Abstract
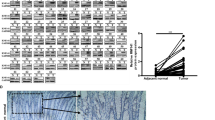
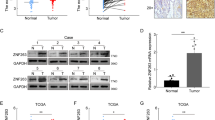
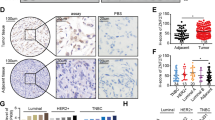
To elucidate the underlying oncogenic mechanism of zinc finger protein 746 (ZNF746), current study was conducted in colorectal cancers (CRCs). Herein, ZNF746 was overexpressed in HCT116, SW620, and SW480 cells, which was supported by CRC tissue microarray and TCGA analysis. Also, DNA microarray revealed the differentially expressed gene profile particularly related to cell cycle genes and c-Myc in ZNF746 depleted HCT116 cells. Furthermore, ZNF746 enhanced the stability of c-Myc via their direct binding through nuclear colocalization by immunoprecipitation and immunofluorescence, while ZNF746 and c-Myc exist mainly in nucleoplasm. Conversely, ZNF746 depletion attenuated phosphorylation of c-Myc (S62) and glycogen synthase kinase 3β (GSK3β) (S9) and also activated p-c-Myc (T58), which was reversed by GSK3 inhibitors such as SB-216763 and Enza. Also, c-Myc degradation by ZNF746 depletion was blocked by knockdown of F-box/WD repeat-containing protein 7 (FBW7) ubiquitin ligase or proteosomal inhibitor MG132. Additionally, the growth of ZNF746 depleted HCT116 cancer cells was retarded with decreased expression of ZNF746 and c-Myc. Overall, these findings suggest that ZNF746 promotes CRC progression via c-Myc stability mediated by GSK3 and FBW7.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Center MM, Jemal A, Ward E. International trends in colorectal cancer incidence rates. Cancer Epidemiol Biomark Prev. 2009;18:1688–94.
Bommer GT, Fearon ER. Role of c-Myc in Apc mutant intestinal phenotype: case closed or time for a new beginning? Cancer Cell. 2007;11:391–4.
He TC, et al. Identification of c-MYC as a target of the APC pathway. Science. 1998;281:1509–12.
Roy UK, et al. Wild-type APC regulates caveolin-1 expression in human colon adenocarcinoma cell lines via FOXO1a and C-myc. Mol Carcinog. 2008;47:947–55.
van de Wetering M, et al. The beta-catenin/TCF-4 complex imposes a crypt progenitor phenotype on colorectal cancer cells. Cell. 2002;111:241–50.
Garte SJ. The c-myc oncogene in tumor progression. Crit Rev Oncog. 1993;4:435–49.
Rothberg PG. The role of the oncogene c-myc in sporadic large bowel cancer and familial polyposis coli. Semin Surg Oncol. 1987;3:152–8.
Kelly K, Siebenlist U. The regulation and expression of c-myc in normal and malignant cells. Annu Rev Immunol. 1986;4:317–38.
Morello D, Asselin C, Lavenu A, Marcu KB, Babinet C. Tissue-specific post-transcriptional regulation of c-myc expression in normal and H-2K/human c-myc transgenic mice. Oncogene. 1989;4:955–61.
Nepveu A, Marcu KB, Skoultchi AI, Lachman HM. Contributions of transcriptional and post-transcriptional mechanisms to the regulation of c-myc expression in mouse erythroleukemia cells. Genes Dev. 1987;1:938–45.
Spencer CA, Groudine M. Control of c-myc regulation in normal and neoplastic cells. Adv Cancer Res. 1991;56:1–48.
Gregory MA, Qi Y, Hann SR. Phosphorylation by glycogen synthase kinase-3 controls c-myc proteolysis and subnuclear localization. J Biol Chem. 2003;278:51606–12.
McCubrey JA, et al. GSK-3 as potential target for therapeutic intervention in cancer. Oncotarget. 2014;5:2881–911.
