Abstract
Bromodomain and extra-terminal (BET) protein inhibitors have been reported as treatment options for acute myeloid leukemia (AML) in preclinical models and are currently being evaluated in clinical trials. This work presents a novel potent and selective BET inhibitor (BI 894999), which has recently entered clinical trials (NCT02516553). In preclinical studies, this compound is highly active in AML cell lines, primary patient samples, and xenografts. HEXIM1 is described as an excellent pharmacodynamic biomarker for target engagement in tumors as well as in blood. Mechanistic studies show that BI 894999 targets super-enhancer-regulated oncogenes and other lineage-specific factors, which are involved in the maintenance of the disease state. BI 894999 is active as monotherapy in AML xenografts, and in addition leads to strongly enhanced antitumor effects in combination with CDK9 inhibitors. This treatment combination results in a marked decrease of global p-Ser2 RNA polymerase II levels and leads to rapid induction of apoptosis in vitro and in vivo. Together, these data provide a strong rationale for the clinical evaluation of BI 894999 in AML.
Similar content being viewed by others
Introduction
Multiple protein complexes and enzymes are responsible for the patterning of chromatin modifications, and many of these molecules are functionally involved in cancer biology, thus representing promising cancer targets [1, 2]. BRD4 is a key epigenetic regulator and plays an important role in activating p-TEFb [3, 4]. This complex is activated by BRD4 and is a central mediator of transcriptional elongation [5]. Mechanistically, BRD4 was shown to govern the expression of various oncogenes, including MYC, by contributing to multi-protein complexes forming the so-called super-enhancers (SE) [6,7,8]. Furthermore, BRD4 was reported to contribute to the expression of lineage-specific genes by interacting with master-transcriptional regulators such as C/EBPb or FOSL2 in osteoblasts or Oct4, Sox2, and Nanog in embryonic stem cells [9,10,11,12].
Shortly after the generation of the first BET inhibitor [13], BRD4 has been demonstrated to be an exploitable dependency in acute myeloid leukemia (AML) [14, 15]. More recently, based on compelling preclinical evidence, BET inhibitors have entered clinical trials for AML and lymphomas with promising results [16, 17].
We studied the effects of the novel selective and potent BET inhibitor BI 894999, showing antitumor activity in cell lines, primary patient samples, and xenograft models of AML. Treatment with BI 894999 induces the expression of HEXIM1 mRNA, which we confirmed to be an excellent pharmacodynamic (PD) biomarker. Importantly, we report that combination treatment of BI 894999 and a CDK9 inhibitor leads to markedly improved efficacy by eliciting a rapid apoptotic response.
First promising clinical results, including antitumor efficacy and PD modulation, in a dose-escalation study of BI 894999 in solid cancers were recently reported [18].
Results
BI 894999 shows strong anti-proliferative activity in cell lines and primary patient samples
Our BRD4 structure-based drug design efforts resulted in the discovery of BI 894999, which belongs to the class of triazolopyrazines (Fig. 1a) and is structurally distinct from BET inhibitors with a benzodiazepine scaffold such as JQ1, OTX-015, or I-BET 762 [19]. BI 894999 was synthesized according to procedures described in WO 2014076237.
Structure and selectivity of BI 894999. a The structure of BI 894999. b AlphaLISA® assay for the respective bromodomains. Values for BRD4-BD1 and BD2 were determined in nine different experiments. c Selectivity screening by BROMOscan® ligand-binding site-directed competition assay. Image developed using TREEspotTM software (courtesy of DiscoverX, now Eurofins). Kd (nM) red < 2, gold < 200, blue < 10,000, gray > 10,000
The compound BI 894999 inhibits the binding of the BRD4-BD1 and BRD4-BD2 bromodomains to acetylated histones with an IC50 of 5 ± 3 nM and 41 ± 30 nM, respectively (Fig. 1b). Testing an extended number of bromodomains (n = 24), BI 894999 was highly selective for BRD2/3/4 and BRDT (Kd of 0.49–1.6 nM; Fig. 1c and Suppl. Table 1), with at least a 200-fold selectivity vs. BRD4-BD1.
The anti-proliferative activity of BI 894999 in various hematological malignancies is summarized in Table 1. Among 48 tested cell lines, all except the chronic myeloid leukemia (CML) line K562, which is known to be resistant to BET inhibition [20, 21], showed an EC50 < 100 nM. About half of all tested cell lines displayed an EC50 < 10 nM. BI 894999 consistently showed a 10–20-fold increased potency compared to OTX-015 on cell lines (Suppl. Table 2). In addition, firefly luciferase transduced cell lines used for in vivo monitoring of xenografts showed comparable EC50 values as the parental lines (Suppl. Table 3).
Analysis of MYC and HEXIM1 protein levels as candidate PD biomarkers [14, 15, 22] demonstrated variable effects for MYC downregulation upon treatment, whereas HEXIM1 was consistently induced in almost all cell lines within 24 h (Suppl. Figure 1).
In addition to cell lines, we also treated primary AML and multiple myeloma (MM) samples ex vivo with BI 894999 (Table 2 and Suppl. Table 4). While 16/20 AML patient samples showed EC50 values <10 nM, the results for MM were more heterogeneous with 4/15 moderate responders.
Treatment of MV-4-11B cells with BI 894999 for 72 h leads to induction of a G1 arrest with an increase in cells in sub-G1, indicative for apoptosis (Suppl. Figure 2). Significant inhibition of proliferation already at concentrations of 10 nM BI 894999 is demonstrated by IncuCyte® Live-Cell Analysis System. Dose-dependent apoptosis, measured by the IncuCyte® Caspase-3/7 apoptosis reagent, peaked at around 40 h after the start of treatment (Suppl. Figure 3A, B).
In vivo potency of BI 894999 as single agent or in combination with decitabine in AML models
For in vivo studies, BI 894999 was administered at daily oral doses of 2 or 4 mg/kg to mice engrafted intravenously with human MV-4-11B AML cells (Suppl. Table 5). This treatment resulted in tumor growth inhibition (TGI) of 96 or 99% and survival prolongation of 29.5 or 52 days, respectively (Fig. 2a). In another AML model (THP-1; Suppl. Figure 4), BI 894999 (daily, 4 mg/kg p.o.) was combined with the DNA methyltransferase inhibitor decitabine (twice per week, 0.5 mg/kg). Monotherapies of BI 894999 and decitabine (TGIs 94 and 99%; survival prolongation of 22 and 26 days) were inferior to the combination (TGI 100%; survival prolongation of 49 days; Suppl. Table 5). Together, these results demonstrate that BI 894999 is efficacious in a range of AML models in vivo.
