Abstract
The AP-1 transcription factor JunB plays crucial roles in multiple biological processes, including placental formation and bone homeostasis. We recently reported that JunB is essential for development of Th17 cells, and thus Junb-deficient mice are resistant to experimental autoimmune encephalomyelitis. However, the role of JunB in CD4+ T cells under other inflammatory disease conditions is unknown. Here we show that mice lacking JunB in CD4+ T cells (Junbfl/flCd4-Cre mice) were more susceptible to dextran sulfate sodium (DSS)-induced colitis because of impaired development of regulatory T (Treg) cells. Production of interleukin (IL)-2 and expression of CD25, a high affinity IL-2 receptor component, were decreased in Junb-deficient CD4+ T cells in vitro and in vivo. Naive CD4+ T cells from Junbfl/flCd4-Cre mice failed to differentiate into Treg cells in the absence of exogenously added IL-2 in vitro. A mixed bone marrow transfer experiment revealed that defective Treg development of Junb-deficient CD4+ T cells was not rescued by co-transferred wild-type cells, indicating a significance of the cell-intrinsic defect. Injection of IL-2-anti-IL-2 antibody complexes induced expansion of Treg cells and alleviated DSS-induced colitis in Junbfl/flCd4-Cre mice. Thus JunB plays a crucial role in the development of Treg cells by facilitating IL-2 signaling.
Similar content being viewed by others
Introduction
The AP-1 transcription factor JunB, which is encoded by the immediate early gene Junb, mediates multiple biological processes including placentation, suppression of myeloid cell proliferation, and maintenance of bone and skin homeostasis.1,2,3,4 JunB is a basic leucine zipper (bZIP) protein and forms an AP-1 complex by dimerizing with other bZIP proteins, such as Fos or BATF family members. Recently we and other groups have reported that JunB is essential for the development of IL-17-producing helper T (Th17) cells, and thus Junb-deficient mice are completely resistant to experimental autoimmune encephalomyelitis (EAE).5,6,7 In addition, the transfer of CD45RBhiCD25−CD4+ T cells from Junb-deficient mice to Rag1-deficient mice fails to induce colitis,6 suggesting that JunB is indispensable for pathogenicity of T cells in these autoimmune diseases. On the other hand, it is unclear whether JunB in CD4+ T cells is involved in other inflammatory diseases such as those induced by innate immune cells.
Inflammatory bowel diseases (IBDs) are inflammatory disorders that are characterized by mucosal damage and intestinal inflammation.8,9 Although the etiology of the IBDs is poorly understood, aberrant activation of immune cells against intestinal microbiota has been proposed to be a cause of the disease. In addition to the genetic and environmental factors, imbalance of the microbiota composition, called dysbiosis, is suggested to play a key role in the pathology of IBDs.10 Ulcerative colitis (UC) and Crohn’s disease (CD) are two major types of IBDs. The dextran sulfate sodium (DSS)-induced colitis is an experimental mouse model of human IBDs.11,12 In this model, administration of DSS via drinking water causes epithelial barrier damages, which allow penetration of intestinal microbiota into injured mucosa to induce inflammation through infiltration of innate immune cells. DSS-induced colitis shares several aspects of clinical pathology with human UC such as confined symptoms at the colon. Because severe combined immunodeficient (SCID) mice are not resistant to DSS-induced colitis,13 this disease model is considered to be evoked by innate immune cells. Rag2−/− mice are more susceptible to DSS-induced colitis, and the pathology is alleviated by transfer of wild-type CD4+ T cells,14 indicating a protective role for CD4+ T cells in the DSS-induced colitis.
Regulatory T (Treg) cells, specified by expression of the transcription factor forkhead box P3 (Foxp3), play crucial roles in maintenance of homeostasis by limiting multiple immune responses.15,16,17 It has been shown that Treg cells ameliorate DSS-induced colitis.18,19 Development of Treg cells requires IL-2 signaling, as indicated by the fact that mice deficient in IL-2 or its receptor subunits exhibit a marked reduction in Treg cells, leading to lethal autoimmune disorders.20 Among the three subunits composing the IL-2 receptor, the inducible α chain (CD25), encoded by Il2ra, is essential for the response to the low levels of IL-2 that are produced physiologically.21 While CD25 is hardly detectable in resting T cells, its expression is upregulated under the Treg-inducing conditions where cells are activated through T cell receptor (TCR) in the presence of TGF-β1 and IL-2.21,22,23 Since Il2ra is a target gene of Foxp3,24,25 accumulation of Foxp3 during the course of Treg development accelerates upregulation of CD25 expression, leading to a further increase in sensitivity to IL-2.