Farrell AS, et al. Pin1 regulates the dynamics of c-Myc DNA binding to facilitate target gene regulation and oncogenesis. Mol Cell Biol. 2013;33:2930–49.
Popov N, Schulein C, Jaenicke LA, Eilers M. Ubiquitylation of the amino terminus of Myc by SCF(beta-TrCP) antagonizes SCF(Fbw7)-mediated turnover. Nat Cell Biol. 2010;12:973–81.
Shin JH, et al. PARIS (ZNF746) repression of PGC-1alpha contributes to neurodegeneration in Parkinson’s disease. Cell. 2011;144:689–702.
Veeriah S, et al. Somatic mutations of the Parkinson’s disease-associated gene PARK2 in glioblastoma and other human malignancies. Nat Genet. 2010;42:77–82.
Poulogiannis G, et al. PARK2 deletions occur frequently in sporadic colorectal cancer and accelerate adenoma development in Apc mutant mice. Proc Natl Acad Sci USA. 2010;107:15145–50.
Kim B, et al. Inhibition of ZNF746 suppresses invasion and epithelial to mesenchymal transition in H460 non-small cell lung cancer cells. Oncol Rep. 2014;31:73–78.
Karuppagounder SS, et al. The c-Abl inhibitor, nilotinib, protects dopaminergic neurons in a preclinical animal model of Parkinson’s disease. Sci Rep. 2014;4:4874.
Chen TC, et al. Nilotinib reduced the viability of human ovarian cancer cells via mitochondria-dependent apoptosis, independent of JNK activation. Toxicol Vitr. 2016;31:1–11.
Yuge R, et al. mTOR and PDGF pathway blockade inhibits liver metastasis of colorectal cancer by modulating the tumor microenvironment. Am J Pathol. 2015;185:399–408.
Won SH, Lee HJ, Jeong SJ, Lu J, Kim SH. Activation of p53 signaling and inhibition of androgen receptor mediate tanshinone IIA induced G1 arrest in LNCaP prostate cancer cells. Phytother Res PTR. 2012;26:669–74.
Bollag G, et al. Clinical efficacy of a RAF inhibitor needs broad target blockade in BRAF-mutant melanoma. Nature. 2010;467:596–9.
Conrad T, Orom UA. Cellular fractionation and isolation of chromatin-associated RNA. Methods Mol Biol. 2017;1468:1–9.
Gao SP, et al. Loss of TIM50 suppresses proliferation and induces apoptosis in breast cancer. Tumour Biol. 2016;37:1279–87.
Wang X, et al. Recurrent amplification of MYC and TNFRSF11B in 8q24 is associated with poor survival in patients with gastric cancer. Gastric Cancer. 2016;19:116–27.
Aslan JE, et al. Akt and 14-3-3 control a PACS-2 homeostatic switch that integrates membrane traffic with TRAIL-induced apoptosis. Mol Cell. 2009;34:497–509.
Yada M, et al. Phosphorylation-dependent degradation of c-Myc is mediated by the F-box protein Fbw7. EMBO J. 2004;23:2116–25.
Geng J, Xia L, Li W, Zhao C, Dou F. Cycloheximide treatment causes a ZVAD-sensitive protease-dependent cleavage of human Tau in drosophila cells. J Alzheimers Dis. 2016;49:1161–8.
Koukourakis GV, Sotiropoulou-Lontou A. Targeted therapy with bevacizumab (Avastin) for metastatic colorectal cancer. Clin Transl Oncol. 2011;13:710–4.
Kumar M, et al. Targeted cancer therapies: the future of cancer treatment. Acta Biomed. 2012;83:220–33.
Ji L, Wei Y, Jiang T, Wang S. Correlation of Nrf2, NQO1, MRP1, cmyc, and p53 in colorectal cancer and their relationships to clinicopathologic features and survival. Int J Clin Exp Pathol. 2014;7:1124–31.
de Mello RA, Marques AM, Araujo A. Epidermal growth factor receptor and metastatic colorectal cancer: insights into target therapies. World J Gastroenterol. 2013;19:6315–8.