In vivo potency of BI 894999 and HEXIM1 modulation in AML xenografts. a Kaplan–Meier curve of CIEA-NOG mice intravenously injected with 1 × 107 MV-4-11B AML cells shows prolonged survival of animals daily treated with 2 mg/kg (light blue) or 4 mg/kg (blue) BI 894999 compared to vehicle (gray line). b Tumor volumes of NMRI-nude mice carrying MV-4-11B s.c. xenografts daily treated with either vehicle control (gray), 2 mg/kg (light blue), 4 mg/kg (blue), or 12 mg/kg (dark blue) BI 894999. c Hexim1/HEXIM1 modulation in the blood (red) and tumors (blue) of mice from the respective treatment groups in the experiment shown in Fig. 2b, measured on the indicated days of treatment
Characterization of HEXIM1 mRNA induction as a PD biomarker
We evaluated HEXIM1 expression as a potential PD biomarker candidate marker for BET inhibition (Suppl. Figure 1 and refs. [21, 22]). To explore the relationship between BI 894999 dose, HEXIM1 modulation, and efficacy, we first treated MV-4-11B cells with increasing concentrations of the drug and measured HEXIM1 transcript levels by qRT-PCR as well as apoptosis (cleaved PARP, MSD assay) after 24 h (Suppl. Figure 5). Both, HEXIM1 transcript and cleaved PARP, show a stable, dose-dependent induction across the different doses used.
Additionally, we performed PK/PD analyses measuring HEXIM1 in subcutaneous xenografts, as tumor sampling is easily feasible compared to the disseminated setting. Daily treatment of MV-4-11B xenograft bearing mice with different doses of BI 894999 led to a TGI of ~50% in 2 and 4 mg/kg treatment groups, and a TGI of ~85% in the 12 mg/kg treatment group (Fig. 2b). Analysis of Hexim1 expression in the blood on days 2, 8, and 15 of treatment, always 6 h after dosing, demonstrated a dose-dependent increase in Hexim1 mRNA (Fig. 2c). Importantly, Hexim1 levels were back to baseline after 24 h, correlating with the PK (data not shown). The analysis of human HEXIM1 levels in tumors was comparable to Hexim1 kinetics in blood (Fig. 2c). In agreement with other reports, these data support HEXIM1 levels as an excellent PD biomarker for BET inhibitors [23,24,25].
Effect of BET inhibition on transcription across five AML cell lines
We performed transcriptional profiling (RNA-seq) of five AML cell lines (MV-4-11B, KASUMI-1, OCI-AML3, MOLM-13, and HL-60) treated with 35 nM BI 894999 for 4 h (Suppl. Figure 6A). Although there is some degree of variability across cell lines, we see a comparable transcriptional response for genes with similar baseline expression (see e.g., Suppl. Fig. 6B, C and Suppl. Table 6).
To explore the biological consequences of BET inhibition across all five AML cell lines, we performed gene set enrichment analyses (GSEA) based on the treatment-induced differential gene sets (Fig. 3a). Overall, we detected a significant overlap of BETi-repressed gene sets and a MYC hallmark target signature (Fig. 3b and ref. [14]) as well as signatures related to inflammation, IL2-STAT5 signaling, and myeloid cell development.
Gene set enrichment analysis for BETi treatment in five AML cell lines. a Gene sets significantly (FDR q-value ≤ 0.05) associated with BETi downregulated genes across five AML cell lines. Gene sets are ordered by hierarchical clustering, cell lines are ordered manually. In addition to RNA-seq, we performed H3K27ac ChIP-seq for cell lines marked by an asterisk. b GSEA plot showing a MYC target signature, which is heavily downregulated upon BETi treatment
BI 894999 efficiently targets super-enhancer-associated transcripts
From the above-mentioned gene expression analysis, we selected ten genes with a high baseline expression and a downregulation of ≥2-fold in at least two of three AML cell lines (Suppl. Figure 7A). In order to understand the role enhancers [7] play in the regulation of these genes, we performed H3K27ac chromatin immunoprecipitation sequencing (ChIP-seq) in these AML cell lines and the NCI-H2171 (SCLC, “non-AML” control). We generated “super-enhancer maps” with these data and found that the above-mentioned ten genes, which are repressed by BI 894999, are associated with a super-enhancer in the respective AML cell lines (10/10 MV-4-11B, 9/10 MOLM-13, and 8/10 OCI-AML3), but not in the SCLC cell line (0/10 NCI-H2171) (Fig. 4a, b and Suppl. Figure 7B). Although these genes are strongly repressed in the tested AML cell lines, modest to no effect in expression changes were observed in the SCLC cell line. This supports previous findings that several lineage-specific genes, associated with super-enhancers, are indeed major targets of BETi [7, 26].
Transcriptional (RNA-seq) and chromatin (H3K27ac ChIP-seq) profiling in AML and SCLC cell lines. a Super-enhancer hockey plots based on H3K27ac ChIP-seq signal. A set of pre-selected genes associated with super-enhancers in MV-4-11B are marked on all plots. Those genes that have no enhancer/super-enhancer associated in the respective cell lines are not shown in the hockey plots. b Jaccard distances measuring the similarity across super-enhancer sets of four cell lines and number of intersection between the respective super-enhancers (see a). c Top five DNA-binding motifs enriched within BRD4 ChIP-seq called super-enhancers in MV-4-11B and NCI-H2171
We could further confirm this association between lineage-specific genes/transcription factors and BRD4 binding [10] by investigating enriched sequence motifs located in BRD4-binding sites in MV-4-11B (AML) and NCI-H2171 (SCLC). This analysis shows that in both indications, BRD4 binds chromatin at sites that harbor DNA-binding motifs for transcription factors, implicated in the specific biology of the two respective cell lineages (Fig. 4c).
Super-enhancer-bound BRD4 is released from chromatin by BI 894999 in a dose-dependent manner
To test the effect of BI 894999 on chromatin-bound BRD4, we performed BRD4 ChIP-seq in MV-4-11B cells. About 40% of total bound BRD4 accumulated in 576 super-enhancer regions, while the rest of the signal was distributed across the remaining 11,747 normal enhancers (Fig. 5a).