In the present study, we show that DSS-induced colitis was exacerbated in CD4+ T cell-specific Junb-deficient mice (Junbfl/flCd4-Cre mice). Although the number of Th17 cells was reduced, the expression of Th17-associated cytokines, such as IL-22 and IL-17A, was not diminished in the colon of Junbfl/flCd4-Cre mice. Notably, Junbfl/flCd4-Cre mice exhibited a significant reduction in the number of Treg cells. Junb−/− CD4+ T cells failed to differentiate into Treg cells in the absence of exogenously added IL-2 in vitro, because the production of IL-2 and upregulation of CD25 were impaired in Junb−/− CD4+ T cells. A mixed bone marrow transfer experiment revealed that defect in Treg development of Junb−/− CD4+ T cells was not rescued in the presence of wild-type CD4+ T cells, indicating a significance of the cell-intrinsic defect in vivo. On the other hand, injection of immune complexes composed of IL-2 and anti-IL-2 antibody induced upregulation of CD25 and subsequent expansion of Treg cells, resulting in attenuation of DSS-induced colitis in Junbfl/flCd4-Cre mice. Thus, JunB plays a crucial role in the development of Treg cells by upregulating IL-2 signaling, thereby attenuating DSS-induced colitis.
Results
Mice lacking Junb in CD4+ T cells are resistant to T cell-mediated autoimmune disease models
We previously reported that Junbfl/flMeox2Cre mice, in which Junb is deleted in all tissues, are resistant to EAE.5 Consistent with previous studies,6,7 we confirmed that mice lacking Junb specifically in CD4+ T cells (Junbfl/flCd4-Cre mice) also did not develop EAE (Fig. 1a). To examine the role for JunB in other T cell-mediated disease models, we employed an intestinal inflammation model induced by injection of the anti-CD3 agonistic antibody.26 Upon injection of the anti-CD3 antibody, activated T cells induce severe tissue damages of the small intestine, followed by establishment of tolerized state.27 As expected, injection of the antibody led to body weight loss, but the extent was markedly attenuated in Junbfl/flCd4-Cre mice compared to control (Junbfl/fl) mice (Fig. 1b). Thus JunB plays a crucial role in the development of T cell-dependent autoimmune diseases in a CD4+ T cell-dependent manner.
JunB is required for the development of T cell-mediated autoimmune disease models. a Control (Junbfl/fl) or Junbfl/flCd4-Cre mice were immunized with MOG31-55 peptide, and the clinical scores were measured. Results are means ± SEMs (n = 6–7 mice per each genotype). b Control or Junbfl/flCd4-Cre mice were intraperitoneally injected with anti-CD3 antibody (20 μg per mouse) at day 0, 2, and 4. Body weight was measured every 24 h. The average of body weight is shown as percentage relative to the initial value. Results are means ± SEMs (n = 7 mice per each genotype). *p < 0.05, **p < 0.01, and ***p < 0.001 (two-tailed Student’s unpaired t test)
DSS-induced colitis is exacerbated in Junb fl/fl Cd4-Cre mice
While JunB is required for the development of T cell-dependent diseases, its role in other inflammatory conditions is unknown. To elucidate the role for JunB in CD4+ T cells in inflammatory diseases that are induced by innate immune cells, we employed the DSS-induced colitis model. Following DSS administration, control mice started to have bloody stool at day 2 or day 3, lost their body weight up to 5% during the first 7 days, and then recovered by day 10. Intriguingly, body weight loss of Junbfl/flCd4-Cre mice was more severe compared to littermate control mice following DSS administration (Fig. 2a). The colon length of Junbfl/flCd4-Cre mice was significantly shorter than that of control mice at day 5 after DSS administration (Fig. 2b), which is a hallmark of severity of the colitis.11 Histological analysis revealed that destruction of villi structure and the edema between the mucosal and the muscular layers were worsened in the colon of Junbfl/flCd4-Cre mice (Fig. 2c). Moreover, the percentage of infiltrated neutrophils (Ly6G+CD11b+ cells) was increased in the colon of Junbfl/flCd4-Cre mice (Fig. 2d). On the other hand, deletion of Junb in myeloid cells or intestinal epithelial cells did not lead to exacerbation of the colitis (Supplementary Figure 1a, b). Thus the expression of JunB in CD4+ T cells is required for attenuating DSS-induced colitis.
