Abstract
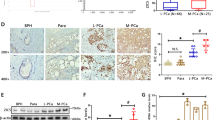
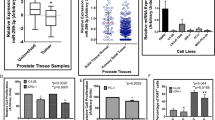
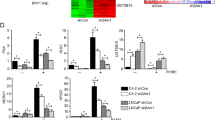
Prostate cancer growth is promoted by the gene regulatory action of androgen receptor (AR) and its downstream signals. The aberrant dysfunction of tumor suppressor p53 has an important role in the prognosis of cancer. We previously found that androgen treatments translocate p53 to the cytoplasm. The mechanism of this translocation depends on sumoylation of p53 by complex of SUMO E3 ligase RanBP2 with androgen-induced GTPase-activating protein-binding protein 2 (G3BP2). Here, we identified tripartite motif-containing protein 25 (TRIM25)/estrogen-responsive finger protein (Efp) as a novel interacting partner of G3BP2 protein complex. Then, we demonstrated that TRIM25 knockdown resulted in p53 downstream activation for cell cycle inhibition and apoptosis induction in LNCaP and 22Rv1 cells. In contrast, overexpression of TRIM25 promoted prostate cancer cell proliferation and inhibited apoptosis by docetaxel treatment in LNCaP cells. We observed that p53 activity was reduced by mechanism of G3BP2-mediated nuclear export in TRIM25-overexpressing prostate cancer cells. We also found TRIM25 is important for G3BP2/RanBP2-mediated p53 modification. Clinically, we newly demonstrated that TRIM25 is a prognostic factor for prostate cancer patients. Expression of TRIM25 is significantly associated with cytoplasmic p53 expression and G3BP2. Moreover, TRIM25 knockdown results in reduced tumor growth and increased p53 activity in the mouse xenograft model of prostate cancer. Thus, our findings show that overexpression of TRIM25 promoted prostate cancer cell proliferation and cell survival by modulating p53 nuclear export mechanism with G3BP2 interaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Vousden KH, Prives C. Blinded by the light: the growing complexity of p53. Cell. 2009;137:413–31.
Melchior F, Hengst L. SUMO-1 and p53. Cell Cycle. 2002;1:245–9.
Müller S, Ledl A, Schmidt D. SUMO: a regulator of gene expression and genome integrity. Oncogene. 2004;23:1998–2008.
Pant V, Lozano G. Limiting the power of p53 through the ubiquitin proteasome pathway. Genes Dev. 2014;28:1739–51.
Yang Y, Li CC, Weissman AM. Regulating the p53 system through ubiquitination. Oncogene. 2004;23:2096–106.
Carter S, Bischof O, Dejean A, Vousden KH. C-terminal modifications regulate MDM2 dissociation and nuclear export of p53. Nat Cell Biol. 2007;9:428–35.
Muller PA, Vousden KH. Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell. 2014;25:304–17.
Cai C, Yuan X, Balk SP. Androgen receptor epigenetics. Transl Androl Urol. 2013;2:148–57.
Yuan X, Cai C, Chen S, Chen S, Yu Z, Balk SP. Androgen receptor functions in castration-resistant prostate cancer and mechanisms of resistance to new agents targeting the androgen axis. Oncogene. 2014;33:2815–25.
Chen CD, Welsbie DS, Tran C, Baek SH, Chen R, Vessella R, et al Molecular determinants of resistance to antiandrogen therapy. Nat Med. 2004;10:33–9.
Cai C, He HH, Chen S, Coleman I, Wang H, Fang Z, et al. Androgen receptor gene expression in prostate cancer is directly suppressed by the androgen receptor through recruitment of lysine-specific demethylase 1. Cancer Cell. 2011;20:457–71.
Wang Q, Li W, Zhang Y, Yuan X, Xu K, Yu J, et al. Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell. 2009;138:245–56.
Linja MJ, Savinainen KJ, Saramäki OR, Tammela TL, Vessella RL, Visakorpi T. Amplification and overexpression of androgen receptor gene in hormone-refractory prostate cancer. Cancer Res. 2001;61:3550–5.
Takayama K, Inoue S. The emerging role of noncoding RNA in prostate cancer progression and its implication on diagnosis and treatment. Brief Funct Genom. 2016;15:257–65.
Takayama K, Inoue S. Transcriptional network of androgen receptor in prostate cancer progression. Int J Urol. 2013;20:756–68.
Ashikari D, Takayama K, Tanaka T, Obinata D, Fujimura T, et al. Androgen induces G3BP2 and SUMO-mediated p53 nuclear export in prostate cancer. Oncogene. 2017;36:6272–81.
Ikeda K, Inoue S, Orimo A, Tsutsumi K, Muramatsu M. Promoter analysis of mouse estrogen-responsive finger protein (efp) gene: mouse efp promoter contains an E-box that is also conserved in human. Gene. 1998;216:155–62.
Orimo A, Inoue S, Minowa O, Tominaga N, Tomioka Y, Sato M, et al Underdeveloped uterus and reduced estrogen responsiveness in mice with disruption of the estrogen-responsive finger protein gene, which is a direct target of estrogen receptor alpha. Proc Natl Acad Sci USA. 1999;96:12027–32.