BI 894999 represses expression of super-enhancer-driven genes in a dose-dependent manner. a BRD4 ChIP-seq-based super-enhancer hockey plot in MV-4-11B with key oncogenes. b Line graph showing the percentage of BRD4 chromatin occupancy normalized to DMSO after 4 h treatment with increasing doses of BI 894999. c Line graph showing the effect of increasing doses of BI 894999 on gene expression after 4 h treatment. The log2-fold-change of gene expression (y-axis) is plotted against the different treatment dosing groups (x-axis). Enhancers were associated with their closest gene based on transcriptional start site (TSS) to enhancer distances. d Gene tracks of BRD4 ChIP-seq occupancy after treatment with increased doses of BI 894999 at the MYC super-enhancer complex. Invidiual enhancer elements are denotated as E1–E5. Three major sub-super-enhancers are shown as a solid dark red bar on top of the graph. All tracks are normalized to the library sizes with identical scales on the y-axis. The percentage of BRD4 signal (summed up within the dashed box) normalized to BRD4 DMSO is annotated to the right of the respective tracks
To correlate the sensitivity of super-enhancers and changes of gene expression, we exposed MV-4-11B cells to a concentration series of BI 894999 and analyzed transcriptional as well as chromatin-bound BRD4 effects. At subefficacious concentration of BI 894999 (1 nM), we observed minor effects, both on BRD4 chromatin association (Fig. 5b, d) as well as on gene expression (Fig. 5c). Increasing the dose, we observed a dose-dependent reduction of BRD4 levels at normal enhancers, and a stronger effect at super-enhancers (Fig. 5b, d). Interestingly, we also saw a strong effect on the expression of super-enhancer-associated genes at 35 nM BI 894999 treatment and higher, which was less pronounced for genes associated with normal enhancers (Fig. 5c). These data are in line with previous reports that super-enhancer-driven genes are exquisitely vulnerable to BETi [7, 27, 28].
Finally, to compare the effects of BI 894999 on super-enhancer-regulated gene expression with those of another BET inhibitor (JQ1 [13]), we treated MV-4-11B cells with biologically equivalent concentrations of the two drugs, 10 and 200 nM, respectively (corresponding to anti-proliferative EC50 values). Analysis of gene expression by RNA-seq demonstrates that both compounds regulate the same set of transcripts (Suppl. Figs. 8–9).
BI 894999 synergizes with CDK9 inhibition, stopping transcriptional elongation
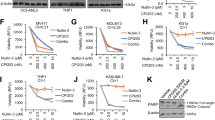
The notion that at an efficacious concentration of 35 nM does not completely repress super-enhancer-regulated gene expression, and the finding that single-agent treatment inhibits tumor growth, but does not lead to regressions in MV-4-11B xenografts (Fig. 2b), prompted us to evaluate potential combination partners to enhance efficacy. Since BRD4 has been proposed to contribute to transcriptional elongation by recruiting the positive transcription elongation factor (p-TEFb) [3], a protein complex containing the cyclin-dependent kinase 9 (CDK9), we explored the therapeutic potential of combining BI 894999 with CDK9 inhibition. We reasoned that additionally inhibiting p-TEFb/CDK9 activity might result in increased cellular activity of BI 894999. MV-4-11B cells treated with LDC000067, a CDK9i with a known selectivity profile [29], resulted only in a moderate EC50 of 5 µM. However, treatment of MV-4-11B cells with BI 894999 and LDC000067 resulted in strong synergy between the two drugs (Fig. 6a and Suppl. Fig. 10A). Notably, we confirmed this finding in vivo, as the combination of the two drugs led to regressions in the subcutaneous MV-4-11B model (Suppl. Fig. 10B).
CDK9 inhibition synergizes with BI 894999 treatment in vitro and in vivo. a Combination matrix of cellular growth inhibition of MV-4-11B cells treated with the indicated amounts of BI 894999 and Flavopiridol. b p-Ser2 POL-II levels in MV-4-11B cells treated for 2 h with the indicated drugs/combination. c RNA Pol-II ChIP-seq mean binding signals (top panels) across all length-normalized protein-coding genes in MV-4-11B cells treated with DMSO, BI 894999, LDC000067, and the combination (4 h). Read density heat maps (lower panels) visualizing individual genes grouped into four classes based on k-means clustering (Group 4: inactive genes; Group 3: transcriptionally engaged genes, with paused POL-II at promotor; Group 2: actively transcribed genes with paused POL-II at TSS/TES; Group 1: short highly transcriptional active genes)
CDK9 phosphorylates serine 2 in the YSPTSPS-heptad repeats of the carboxy-terminal-domain (CTD) of RNA polymerase II (POL-II) [30]. We therefore tested whether p-Ser2 POL-II levels are affected by combined BETi/CDK9i treatment. Treatment with 35 nM BI 894999 resulted in minor reduction of p-Ser2 POL-II levels (~86% of control), treatment with 5 µM LDC000067 led to ~64% of control, the combination of both showed the strongest effect (~46%) (Fig. 6b), suggesting that the combination might exert synergistic effects on transcriptional elongation.
To test the effects of the BETi/CDK9i combination on transcription, we treated MV-4-11B cells with 35 nM of BI 894999, 5 µM of LDC000067, or a combination (4 h treatment) and performed RNA-seq. BETi as well as CDK9i affected distinct sets of transcripts (Suppl. Fig. 11A, B and Suppl. Table 7), however, their combination led to the pronounced repression of many super-enhancer-driven genes (Suppl. Fig. 11B). In addition, combined BETi/CDK9i treatment led to strong downregulation of combination-specific genes (Suppl. Fig. 11C).
A caveat of this analysis is that RNA-seq might not reveal direct effects of interfering with POL-II elongation, rather reflecting bulk mRNA levels, which are affected by transcript-specific half-lives. To understand this better, we monitored POL-II localization by ChIP-seq. Unlike single agent treatments, the combination of BI 894999 and LDC000067 resulted in a drastic decrease of POL-II occupancy at the transcriptional end site (TES) of virtually all genes expressed in MV-4-11B cells (Fig. 6c). This underscores the necessity of applying various technologies to discriminate direct from indirect effects on transcription.