DSS-induced colitis is exacerbated in Junbfl/flCd4-Cre mice. a Control or Junbfl/flCd4-Cre mice were administrated with 2.0% DSS in drinking water for 5 days, and then they were given by normal water in the following days. The average of body weight is shown as percentage relative to the initial value. Results are means ± SEMs (n = 9–11 mice per each genotype). b Representative image of macroscopic observation of the colons at day 5 after administration of DSS. The graph shows the average length of the colons. Results are means ± SEMs (n = 13 mice per each genotype). c Hematoxylin and eosin staining of the colon sections (n = 4 mice per each genotype). Scale bars = 100 μm. d Colonic lamina propria cells were isolated at day 9 after administration of DSS, and analyzed by flow cytometry. Representative results of flow cytometry are shown (n = 4 mice, left). The graph shows the average percentage of CD11b+Ly6G+ neutrophils among CD45.2+ cells. Results are means ± SEMs (n = 4 mice per each genotype). *p < 0.05, **p < 0.01, ****p < 0.0001 (two-tailed Student’s unpaired t test)
IFN-γ-producing colitogenic T cells are not increased in the colon of Junb fl/fl Cd4-Cre mice after DSS administration
It might be possible that deletion of Junb in CD4+ T cells accelerates the development of pathogenic T cells such as IFN-γ-producing Th1 cells under certain conditions. To test this possibility, we analyzed the numbers of IFN-γ- and IL-17A-producing CD4+ T cells in colonic lamina propria at day 5 after DSS treatment. The percentage of IFN-γ-producing CD4+ T cells in the colon of Junbfl/flCd4-Cre mice was comparable to that of the control mice (Fig. 3a, b). Consistent with our previous study,5 the proportion of Th17 cells was markedly reduced in the colon of DSS-induced Junbfl/flCd4-Cre mice (Fig. 3a,c and Supplementary Figure 2a). Thus, the deletion of Junb in CD4+ T cells does not cause an increase in colitogenic Th1 cells.
The number of IFN-γ-producing pathogenic T cells is not increased in the colon of Junbfl/flCd4-Cre mice after DSS administration. a Colonic lamina propria cells were isolated from untreated mice or mice administrated with DSS for 5 days, and analyzed by flow cytometry. Representative results of flow cytometry are shown (n = 4 mice). As shown in Supplementary Figure 2, FVD506−CD45.2+CD4+TCRβ+ cells (CD4+ T cells) are presented. b, c Average percentages of IFN-γ+ cells (b) or IL-17A+ cells (c) among CD4+ T cells are shown. Results are means ± SEMs (n = 4–6 mice per each genotype). *p < 0.05, **p < 0.01, ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
The expression of Th17-associated cytokines is not reduced in the colon of Junb fl/fl Cd4-Cre mice
To further characterize the exacerbation of DSS-induced colitis in Junbfl/flCd4-Cre mice, we performed genome-wide transcriptome analysis using total RNA prepared from the colon of control and Junbfl/flCd4-Cre mice after DSS administration (Fig. 4a and Supplementary Table 1). The expression of genes encoding proinflammatory cytokines, including Il6, Ifng, and Il1b, was elevated in the colon of DSS-treated Junbfl/flCd4-Cre mice compared to control mice, whereas the expression of Il10 was not largely affected (Fig. 4b). The elevation of Ifng expression in the colon of Junbfl/flCd4-Cre mice was likely to be contributed by cells in the CD4− TCRβ− fraction (Supplementary Figure 2b). Intriguingly, while the percentage of Th17 cells was drastically reduced in the colon of Junbfl/flCd4-Cre mice (Fig. 3a, c), the expression of Il17a and Il17f was comparable to that in control mice (Fig. 4c). This suggests that cells other than Th17 cells produce high levels of these cytokines (Supplementary Figure 2c).28,29 Indeed, the majority of IL-17A-producing cells was not CD4+TCRβ+ (Th17), but CD4−TCRβ− cells (Supplementary Figure 2d), and the production of IL-17A from the CD4−TCRβ− cells was not different between control and Junbfl/flCd4-Cre mice (Supplementary Figure 2c). We also found that 20–30% of the population was γδT cells (Supplementary Figure 2d), whereas the rest of the population was non-T cells, possibly including group 3 innate lymphoid cells (ILC3s).29 While IL-22 is known to play a protective role in DSS-induced colitis by stimulating production of the cytoprotective proteins Reg3β and Reg3γ from intestinal epithelial cells,30,31 expression of Il22 was not decreased and tended to be increased in the colon of Junbfl/flCd4-Cre mice compared to that of control mice (Fig. 4c). The expression of Reg3b and Reg3g was elevated in the colon of Junbfl/flCd4-Cre mice (Fig. 4c). Thus, the exacerbation of DSS-induced colitis in Junbfl/flCd4-Cre mice is not caused by a defect in IL-22-dependent signaling.