Mansi L, Viel E, Curtit E, Medioni J, Le Tourneau C. Targeting the RAS signalling pathway in cancer. Bull Cancer. 2011;98:1019–28.
Schweiger T, et al. EGFR, BRAF and KRAS status in patients undergoing pulmonary metastasectomy from primary colorectal carcinoma: a prospective follow-up study. Ann Surg Oncol. 2014;21:946–54.
Chen BJ, Wu YL, Tanaka Y, Zhang W. Small molecules targeting c-Myc oncogene: promising anti-cancer therapeutics. Int J Biol Sci. 2014;10:1084–96.
Huang H, Weng H, Zhou H, Qu L. Attacking c-Myc: targeted and combined therapies for cancer. Curr Pharm Des. 2014;20:6543–54.
Prochownik EV. c-Myc as a therapeutic target in cancer. Expert Rev Anticancer Ther. 2004;4:289–302.
Robson S, Pelengaris S, Khan M. c-Myc and downstream targets in the pathogenesis and treatment of cancer. Recent Pat Anticancer Drug Discov. 2006;1:305–26.
Choi SH, Wright JB, Gerber SA, Cole MD. Myc protein is stabilized by suppression of a novel E3 ligase complex in cancer cells. Genes Dev. 2010;24:1236–41.
Koch HB, et al. Large-scale identification of c-MYC-associated proteins using a combined TAP/MudPIT approach. Cell Cycle. 2007;6:205–17.
Liu PY, et al. The histone deacetylase SIRT2 stabilizes Myc oncoproteins. Cell Death Differ. 2013;20:503–14.
Paul I, Ahmed SF, Bhowmik A, Deb S, Ghosh MK. The ubiquitin ligase CHIP regulates c-Myc stability and transcriptional activity. Oncogene. 2013;32:1284–95.
Schwamborn JC, Berezikov E, Knoblich JA. The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell. 2009;136:913–25.
Li M, et al. E3 ubiquitin ligase FBW7alpha inhibits cholangiocarcinoma cell proliferation by downregulating c-Myc and cyclin E. Oncol Rep. 2017;37:1627–36.
Welcker M, Clurman BE. Fbw7/hCDC4 dimerization regulates its substrate interactions. Cell Div. 2007;2:7.
Acknowledgements
This work was supported by the National Research Foundation of Korea (NRF) grant (no. 2017R1A2A1A17069297) and also we appreciate Dr. Han YH in KIRMAS for pcDNA3 reporter, 5′-V5-Myc, HA-FBW7, HA-Myc(S62A), and HA-Myc(T58A) and Dr. Lee ES for c-Myc mutant plasmids used in this project.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Jung, J.H., Jung, DB., Kim, H. et al. Zinc finger protein 746 promotes colorectal cancer progression via c-Myc stability mediated by glycogen synthase kinase 3β and F-box and WD repeat domain-containing 7. Oncogene 37, 3715–3728 (2018). https://doi.org/10.1038/s41388-018-0225-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0225-0
This article is cited by
-
Tespa1 facilitates hematopoietic and leukemic stem cell maintenance by restricting c-Myc degradation
Leukemia (2023)
-
Significant position of C-myc in colorectal cancer: a promising therapeutic target
Clinical and Translational Oncology (2022)
-
Actin-like protein 6A/MYC/CDK2 axis confers high proliferative activity in triple-negative breast cancer
Journal of Experimental & Clinical Cancer Research (2021)
-
ZNF746/PARIS overexpression induces cellular senescence through FoxO1/p21 axis activation in myoblasts
Cell Death & Disease (2020)
-
DUF3669, a “domain of unknown function” within ZNF746 and ZNF777, oligomerizes and contributes to transcriptional repression
BMC Molecular and Cell Biology (2019)