A combination of BI 894999 and the CDK9i flavopiridol is highly efficacious in vivo and rapidly induces apoptosis
To further address the consequences of combining BETi/CDK9i, we performed additional in vitro studies. For this, we switched to the less selective, but more potent CDK9i flavopiridol, which, unlike LDC000067, is currently tested for the treatment of AML (e.g., NCT02520011). Importantly, we could recapitulate the results obtained for LDC000067 with flavopiridol, i.e., the strong synergy with BI 894999 in proliferation assays (Suppl. Fig. 12A, B) and a strong effect on p-Ser2 POL-II levels 2 h after treatment (Suppl. Fig. 12C and 13A). Hence, the results obtained with both CDK9 inhibitors and BI 894999 are well comparable and support the notion that BETi/CDK9i can be efficiently combined [31, 32].
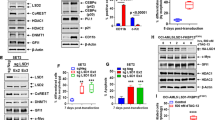
The same holds true for an in vivo experiment, as BI 894999 monotherapy (1 mg/kg qd) resulted in a TGI of 72%, flavopiridol monotherapy (2.5 mg/kg qdx5) in a TGI of 73%, and the combination of both drugs led to significantly enhanced efficacy (TGI 103%) and tumor regressions (Fig. 7a). While neither flavopiridol nor BETi single-agent treatment had an effect on p-Ser2 POL-II levels in tumors 2 h post dose, the combination led to a strong reduction of the p-Ser2 POL-II signal (Fig. 7b). In order to identify transcripts that could serve as pharmacodynamic biomarkers for the BETi/CDK9i combination, we performed RNA-seq of the tumors after 2 and 6 h and found combination-specific-regulated genes (Fig. 7c).
The effect of the BETi/CDK9i combination is BAX and BAK1 dependent and results in a global inhibition of transcriptional elongation. a Tumor volumes of NMRI-nude mice carrying MV-4-11B s.c. xenografts daily treated with either vehicle control (gray), 2.5 mg/kg flavopiridol (blue), 1 mg/kg BI 894999 (green), or a combination of both (red). Triangles show the duration of treatment. b p-Ser2 POL-II levels in MV-4-11B tumors 2 h after the indicated treatments. c Heat maps visualizing log2-fold-change gene expression values of the top 20 combination-specific differentially expressed genes in MV-4-11B tumors at the indicated treatments at 2 and 6 h. d Cleaved PARP levels in MV-4-11B tumors 6 h after the indicated treatments. e Log2-fold-changes of apoptotic regulator transcripts in MV-4-11B tumors treated with the respective doses of BI 894999, flavopiridol, and the combination 2 and 6 h post dose. f NOMO-1 cells were transduced with GFP-labeled CRISPR/Cas9 constructs targeting BAX or BAK1. From day 6, cells were treated as indicated and the fraction of GFP-positive viable cells determined by FACS
Upon 6 h of combination therapy, we detected a marked induction of apoptosis in the xenograft tumors, as indicated by elevated levels of cleaved PARP, whereas only modest effects were observed for the respective single-agent treatments (Fig. 7d). These findings recapitulate the results obtained in vitro (Suppl. Fig. 12D) and suggest that efficacy of the BI 894999/CDK9i combination is probably caused by rapid induction of apoptosis.
The apoptotic effector BAX contributes to the anti-proliferative effect of combined BETi/CDK9i treatment
To explore a mechanistic link between the induction of apoptosis and the transcriptional response elicited by the BETi/CDK9i combination, we conducted an analysis of apoptotic pathway transcripts in MV-4-11B xenografts. Neither pro- nor anti-apoptotic transcripts were found to be consistently altered by combined BETi/CDK9i (Fig. 7e). This interpretation is further supported by an analysis of pro- and anti-apoptotic proteins in BETi/CDK9i-treated cells (Supplemental Fig. 13A, B). Combined BETi and CDK9i results in decreased anti-apoptotic protein levels (MCL1 and BCLxL) as well as pro-apoptotic protein levels (PUMA and BIM) at early time points (2 and 4 h).
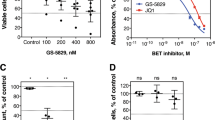
To nevertheless gauge the contribution of an apoptotic response to the effects of the BETi/CDK9i combination, we performed growth competition assays with cells expressing CRISPR/Cas9 constructs targeting BAX or BAK1, two essential effectors of the mitochondrial apoptosis pathway [33] (Fig. 7f and Suppl. Fig. 14). An increased fraction of BAX or BAK1 depleted living cells upon selection with the combination of BI 894999 and flavopiridol indicates a survival advantage of cells with defective apoptotic effectors. Neither depletion of BAX nor BAK1 rescued viability of NOMO-1 cells to a comparable degree for single-agent treatment (Suppl. Fig. 14B, C). As a positive control for BAX-dependent apoptosis [34], we also performed the assays with ABT-199 (Suppl. Fig. 14A).
FACS-based analysis of cells treated with the BETi/CDK9i combination resulted in a substantial increase of Annexin-V-positive cells indicative of apoptosis (Suppl. Fig. 14G). In addition, a fraction of Annexin-V-negative dead cells was detected upon BETi/CDK9i, indicating other, non-apoptotic cell death.
Discussion
Herein, we describe the characteristics of BI 894999, a novel BET bromodomain inhibitor, both in monotherapy as well as in combination therapies, focusing on CDK9i as an active partner for BET inhibition.
BI 894999 is distinct from other published BET inhibitors, as it belongs to the class of [1,2,4]triazolo[4,3-a]pyrazines [19]. Our molecule, which like other BET inhibitors [20, 35,36,37] also targets BRD2/3/T in addition to BRD4, exhibits a broad activity on cell lines and ex vivo-treated samples of various hematological indications, regulating the same genes as JQ1 at biologically comparable doses. BI 894999 is, however, a particularly potent BET inhibitor, as demonstrated by its high efficacy when tested in AML cell lines and patient samples, as well as in AML xenograft studies. We anticipate that the therapeutic window, based on the preferential effect of BET inhibitors on tumor cells rather than normal (hematopoietic) cells, with thrombocytopenia being a commonly observed dose-limiting toxicity [38] might be different for individual BET inhibitors and caused by distinct characteristics like potency or PK parameters.
Our studies have underscored the role of HEXIM1 as an excellent PD biomarker for BETi treatment, and indeed the translation of our preclinical findings has demonstrated that HEXIM1 is modulated in the blood of patients in our clinical trial (NCT02516553) and in ref. [18].