The expression of Th17-associated cytokines is not reduced in the colon of Junbfl/flCd4-Cre mice after DSS administration. a–c Total RNA was extracted from the colon of control or Junbfl/flCd4-Cre mice at day 5 after DSS administration. Gene expression profile was analyzed using microarray analysis (n = 2 mice per each genotype). Heat map of the gene expression is shown (a). The expression of the indicated genes was analyzed by qPCR (b, c). Results are means ± SEMs (n = 5–10 mice per each genotype). *p < 0.05, ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
The number of Treg cells is reduced in Junb fl/fl Cd4-Cre mice
Foxp3+ Treg cells are most abundantly present in the intestines and play a crucial role in the protection against inflammatory bowel diseases including DSS-induced colitis.18 We found that the percentage of Foxp3+ cells in CD4+ T cells was decreased in the colon of Junbfl/flCd4-Cre mice compared to that of control mice at day 5 after DSS administration (Fig. 5a). Reduction of Foxp3+ cells was also observed in the colon, small intestine, thymus, and spleen of Junbfl/flCd4-Cre mice even before DSS administration (Fig. 5b). We found that JunB is required for expression of RORγt in not only Th17 cells (Foxp3− CD4+ T cells)5 but also Foxp3+ Treg cells (Fig. 5c, d). The RORγt+ Treg subset was reported to represent a stable effector lineage with a potent suppression activity.32 Among molecules involved in effector Treg functions,33 the expression of CD103, ICOS, and CTLA4 was reduced in Treg cells of Junbfl/flCd4-Cre mice compared to those of control mice, while the expression of GITR in Treg cells were comparable between the two groups (Supplementary Figure 3 and Fig. 5e–h). Thus, the exacerbation of DSS-induced colitis in Junbfl/flCd4-Cre mice appear to be due to both a reduction in numbers of Treg cells and impairment of their effector functions. Previous studies have reported that gut microbiota affects the development of Treg cells in the intestine,34,35 raising a possibility that the reduction of Treg cells in Junbfl/flCd4-Cre mice is caused by alteration of gut microbiota. To test this possibility, we compared the composition of gut microbiota between control and Junbfl/flCd4-Cre mice by performing Illumina MiSeq 16 S ribosomal DNA sequencing (Supplementary Figure 4a–c). Comparison of fecal microbiota composition did not show any differences in α-diversity of species evenness (Shannon index) and richness (Number of OTUs), or genotype-dependent clustering of β-diversity between control and Junbfl/flCd4-Cre mice (Supplementary Figure 4d–f). It has been also reported that specific bacteria colonized in the intestine, such as Clostridium and Lactobacillus species, facilitate differentiation of Treg cells.18,19 We analyzed relative abundance of these six Clostridium strains and total Lactobacillus by quantifying the 16 S ribosomal DNA in feces, but their frequency was not significantly reduced in Junbfl/flCd4-Cre mice (Supplementary Figure 4g). These results suggest that reduction of Treg cells is not caused by the alterations of microbiota composition in the colon of Junbfl/flCd4-Cre mice.