Inoue S, Orimo A, Matsuda Y, Inazawa J, Emi M, Nakamura Y, et al Chromosome mapping of human (ZNF147) and mouse genes for estrogen-responsive finger protein (efp), a member of the RING finger family. Genomics. 1995;25:581–3.
Shimada N, Suzuki T, Inoue S, Kato K, Imatani A, Sekine H, et al. Systemic distribution of estrogen-responsive finger protein (Efp) in human tissues. Mol Cell Endocrinol. 2004;218:147–53.
Urano T, Saito T, Tsukui T, Fujita M, Hosoi T, Muramatsu M, et al Efp targets 14-3-3 sigma for proteolysis and promotes breast tumour growth. Nature. 2002;417:871–5.
Ikeda K, Orimo A, Higashi Y, Muramatsu M, Inoue S. Efp as a primary estrogen-responsive gene in human breast cancer. FEBS Lett. 2000;472:9–13.
Suzuki T, Urano T, Tsukui T, Horie-Inoue K, Moriya T, Ishida T, et al. Estrogen-responsive finger protein as a new potential biomarker for breast cancer. Clin Cancer Res. 2005;11:6148–54.
Sakuma M, Akahira J, Suzuki T, Inoue S, Ito K, Moriya T, et al. Expression of estrogen-responsive finger protein (Efp) is associated with advanced disease in human epithelial ovarian cancer. Gynecol Oncol. 2005;99:664–70.
Wang S, Kollipara RK, Humphries CG, Ma SH, Hutchinson R, Li R, et al The ubiquitin ligase TRIM25 targets ERG for degradation in prostate cancer. Oncotarget. 2016;7:64921–31.
Horie-Inoue K, Inoue S. Epigenetic and proteolytic inactivation of 14-3-3sigma in breast and prostate cancers. Semin Cancer Biol. 2006;16:235–9.
Zurita M, Lara PC, del Moral R, Torres B, Linares-Fernández JL, Arrabal SR, et al. Hypermethylated 14-3-3-sigma and ESR1 gene promoters in serum as candidate biomarkers for the diagnosis and treatment efficacy of breast cancer metastasis. BMC Cancer. 2010;10:217
Moreira JM, Ohlsson G, Rank FE, Celis JE. Down-regulation of the tumor suppressor protein 14-3-3sigma is a sporadic event in cancer of the breast. Mol Cell Proteom. 2005;4:555–69.
Simooka H, Oyama T, Sano T, Horiguchi J, Nakajima T. Immunohistochemical analysis of 14-3-3 sigma and related proteins in hyperplastic and neoplastic breast lesions, with particular reference to early carcinogenesis. Pathol Int. 2004;54:595–602.
Umbricht CB, Evron E, Gabrielson E, Ferguson A, Marks J, Sukumar S. Hypermethylation of 14-3-3 sigma (stratifin) is an early event in breast cancer. Oncogene. 2001;20:3348–53.
Ferguson AT, Evron E, Umbricht CB, Pandita TK, Chan TA, Hermeking H, et al High frequency of hypermethylation at the 14-3-3 sigma locus leads to gene silencing in breast cancer. Proc Natl Acad Sci USA. 2000;97:6049–54.
Henrique R, Jerónimo C, Hoque MO, Carvalho AL, Oliveira J, Teixeira MR, et al. Frequent 14-3-3 sigma promoter methylation in benign and malignant prostate lesions. DNA Cell Biol. 2005;24:264–9.
Cheng L, Pan CX, Zhang JT, Zhang S, Kinch MS, Li L, et al. Loss of 14-3-3sigma in prostate cancer and its precursors. Clin Cancer Res. 2004;10:3064–8.
Urano T, Takahashi S, Suzuki T, Fujimura T, Fujita M, Kumagai J, et al. 14-3-3sigma is down-regulated in human prostate cancer. Biochem Biophys Res Commun. 2004;319:795–800.
Jain S, Wheeler JR, Walters RW, Agrawal A, Barsic A, Parker R. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell. 2016;164:487–98.
Zhang P, Elabd S, Hammer S, Solozobova V, Yan H, Bartel F, et al. TRIM25 has a dual function in the p53/Mdm2 circuit. Oncogene. 2015;34:5729–38.
Liu C, Zhu Y, Lou W, Nadiminty N, Chen X, Zhou Q, et al Functional p53 determines docetaxel sensitivity in prostate cancer cells. Prostate. 2013;73:418–27.
Zhao KW, Sikriwal D, Dong X, Guo P, Sun X, Dong JT. Oestrogen causes degradation of KLF5 by inducing the E3 ubiquitin ligase EFP in ER-positive breast cancer cells. Biochem J. 2011;437:323–33.
Lee NR, Kim HI, Choi MS, Yi CM, Inn KS. Regulation of MDA5-MAVS antiviral signaling axis by TRIM25 through TRAF6-mediated NF-κB activation. Mol Cells. 2015;38:759–64.