Although the precise mechanism of how HEXIM1 is induced by BET inhibition is still not understood, CDK9 is implicated in its induction, and is the reason why HEXIM1 cannot be considered as a PD marker for the BETi/CDK9i combination [39, 40].
While constituting an excellent PD marker, HEXIM1 induction (like MYC repression) is not a marker for efficacy (unpublished data and ref. [24]). In AML, it is difficult to disentangle cause or consequence for this as all cell lines tested are exquisitely sensitive to BI 894999 (Table 1), and we did not have the possibility to analyze patient-derived samples molecularly. More generally, although mechanisms of resistance are emerging [21, 41, 42], so far no patient selection biomarker for BETi treatment is known. A recent study, in which JQ1 was used in over 1000 cancer cell lines to identify pharmacogenomics markers of sensitivity, has led to the identification of FAT1 mutations in HNSCC and POLR2B mutations in breast cancer as predictors for sensitivity [43]. Likewise SPOP mutations appear to render prostate cancer cell lines resistant to BET inhibition. The clinical relevance of these findings will likely be addressed in the near future. Analysis of mutations in our AML cell line panel does neither allow the identification of predictors of response (e.g., TP53, NPM1; Suppl. Table 8), as all cell lines are sensitive.
One hypothesis that has been proposed for several years, but never formally proven, is that the sensitivity of cancer cells to BET inhibition is mediated by the repression of cancer-specific super-enhancer-driven oncogenes [6, 7, 26, 28]. The nature of these super-enhancers is still debated [44,45,46], and it is becoming apparent that they consist of many small enhancers with individual, lineage-specific transcription factor binding sites and dependencies [10, 47].
Although we and others have shown a connection between super-enhancers and gene expression [7], future studies on this topic are needed, as currently the correlation between super-enhancer presence and the effects on gene expression are mainly based on data from ChIP-seq and RNA-seq analyses, which does not differentiate between direct and indirect effects on transcription (e.g., for short-lived proteins like MYC).
Initially, we had hypothesized that synergy on the repression of super-enhancer-regulated transcripts might constitute a potential mechanism behind the synergy, we observed in vitro and in vivo for the combination of BI 894999 and CDK9i, as these super-enhancers were reported to regulate pause release at associated genes [48,49,50].
When we compared RNA-seq data with POL-II ChIP-seq data of cells treated with a combination of BI 894999 and a CDK9i or the respective single agents, we indeed found that the RNA-seq analysis suggests that many super-enhancer-regulated genes are strongly affected by combination treatment. However, POL-II ChIP-seq data rather point at a pronounced global effect on POL-II elongation caused exclusively by BET/CDK9 inhibition [50]. Future studies employing analysis of nascent RNA will hopefully clarify this and other related issues.
The finding that BI 894999/CDK9i combinations lead to globally reduced p-Ser2 POL-II levels measured from protein lysate support the notion that indeed this treatment leads to a global arrest of transcriptional elongation. We could actually identify some transcripts specific for the combination as potential PD markers, although we think that due to the underlying mechanism these transcripts might be heavily influenced by tumor-to-tumor variabilities in mRNA half-lives and should be treated with caution.
With the precise mechanistic basis of the BETi/CDK9i combination being complex [5, 51, 52], its exquisite effect on AML cell and tumor viability seems clearer. The combination of BI 894999 with either flavopiridol or LDC000067 results in a rapid induction of apoptosis in vivo and in cultured cells. Our CRISPR-based apoptotic effector inactivation experiments as well as apoptotic marker protein analyses establish that BAX-dependent apoptosis contributes to the remarkable efficacy of the BETi/CDK9i combination. As combined BETi/CDK9i leads to induction of apoptosis in cell lines with complete loss of a functional TP53 gene (e.g., for NOMO-1 Cosmic sample ID COSS908451, confirmed by in-house analyses), we conclude that induction of apoptosis by the BETi/CDK9i combination does not depend on a functional TP53 pathway. However, given the broad effect of the inhibitor combination on global transcriptional elongation, it is not surprising that also non-apoptotic cell death appears to contribute to the anti-proliferative effects of suppression of transcriptional elongation. In line with the observed global effects on transcription, we could not identify individual mediators of apoptosis on the protein levels specifically regulated by combination treatment.
In addition to CDK9i, BET inhibition may be effectively combined with various other treatment approaches [29, 36, 53,54,55,56,57], and we have confirmed this for the combination with decitabine and volasertib (manuscript under review).
While many aspects remain to be understood in mechanistic terms, both for BET inhibition as a single agent therapy, and even more for the combination with CDK9 inhibition, our data further substantiate that this novel therapeutic concept deserves careful clinical evaluation.
Material and methods
BRD4 AlphaLISA assay
This bead-based proximity assay is based on the competitive displacement of a histone peptide (acetyl-histone H4/ Lys5, 8, 12, and 16) from human bromodomain by the test compound as described previously [58].
Analysis of drug combinations
Effects of drug combinations (antagonistic, antagonistic, or synergistic) were analyzed using the Bliss Independence Model [59,60,61].
p-Ser2 POL-II custom MSD assay
p-Ser2 levels of POL-II were measured by using a custom MSD assay from Meso Scale Diagnostics (Rockville, US-MD) on multi-array 96-well plates.
Efficacy studies in mice
All animal studies were approved by the internal ethics committee and the local governmental committee.
This analysis follows largely the procedures described in refs. [58, 62] with extensions outlined in the supplementary information.
Cell culture
Tumor cell lines were obtained from the American Type Culture Collection (Manassas, US-VA) or the German Collection of Microorganisms and Cell Culture (DSMZ, Braunschweig, Germany). MV-4-11B is a clone derived from the AML cell line MV-4-11 from ATCC (CRL-9591), carrying a heterozygous mutation in TP53 (c.742C>T, p.R248W). All cell lines used in this study were cultured according to the manufacturer’s instruction and authenticated by short tandem repeat (STR) analysis at Boehringer Ingelheim (Suppl. Table 9).
Viability assays
Viability assays were performed on 96-well flat-bottom microtiter plates and incubated with alamarBlue® solution (Invitrogen, Carlsbad, US-CA).
Assays with primary bone marrow samples from AML patients
Studies were performed at Vivia Biotech, Madrid, Spain and were done as described before [58].
Incucyte assay
Incucyte assays were performed as described previously [58].