The number of Treg cells is reduced in Junbfl/flCd4-Cre mice. a Colonic lamina propria cells were isolated from control or Junbfl/flCd4-Cre mice after administration with DSS for 5 days, and analyzed by flow cytometry. The expression of Foxp3 in FVD506−CD45.2+CD4+TCRβ+ cells (CD4+ T cells) were analyzed. The graph shows the average percentage of Foxp3+ cells among CD4+ T cells. Results are means ± SEMs (n = 6 mice per each genotype). b Cells were isolated from the indicated tissues of untreated mice, and analyzed by flow cytometry. The average percentages of Foxp3+ cells among CD4+ T cells are shown as in (a). Results are means ± SEMs (n = 5–13 mice per each genotype). c Colonic lamina propria cells were isolated from untreated mice, and analyzed by flow cytometry. FVD506−CD45.2+CD4+TCRβ+ cells (CD4+ T cells) are presented. The plots show representative results of flow cytometry. The graph represents the average percentages of RORγt+Foxp3− cells (Th17 cells) in CD4+ T cells. d The histograms show representative results of flow cytometry. The graph represents the average percentages of RORγt+ cells among Foxp3+CD4+ T cells. Results are means ± SEMs (n = 7 mice per each genotype) (c, d). e–h Cells were isolated from the indicated tissues of untreated mice, and analyzed by flow cytometry. Representative results of flow cytometry are shown in Supplementary Figure 3. The average percentages of CD103+ cells among Foxp3+CD4+ T cells (e) and mean fluorescence intensities (MFI) of ICOS (f), CTLA4 (g), and GITR (h) in Foxp3+CD4+ T cells are shown. Results are means ± SEMs (n = 4–8 mice per each genotype). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
Junb-deficient CD4+ T cells are defective in the development of Treg cells due to an impairment of IL-2 signaling
To investigate the mechanisms underlying a decrease of Treg cells in Junbfl/flCd4-Cre mice, we carried out in vitro Treg differentiation experiments using naive CD4+ T cells. While Junb−/− CD4+ T cells normally differentiated to Foxp3+ Treg cells when IL-2 was exogenously supplemented in the culture, they were incapable of efficiently differentiating into Treg cells in the absence of exogenous IL-2 (Fig. 6a, g). Given that IL-2 signaling is essential for the development of Treg cells,20 Junb−/− CD4+ T cells were considered to have a defect in production of IL-2 or sensitive recognition of IL-2, or both. We found that the amount of IL-2 released from Junb−/− CD4+ T cells was significantly reduced compared to control cells (Fig. 6b).
CD4+ T cells lacking Junb exhibit an impairment in production of IL-2 and expression of CD25 under Treg-inducing conditions in vitro. a Naive CD4+ T cells isolated from control or Junbfl/flCd4-Cre mice were cultured for 3 days under Treg-inducing conditions in the absence or presence of recombinant IL-2 (50 ng/ml), and then analyzed by flow cytometry. b Naive CD4+ T cells isolated from the indicated mice were cultured for 9 h under the Treg-inducing conditions in the absence of exogenous IL-2. The concentrations of IL-2 in the culture supernatant were measured by ELISA. Results are means ± SDs of triplicate samples and are representative of three independent experiments. c Naive CD4+ T cells from the indicated mice were cultured for 3 days under Treg-inducing conditions in the presence of neutralizing anti-IL-2 antibody (20 μg/ml), and then analyzed by flow cytometry. d Mean fluorescence intensities (MFIs) of CD25 are shown. Pooled results of 4 or 6 independent experiments performed under the conditions in (a, c) are presented. e Naive CD4+ T cells from the indicated mice were cultured for 3 days under the Th0 conditions in the absence or presence of neutralizing anti-IL-2 antibody (20 μg/ml), and then analyzed by flow cytometry. f MFIs of CD25 are shown. Pooled results of 4 independent experiments performed under the conditions in (e) are presented. g Average percentages of Foxp3+ cells under the indicated culture conditions are presented. Results are means ± SEMs (n = 4–6 per each genotype). Representative results of flow cytometry are shown (n = 4–6 mice per each genotype) (a, c, e). h The nucleotide sequence of Il2ra promoter. Potential AP-1 binding sites are boxed. i Naive CD4+ T cells were cultured under Treg-inducing conditions for 3 days, and subjected to the ChIP assay using the indicated antibodies. The precipitated DNA was analyzed by quantitative PCR using primer pairs corresponding to the indicated genomic regions. The enrichment of the precipitated DNA relative to input is shown. Results are means ± SDs of triplicate samples and are representative of at least four independent experiments. Statistical significance was determined by two-tailed Student’s unpaired t test (b, g, i) or two-tailed Student’s paired t test (d, f). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, n.s. not significant
We next investigated whether Junb−/− CD4+ T cells also have an impairment in responsiveness to IL-2. Given that expression of the α-chain of the IL-2 receptor (CD25) is upregulated under Treg-inducing conditions,21,36 we surmised that the upregulation of CD25 expression was impaired in Junb−/− CD4+ T cells. In the absence of exogenous IL-2, the expression of CD25 was lower in Junb−/− CD4+ T cells than control cells under Treg-inducing conditions (Fig. 6a, d). Although the expression of Il2ra is directly regulated by Foxp3,24,25 the expression of CD25 was still lower in Junb−/− CD4+ T cells even when the development to Foxp3+ Treg cells was blocked by a neutralizing antibody against IL-2 (Fig. 6c, d). This suggests that the impairment of CD25 expression in Junb−/− CD4+ T cells caused a defect in differentiation to Treg cells. In contrast to the TGF-β1-containing conditions, the expression of CD25 was not lowered in Junb−/− CD4+ T cells under Th0-polarizing conditions (Fig. 6e, f), substantiating a specific role for JunB in the upregulation of CD25 under Treg-inducing conditions. As the Il2ra promoter contains potential AP-1 binding sites (Fig. 6h), we hypothesized that JunB directly bound to the Il2ra promoter. The chromatin immunoprecipitation (ChIP) assay revealed that JunB specifically bound to the Il2ra promoter (Fig. 6i). These results indicate that JunB directly upregulates the expression of Il2ra to increase sensitivity to IL-2, thereby facilitating Treg differentiation. Together, the reduction of Treg cells in tissues of Junbfl/flCd4-Cre mice is likely to be caused by impairment in both production and recognition of IL-2.