Gack MU, Shin YC, Joo CH, Urano T, Liang C, Sun L, et al TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature. 2007;446:916–20.
Nakasato N, Ikeda K, Urano T, Horie-Inoue K, Takeda S, Inoue S. A ubiquitin E3 ligase Efp is up-regulated by interferons and conjugated with ISG15. Biochem Biophys Res Commun. 2006;351:540–6.
Brenner JC, Ateeq B, Li Y, Yocum AK, Cao Q, Asangani IA, et al Mechanistic rationale for inhibition of poly(ADP-ribose) polymerase in ETS gene fusion-positive prostate cancer. Cancer Cell. 2011;19:664–78.
Kim MM, Wiederschain D, Kennedy D, Hansen E, Yuan ZM. Modulation of p53 and MDM2 activity by novel interaction with Ras-GAP binding proteins (G3BP). Oncogene. 2007;26:4209–15.
Agell L, Hernández S, de Muga S, Lorente JA, Juanpere N, Esgueva R, et al. KLF6 and TP53 mutations are a rare event in prostate cancer: distinguishing between Taq polymerase artifacts and true mutations. Mod Pathol. 2008;21:1470–8.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Yue X, Zhao Y, Xu Y, Zheng M, Feng Z, Hu W. Mutant p53 in cancer: accumulation, gain-of-function, and therapy. J Mol Biol. 2017;429:1595–606.
Valentino E, Bellazzo A, Di Minin G, Sicari D, Apollonio M, Scognamiglio G, et al. Mutant p53 potentiates the oncogenic effects of insulin by inhibiting the tumor suppressor DAB2IP. Proc Natl Acad Sci USA. 2017;114:7623–8.
Takayama K, Horie-Inoue K, Katayama S, Suzuki T, Tsutsumi S, Ikeda K, et al Androgen-responsive long noncoding RNA CTBP1-AS promotes prostate cancer. EMBO J. 2013;32:1665–80.
Takayama K, Suzuki T, Fujimura T, Urano T, Takahashi S, Homma Y, et al CtBP2 modulates the androgen receptor to promote prostate cancer progression. Cancer Res. 2014;74:6542–53.
Takayama K, Misawa A, Suzuki T, Takagi K, Hayashizaki Y, Fujimura T, et al TET2 repression by androgen hormone regulates global hydroxymethylation status and prostate cancer progression. Nat Commun. 2015;6:8219.
Takayama K, Tsutsumi S, Katayama S, Okayama T, Horie-Inoue K, Ikeda K, et al. Integration of cap analysis of gene expression and chromatin immunoprecipitation analysis on array reveals genome-wide androgen receptor signaling in prostate cancer cells. Oncogene. 2011;30:619–30.
Ueyama K, Ikeda K, Sato W, Nakasato N, Horie-Inoue K, Takeda S, et al Knockdown of Efp by DNA-modified small interfering RNA inhibits breast cancer cell proliferation and in vivo tumor growth. Cancer Gene Ther. 2010;17:624–32.
Acknowledgements
This work was supported by grants of the P-DIRECT and the P-CREATE from the Ministry of Education, Culture, Sports, Science and Technology, Japan (to SI); grants from the Japan Society for the Promotion of Science, Japan (to KT, grant 15K15581; to SI, grant 15K15353); a grant of the Program for Promotion of Fundamental Studies in Health Sciences from the National Institute of Biomedical Innovation, Japan (to SI); Grants-in-Aid (to SI) from the Ministry of Health, Labor and Welfare, Japan, and grants from the Takeda Science Foundation (to SI and KT), the Terumo foundation for Life Sciences and Arts (to KT) and the NOVARTIS Foundation for the Promotion of Science (to KT), Japan.
Author contributions
KT designed the study, performed experiments and analyzed data. TS performed IHC analysis. TT analyzed data. TF and ST prepared and analyzed tumor samples. TU and KI provided important materials such as plasmids and siRNAs. SI supervised the study. KT and SI wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Takayama, Ki., Suzuki, T., Tanaka, T. et al. TRIM25 enhances cell growth and cell survival by modulating p53 signals via interaction with G3BP2 in prostate cancer. Oncogene 37, 2165–2180 (2018). https://doi.org/10.1038/s41388-017-0095-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-017-0095-x
This article is cited by
-
TRIM25 promotes glioblastoma cell growth and invasion via regulation of the PRMT1/c-MYC pathway by targeting the splicing factor NONO
Journal of Experimental & Clinical Cancer Research (2024)
-
Tripartite-motif 3 represses ovarian cancer progression by downregulating lactate dehydrogenase A and inhibiting AKT signaling
Molecular and Cellular Biochemistry (2024)
-
TRIM25 promotes temozolomide resistance in glioma by regulating oxidative stress and ferroptotic cell death via the ubiquitination of keap1
Oncogene (2023)
-
USP7- and PRMT5-dependent G3BP2 stabilization drives de novo lipogenesis and tumorigenesis of HNSC
Cell Death & Disease (2023)
-
Therapeutic, diagnostic and prognostic values of TRIM proteins in prostate cancer
Pharmacological Reports (2023)