RNA isolation
Samples were treated with 500 μl TRI Reagent® (Sigma, #T9424, St. Louis, US-MO) and 50 μl of 1-Bromo-3-chloropropane (Sigma, #B9673, St. Louis, US-MO) was added. Total RNA was isolated with RNAeasy Mini Kit (Qiagen, #74106, Venlo, the Netherlands).
RNA isolation and sequencing
Cells were lysed in TRI lysis reagent (Sigma-Aldrich, T9424, St. Louis, US-MO). RNA-seq sequencing libraries were prepared using the TruSeq RNA Library Preparation Kit v2 and subsequently sequenced on the Illumina HiSeq1500.
cDNA synthesis and RT-qPCR
For cDNA, 500 ng of total RNA were with SuperScript® VILO™ cDNA Synthesis Kit (Invitrogen, #11754-010, Waltham, US-MA), and for qPCR2x SYBR® Green PCR Master Mix (AppliedBiosystems by Lifetechnologies # 4309155, Waltham, US-MA) was used according to the manufacturers’ instructions.
ChIP protocol
The protocol is based on ref. [63] with further improvements and extensions as described in the supplementary information. The following antibodies were used: 5 µg H3K27ac (Diagenode, #C15410196, Liège, Belgium) and 10 µg BRD4 (Bethyl, #A301-985A50, Montgomery, US-TX).
Library preparation from ChIP samples
An aliquot of 10 ng of ChIP material per sample was used for library preparation. NEBNext® ChIP-Seq Library Prep Master Mix Set for Illumina® (NEB #E6240L, Ipswich, US-MA) and for the quantity measurement KAPA Library Quantification Kit (KapaBiosystems, # KK4824, Basel, Switzerland) was used according to the manufacturers’ instructions. Samples with 2 nM concentration were multiplexed for sequencing.
Bioinformatics analysis
Data analyses and visualizations were mainly performed using R and Bioconductor packages.
Gene expression analysis
RNA-seq data were processed as described in ref. [62]. If not otherwise stated, an adjusted p-value of 0.05 and an absolute fold-change of 2 were used as cutoffs. Gene set enrichment analyses (GSEA) were performed using the GSEA software version 2.1.0 and MSigDB 5.0.
ChIP-seq analysis
Reads were trimmed with cutadapt and mapped to the genome with bowtie2. MACS and HOMER were used to perform peak calling and super-enhancer calling from uniquely mapped reads. Sequence motif analyses were also performed using HOMER. The deepTools package was used for analyzing and plotting genome-wide ChIP-seq profiles.
Genomics analysis
Bedtools were used for performing genome arithmetics and intersections of data sets.
Super-enhancer plots
Disproportional BRD4/H3K27ac binding to chromatin was visualized via “hockey-plots” as described in ref. [11].
Data availability
Sequencing data was submitted to the Gene Expression Omnibus (accession number GSE101821) database repository.
References
Easwaran H., Tsai H.C., Baylin S.B. Cancer epigenetics: tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol Cell. 2014;54:716–27.
Simo-Riudalbas L., Esteller M. Targeting the histone orthography of cancer: drugs for writers, erasers and readers. Br J Pharmacol. 2015;172:2716–32.
Yang Z., Yik J.H., Chen R., He N., Jang M.K., Ozato K., et al. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol Cell. 2005;19:535–45.
Jang M.K., Mochizuki K., Zhou M., Jeong H.S., Brady J.N., Ozato K. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol Cell. 2005;19:523–34.
Jonkers I., Lis J.T. Getting up to speed with transcription elongation by RNA polymerase II. Nat Rev Mol Cell Biol. 2015;16:167–77.
Hnisz D., Abraham B.J., Lee T.I., Lau A., Saint-Andre V., Sigova A.A., et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Loven J., Hoke H.A., Lin C.Y., Lau A., Orlando D.A., Vakoc C.R., et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell. 2013;153:320–34.
Shi J., Vakoc C.R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol Cell. 2014;54:728–36.
Najafova Z., Tirado-Magallanes R., Subramaniam M., Hossan T., Schmidt G., Nagarajan S., et al. BRD4 localization to lineage-specific enhancers is associated with a distinct transcription factor repertoire. Nucleic Acids Res. 2017;45:127–41.
Roe J.S., Mercan F., Rivera K., Pappin D.J., Vakoc C.R. BET bromodomain inhibition suppresses the function of hematopoietic transcription factors in acute myeloid leukemia. Mol Cell. 2015;58:1028–39.
Whyte W.A., Orlando D.A., Hnisz D., Abraham B.J., Lin C.Y., Kagey M.H., et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–19.
Zhang W., Prakash C., Sum C., Gong Y., Li Y., Kwok J.J., et al. Bromodomain-containing protein 4 (BRD4) regulates RNA polymerase II serine 2 phosphorylation in human CD4+T cells. J Biol Chem. 2012;287:43137–55.
Filippakopoulos P., Qi J., Picaud S., Shen Y., Smith W.B., Fedorov O., et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–73.
Delmore J.E., Issa G.C., Lemieux M.E., Rahl P.B., Shi J., Jacobs H.M., et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–17.
Zuber J., Shi J., Wang E., Rappaport A.R., Herrmann H., Sison E.A., et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature. 2011;478:524–8.
Amorim S., Stathis A., Gleeson M., Iyengar S., Magarotto V., Leleu X., et al. Bromodomain inhibitor OTX015 in patients with lymphoma or multiple myeloma: a dose-escalation, open-label, pharmacokinetic, phase 1 study. Lancet Haematol. 2016;3:e196–204.
Berthon C., Raffoux E., Thomas X., Vey N., Gomez-Roca C., Yee K., et al. Bromodomain inhibitor OTX015 in patients with acute leukaemia: a dose-escalation, phase 1 study. Lancet Haematol. 2016;3:e186–195.
Aftimos P.G., Bechter O., Awada A., Jungels C., Dumez H., Huyvaert N., et al. Phase I first-in-man trial of a novel bromodomain and extra-terminal domain (BET) inhibitor (BI 894999) in patients (Pts) with advanced solid tumors. J Clin Oncol. 2017;35:2504–2504.
Jung M., Gelato K.A., Fernandez-Montalvan A., Siegel S., Haendler B. Targeting BET bromodomains for cancer treatment. Epigenomics. 2015;7:487–501.
Coude M.M., Braun T., Berrou J., Dupont M., Bertrand S., Masse A., et al. BET inhibitor OTX015 targets BRD2 and BRD4 and decreases c-MYC in acute leukemia cells. Oncotarget. 2015;6:17698–712.