To examine how the absence of JunB affects global gene expression in CD4+ T cells under Treg-inducing conditions in the absence of exogenous IL-2, we performed a microarray analysis (Fig. 7a, b and Supplementary Table 2). In addition to Il2, Il2ra, and Foxp3, we found that the expression of other Treg-related genes, including Ctla4, Socs2, Lgals1, Ccr4, and Gpr83,37 was decreased in Junb-deficient cells (Fig. 7a, c). Downregulated genes in Junb-deficient cells also included Il17a, Il17f, Il9, and Il12rb1, which are known to be regulated by JunB in Th17 cells.5
JunB is crucial for regulation of Treg-associated genes. a Naive CD4+ T cells from control or Junbfl/flCd4-Cre mice were cultured for 3 days under Treg-inducing conditions in the absence of exogenous IL-2. Total RNA was extracted, and gene expression profiles were analyzed by microarray analysis (n = 2 per each genotype). Heat map of highly downregulated genes in Junb-deficient cells is shown. Arrows indicate typical Treg-related genes and known JunB-regulated genes in Th17 cells. b Principal component analysis (PCA) for gene expression profiles of control and Junb-deficient cells. c The expression of the indicated genes was analyzed by qPCR. Results are means ± SEMs (n = 6 mice per each genotype). *p < 0.05, **p < 0.01, ***p < 0.001 (two-tailed Student’s unpaired t test)
Expression of CD25 is reduced in thymic Treg cells of Junb fl/fl Cd4-Cre mice
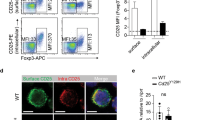
To examine whether JunB-dependent CD25 expression is relevant in vivo, we analyzed the expression of CD25 in tissue-resident Treg cells (Fig. 8a–c). We found that the percentage of CD25+ cells among Foxp3+ cells was markedly reduced in the thymus of Junbfl/fl Cd4-Cre mice compared to that of control mice (Fig. 8b, c). The frequency of Foxp3−CD25+ Treg precursor cells38 tended to be reduced in the thymus of Junbfl/flCd4-Cre mice, although the reduction was not statistically significant (Fig. 8d). The expression of CD25 in Foxp3+ cells was not diminished in the spleen and colon of Junbfl/flCd4-Cre mice (Fig. 8b, c). Given that Treg cells in these peripheral tissues of Junbfl/flCd4-Cre mice were significantly reduced (Fig. 5b), JunB is likely to be involved in upregulation of CD25 expression during Treg development, but is dispensable for maintenance of CD25 expression after Treg differentiation in the spleen and colon. These results indicate that JunB plays an essential role in Treg development by regulating the expression of CD25 to increase a sensitivity to IL-2 in vivo.
JunB is required for maintenance of CD25 expression in thymic Treg cells. a Cells from the indicated tissues of untreated mice were analyzed by flow cytometry. The expression of Foxp3 and CD25 in FVD506−CD45.2+CD4+TCRβ+ cells (CD4+ T cells) was analyzed. b The expression of CD25 in Foxp3+CD4+ T cells was analyzed. Dashed lines indicate signals when isotype control antibody was used instead of anti-CD25 antibody. Representative results of flow cytometry are shown (n = 4–6 mice per each genotype) (a, b). c Average percentages of CD25+ cells among Foxp3+CD4+ T cells are presented. d Average percentages of CD25+Foxp3− cells among CD4+ T cells are presented. Results are means ± SEMs (n = 4–6 mice per each genotype). ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
IL-2 production from T cells is impaired in Junb fl/fl Cd4-Cre mice
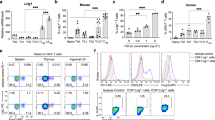
Consistent with a decrease in the production of IL-2 in Junb-deficient CD4+ T cells in vitro (Fig. 6b), the number of IL-2+CD4+ T cells was significantly decreased in the thymus and spleen of Junbfl/flCd4-Cre mice (Supplementary Figure 5a, b and Fig. 9a). In Junbfl/flCd4-Cre mice, Junb is lacking in not only CD4+ T cells but also CD8+ T cells, and IL-2 production in CD8+ T cells (CD4−TCRβ+ cells) also depended on JunB (Supplementary Figure 5a, b and Fig. 9b).
