;29:477–93.
Acknowledgements
We thank Pierre Baillargeon and Lina Deluca at Scripps for their help with compound management. This work was supported in part by the National Cancer Institute of the National Institutes of Health under Award Number R33CA206949 (TPS) and CA124495 (JK).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
These authors contributed equally: Smitha Kota, Shurong Hou, William Guerrant.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kota, S., Hou, S., Guerrant, W. et al. A novel three-dimensional high-throughput screening approach identifies inducers of a mutant KRAS selective lethal phenotype. Oncogene 37, 4372–4384 (2018). https://doi.org/10.1038/s41388-018-0257-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0257-5
This article is cited by
-
Signature-driven repurposing of Midostaurin for combination with MEK1/2 and KRASG12C inhibitors in lung cancer
Nature Communications (2023)
-
A combination of novel NSC small molecule inhibitor along with doxorubicin inhibits proliferation of triple-negative breast cancer through metabolic reprogramming
Oncogene (2022)
-
MISpheroID: a knowledgebase and transparency tool for minimum information in spheroid identity
Nature Methods (2021)
-
A Simple Procedure for Creating Scalable Phenotypic Screening Assays in Human Neurons
Scientific Reports (2019)