Rathert P., Roth M., Neumann T., Muerdter F., Roe J.S., Muhar M., et al. Transcriptional plasticity promotes primary and acquired resistance to BET inhibition. Nature. 2015;525:543–7.
Devaraj S.G.T., Fiskus W., Sharma S., Qi J., Shah B., Schaub L.J., et al. Abstract 3686: HEXIM1 induction exerts a mechanistic role and is a biomarker of lethal activity of BRD4 antagonist against human AML cells. Cancer Res. 2014;74:3686.
Lin X., Huang X., Uziel T., Hessler P., Albert D.H., Roberts-Rapp L.A., et al. HEXIM1 as a robust pharmacodynamic marker for monitoring target engagement of BET family bromodomain inhibitors in tumors and surrogate tissues. Mol Cancer Ther. 2017;16:388–96.
Yeh T.C., O’Connor G., Petteruti P., Dulak A., Hattersley M., Barrett J.C., et al. Identification of CCR2 and CD180 as robust pharmacodynamic tumor and blood biomarkers for clinical use with BRD4/BET inhibitors. Clin Cancer Res. 2017;23:1025–35.
Berenguer-Daize C., Astorgues-Xerri L., Odore E., Cayol M., Cvitkovic E., Noel K., et al. OTX015 (MK-8628), a novel BET inhibitor, displays in vitro and in vivo antitumor effects alone and in combination with conventional therapies in glioblastoma models. Int J Cancer. 2016;139:2047–55.
Henssen A., Althoff K., Odersky A., Beckers A., Koche R., Speleman F., et al. Targeting MYCN-driven transcription by BET-bromodomain inhibition. Clin Cancer Res. 2016;22:2470–81.
Yokoyama Y., Zhu H., Lee J.H., Kossenkov A.V., Wu S.Y., Wickramasinghe J.M., et al. BET inhibitors suppress ALDH activity by targeting ALDH1A1 super-enhancer in ovarian cancer. Cancer Res. 2016;76:6320–30.
Chapuy B., McKeown M.R., Lin C.Y., Monti S., Roemer M.G., Qi J., et al. Discovery and characterization of super-enhancer-associated dependencies in diffuse large B cell lymphoma. Cancer Cell. 2013;24:777–90.
Muralidharan S.V., Bhadury J., Nilsson L.M., Green L.C., McLure K.G., Nilsson J.A. BET bromodomain inhibitors synergize with ATR inhibitors to induce DNA damage, apoptosis, senescence-associated secretory pathway and ER stress in Myc-induced lymphoma cells. Oncogene. 2016;35:4689–97.
Schuller R., Forne I., Straub T., Schreieck A., Texier Y., Shah N., et al. Heptad-specific phosphorylation of RNA polymerase II CTD. Mol Cell. 2016;61:305–14.
Moreno N., Holsten T., Mertins J., Zhogbi A., Johann P., Kool M., et al. Combined BRD4 and CDK9 inhibition as a new therapeutic approach in malignant rhabdoid tumors. Oncotarget. 2017;8:84986–95.
Lee Y.S., Kim W., Soh K.K., Peterson P., Whatcott C.J., Siddiqui-Jain A., et al. Abstract C202: CDK9 inhibition synergizes with BRD4 inhibitor-mediated super enhancer transcriptional repression in multiple preclinical tumor models. Mol Cancer Ther. 2015;14:C202–C202.
Wei M.C., Zong W.X., Cheng E.H., Lindsten T., Panoutsakopoulou V., Ross A.J., et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science. 2001;292:727–30.
Sarosiek K.A., Chi X., Bachman J.A., Sims J.J., Montero J., Patel L., et al. BID preferentially activates BAK while BIM preferentially activates BAX, affecting chemotherapy response. Mol Cell. 2013;51:751–65.
Herrmann H., Blatt K., Shi J., Gleixner K.V., Cerny-Reiterer S., Mullauer L., et al. Small-molecule inhibition of BRD4 as a new potent approach to eliminate leukemic stem- and progenitor cells in acute myeloid leukemia AML. Oncotarget. 2012;3:1588–99.
Boi M., Gaudio E., Bonetti P., Kwee I., Bernasconi E., Tarantelli C., et al. The BET bromodomain inhibitor OTX015 affects pathogenetic pathways in preclinical B-cell tumor models and synergizes with targeted drugs. Clin Cancer Res. 2015;21:1628–38.
Chaidos A., Caputo V., Karadimitris A. Inhibition of bromodomain and extra-terminal proteins (BET) as a potential therapeutic approach in haematological malignancies: emerging preclinical and clinical evidence. Ther Adv Hematol. 2015;6:128–41.
Doroshow D.B., Eder J.P., LoRusso P.M. BET inhibitors: a novel epigenetic approach. Ann Oncol. 2017;28:1776–87.
Bartholomeeusen K., Xiang Y., Fujinaga K., Peterlin B.M. Bromodomain and extra-terminal (BET) bromodomain inhibition activate transcription via transient release of positive transcription elongation factor b (P-TEFb) from 7SK small nuclear ribonucleoprotein. J Biol Chem. 2012;287:36609–16.
Liu P., Xiang Y., Fujinaga K., Bartholomeeusen K., Nilson K.A., Price D.H., et al. Release of positive transcription elongation factor b (P-TEFb) from 7SK small nuclear ribonucleoprotein (snRNP) activates hexamethylene bisacetamide-inducible protein (HEXIM1) transcription. J Biol Chem. 2014;289:9918–25.
Fong C.Y., Gilan O., Lam E.Y., Rubin A.F., Ftouni S., Tyler D., et al. BET inhibitor resistance emerges from leukaemia stem cells. Nature. 2015;525:538–42.
Shu S., Lin C.Y., He H.H., Witwicki R.M., Tabassum D.P., Roberts J.M., et al. Response and resistance to BET bromodomain inhibitors in triple-negative breast cancer. Nature. 2016;529:413–7.
Iorio F., Knijnenburg T.A., Vis D.J., Bignell G.R., Menden M.P., Schubert M., et al. A landscape of pharmacogenomic interactions in cancer. Cell. 2016;166:740–54.
Pott S., Lieb J.D. What are super-enhancers? Nat Genet. 2015;47:8–12.
Dukler N., Gulko B., Huang Y.F., Siepel A. Is a super-enhancer greater than the sum of its parts? Nat Genet. 2016;49:2–3.