Production of IL-2 in T cells is reduced in Junbfl/flCd4-Cre mice. a, b Cells from the indicated tissues of control or Junbfl/flCd4-Cre mice were analyzed by flow cytometry. Representative results of flow cytometry are shown in Supplementary Figure 5. The average percentages of IL-2+ cells among CD4+ T cells (a) and CD4−T cells (CD8+ T cells) (b) are shown. Results are means ± SEMs (n = 6–8 mice per each genotype). c Schematic diagram for generation of bone marrow (BM) chimera mice. BM cells from wild type (CD45.1) and Junbfl/flCd4-Cre (CD45.2) mice were mixed at a ratio of 1:1, and transferred to a recipient wild-type mouse (CD45.1) that had been irradiated at 8.5 Gy. d, e Eight weeks after the BM transfer, cells from the indicated tissues were analyzed by flow cytometry. The average percentages of Foxp3+ cells among CD4+ T cells (d) and CD4+TCRβ+ cells among total CD45+ cells (e) are shown. Results are means ± SEMs (n = 7 mice). *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
Given that IL-2 is mainly produced by T cells,20 the reduction of Treg cells in Junbfl/flCd4-Cre mice could be due to insufficient supply of IL-2 for the development and maintenance of Treg cells. To elucidate the importance of the intrinsic and extrinsic defects of Junb-deficient CD4+ T cells in the defect in Treg development, we mixed bone marrow cells from CD45.2+ Junbfl/flCd4-Cre mice and CD45.1+ wild-type mice at a ratio of 1:1, and transplanted the mixture into CD45.1+ recipient mice (Fig. 9c). At 8 weeks after bone marrow transfer, the number of Treg cells of CD45.2+ population was still lower in the thymus, spleen and colon compared to those of CD45.1+ population (Fig. 9d). These results suggest that IL-2 provided by wild-type CD4+ T cells was not sufficient to rescue the defect of Junb-deficient CD4+ T cells in Treg development in vivo. We also found that the number of total CD4+ T cells of CD45.2+ population was similarly reduced in the spleen and colon, and to a lesser extent, thymus (Fig. 9e). Given that the number of B cells of CD45.2+ population was comparable to those of CD45.1+ population (Supplementary Figure 6a, b), Junb-deficient CD4+ T cells appeared to have a cell-intrinsic defect in the development and the maintenance, thereby failing to be repopulated in peripheral tissues even in the presence of IL-2 derived from wild-type CD4+ T cells.
IL-2-induced expansion of Treg cells protects Junb fl/fl Cd4-Cre mice from exacerbation of DSS-induced colitis
Injection of super-agonist immune complexes composed of IL-2 and an anti-IL-2 monoclonal antibody (clone JES6-1) induces expansion of Treg cells in wild-type mice.39 Despite the lower responsiveness of Junb-deficient CD4+ T cells to IL-2 (Fig. 6), injection of IL-2-anti-IL-2 Ab complexes caused a robust increase in Treg cells in Junbfl/flCd4-Cre mice (Fig. 10a, b). The expression of CD25 was also highly upregulated after injection of IL-2-anti-IL-2 complexes (Fig. 10a). To investigate the relationship between the decrease in Treg cells and the exacerbation of DSS-induced colitis, we injected IL-2-anti-IL-2 complexes and concurrently administrated DSS to Junbfl/flCd4-Cre mice. Notably, the body weight loss was alleviated by injection of IL-2-anti-IL-2 complexes (Fig. 10c). The reduction of Th17 cells in Junbfl/flCd4-Cre mice was not rescued by injection of IL-2-anti-IL-2 complexes (Fig. 10d, e), which is consistent with the notion that the reduction in Treg cells, but not Th17 cells, is responsible for the exacerbation of DSS-induced colitis in Junbfl/flCd4-Cre mice.