Hnisz D., Shrinivas K., Young R.A., Chakraborty A.K., Sharp P.A. A phase separation model for transcriptional control. Cell. 2017;169:13–23.
Hnisz D., Schuijers J., Lin C.Y., Weintraub A.S., Abraham B.J., Lee T.I., et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol Cell. 2015;58:362–70.
Gressel S., Schwalb B., Decker T.M., Qin W., Leonhardt H., Eick D., et al. CDK9-dependent RNA polymerase II pausing controls transcription initiation. Elife. 2017;6:e29736.
Liu W., Ma Q., Wong K., Li W., Ohgi K., Zhang J., et al. Brd4 and JMJD6-associated anti-pause enhancers in regulation of transcriptional pause release. Cell. 2013;155:1581–95.
Winter G.E., Mayer A., Buckley D.L., Erb M.A., Roderick J.E., Vittori S., et al. BET bromodomain proteins function as master transcription elongation factors independent of CDK9 recruitment. Mol Cell. 2017;67:5–18.e19.
Bres V., Yoh S.M., Jones K.A. The multi-tasking P-TEFb complex. Curr Opin Cell Biol. 2008;20:334–40.
Kanno T., Kanno Y., LeRoy G., Campos E., Sun H.W., Brooks S.R., et al. BRD4 assists elongation of both coding and enhancer RNAs by interacting with acetylated histones. Nat Struct Mol Biol. 2014;21:1047–57.
Ceribelli M., Kelly P.N., Shaffer A.L., Wright G.W., Xiao W., Yang Y., et al. Blockade of oncogenic IkappaB kinase activity in diffuse large B-cell lymphoma by bromodomain and extraterminal domain protein inhibitors. Proc Natl Acad Sci USA. 2014;111:11365–70.
Mazur P.K., Herner A., Mello S.S., Wirth M., Hausmann S., Sanchez-Rivera F.J., et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat Med. 2015;21:1163–71.
Siegel M.B., Liu S.Q., Davare M.A., Spurgeon S.E., Loriaux M.M., Druker B.J., et al. Small molecule inhibitor screen identifies synergistic activity of the bromodomain inhibitor CPI203 and bortezomib in drug resistant myeloma. Oncotarget. 2015;6:18921–32.
Yao W., Yue P., Khuri F.R., Sun S.Y. The BET bromodomain inhibitor, JQ1, facilitates c-FLIP degradation and enhances TRAIL-induced apoptosis independent of BRD4 and c-Myc inhibition. Oncotarget. 2015;6:34669–79.
Shahbazi J., Liu P.Y., Atmadibrata B., Bradner J.E., Marshall G.M., Lock R.B., et al. The bromodomain inhibitor JQ1 and the histone deacetylase inhibitor panobinostat synergistically reduce N-Myc expression and induce anticancer effects. Clin Cancer Res. 2016;22:2534–44.
Rudolph D., Impagnatiello M.A., Blaukopf C., Sommer C., Gerlich D.W., Roth M., et al. Efficacy and mechanism of action of volasertib, a potent and selective inhibitor of Polo-like kinases, in preclinical models of acute myeloid leukemia. J Pharmacol Exp Ther. 2015;352:579–89.
Bliss C.I. The toxicity of poisons applied jointly. Ann Appl Biol. 1939;26:585–615.
Bliss C.I. The calculation of microbial assays. Bacteriol Rev. 1956;20:243–58.
Greco W.R., Bravo G., Parsons J.C. The search for synergy: a critical review from a response surface perspective. Pharmacol Rev. 1995;47:331–85.
Waizenegger I.C., Baum A., Steurer S., Stadtmuller H., Bader G., Schaaf O., et al. A novel RAF kinase inhibitor with DFG-out-binding mode: high efficacy in BRAF-mutant tumor xenograft models in the absence of normal tissue hyperproliferation. Mol Cancer Ther. 2016;15:354–65.
Lee T.I., Johnstone S.E., Young R.A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat Protoc. 2006;1:729–48.
Acknowledgements
We want to thank Christoph Albrecht, Alexandra Beran, Alika Cotten, Sandra Döbel, Wolfgang Egermann, Georg Egger, Christian Engelhardt, Monika Fraunhofer, Nina Rodi, Vanessa Roessler, Regina Ruzicka, Christian Salamon, Susanne Schmittner, Jochen Schön, Andreas Schrenk, Susy Straubinger, Eva Strauss, Roland Varecka, Anika Weiss, Corinna Wieshofer, and Andreas Wernitznig for technological and methodology expertise.
Author contributions
Conception, design, and supervision: FS, UT-G, DG, and NK. Development of methodology: FS, UT-G, and DG. Acquisition of data (provided animals, provided facilities, etc.): UT-G, FS, OK, MHH, AB, NS, TG, JB, and JP. Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): UT-G, FS, OK, DG, NS, MHH, AB, DS, HE, and JP. Writing, review, and/or revision of the manuscript: FS, UT-G, DG, JP, JZ, and NK.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors except JZ are employees of Boehringer Ingelheim. The main sponsor of the IMP is Boehringer Ingelheim.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gerlach, D., Tontsch-Grunt, U., Baum, A. et al. The novel BET bromodomain inhibitor BI 894999 represses super-enhancer-associated transcription and synergizes with CDK9 inhibition in AML. Oncogene 37, 2687–2701 (2018). https://doi.org/10.1038/s41388-018-0150-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-018-0150-2
This article is cited by
-
KLF7 regulates super-enhancer-driven IGF2BP2 overexpression to promote the progression of head and neck squamous cell carcinoma
Journal of Experimental & Clinical Cancer Research (2024)
-
Cancer cell plasticity: from cellular, molecular, and genetic mechanisms to tumor heterogeneity and drug resistance
Cancer and Metastasis Reviews (2024)
-
The multi-CDK inhibitor dinaciclib reverses bromo- and extra-terminal domain (BET) inhibitor resistance in acute myeloid leukemia via inhibition of Wnt/β-catenin signaling
Experimental Hematology & Oncology (2024)
-
Molecular basis of GDF15 induction and suppression by drugs in cardiomyocytes and cancer cells toward precision medicine
Scientific Reports (2023)
-
MYC: a multipurpose oncogene with prognostic and therapeutic implications in blood malignancies
Journal of Hematology & Oncology (2021)