IL-2-induced expansion of Treg cells prevents Junbfl/flCd4-Cre mice from exacerbation of DSS-induced colitis. a, b Control IgG or IL-2-anti-IL-2 complex was injected to Junbfl/flCd4-Cre mice on days 0, 1, and 2, and cells from the indicated tissues were analyzed by flow cytometry on day 5. Representative results of flow cytometry (a) and the average percentages of Foxp3+ cells among CD4+ T cells (b) are shown. Results are means ± SEMs (n = 4 mice per each group). c Junbfl/flCd4-Cre mice were administrated with 2.0% DSS in drinking water for 5 days, and then they were given by normal water in the following days. Control IgG or IL-2-anti-IL-2 complex was injected on days 0, 1, 2, 3, and 4. The average of body weight is shown as percentage relative to the initial value. Results are means ± SEMs (n = 7 or 8 mice per each group). d, e Junbfl/flCd4-Cre mice were treated as in (a), and colonic lamina propria cells were analyzed by flow cytometry on day 5. Representative results of flow cytometry (d) and the average percentages of IL-17A+ cells among CD4+ T cells (e) are shown. Results are means ± SEMs (n = 4 mice per each group). *p < 0.05, **p < 0.01, ****p < 0.0001, n.s. not significant (two-tailed Student’s unpaired t test)
Discussion
In the present study, we demonstrated that CD4+ T cell-specific Junb-deficient mice were more susceptible to DSS-induced colitis because of the impaired development of Treg cells. Junb−/− CD4+ T cells failed to fully express IL-2 and CD25, thus incapable of differentiating into Treg cells unless high concentrations of IL-2 were supplemented in vitro. Expression of IL-2 and Treg-related molecules including CD25 was reduced in CD4+ T cells in Junbfl/flCd4-Cre mice. In a mixed bone marrow transfer experiment, the defect of Junb-deficient CD4+ T cells in Treg development was not rescued in the presence of wild-type CD4+ T cells, indicating an importance of the cell-intrinsic defect of Junb-deficient CD4+ T cells in the decreased number of Treg cells in Junbfl/flCd4-Cre mice. On the other hand, injection of IL-2-anti-IL-2 complexes into Junbfl/flCd4-Cre mice lead to expansion of Treg cells and attenuated DSS-induced colitis. Thus we provide a novel role for JunB in the development of Treg cells through promoting IL-2 signaling.
Although Junbfl/flCd4-Cre mice exhibited a reduction in the number of Treg cells, the influence of the reduction is not manifested in T cell-mediated autoimmune disease conditions because these mice also lacked pathogenic T cells.5,6,7 By employing the DSS-induced colitis model, we demonstrated a protective role of JunB in CD4+ T cells in the inflammatory disease mediated by innate immune cells. Although Treg cells have been shown to attenuate DSS-induced colitis,18,19 the mechanisms underlying the attenuation is not fully understood. One might surmise that the reduction of Treg cells caused a decrease in production of the anti-inflammatory cytokine IL-10 in the colon of Junbfl/flCd4-Cre mice. However, the expression of Il10 in the colon of Junbfl/flCd4-Cre mice was not decreased compared to control mice. A recent study has reported that Treg cells suppress release of IL-23 and IL-1β from intestinal CXCR3+ macrophages, thereby inhibiting aberrant activation of ILC3.40 Given that the expression of Il1b was higher in the colon of Junbfl/flCd4-Cre mice than that of control mice after DSS treatment, activation of the CXCR3+ macrophages might be one of the mechanisms how a decrease in Treg cells exacerbated DSS-induced colitis in Junbfl/flCd4-Cre mice.
A mixed bone marrow transfer experiment revealed that the cell-intrinsic defect of Junb-deficient CD4+ T cells played a crucial role in the impaired development of Treg cells in vivo. In contrast, injection of IL-2-anti-IL-2 complexes induced robust expansion of Treg cells in Junbfl/flCd4-Cre mice. These results are apparently inconsistent with each other. Given that the expression of CD25 was highly upregulated in Junbfl/flCd4-Cre mice after injection of IL-2-anti-IL-2 complexes, one plausible explanation would be that strong signals from IL-2 receptor triggered by IL-2-anti-IL-2 complexes, but not IL-2 supplied from wild-type CD4+ T cells, can overcome the intrinsic defect of Junb-deficient CD4+ T cells. The cell-intrinsic defects of Junb-deficient CD4+ T cells may also include a reduction in expression of CD103, ICOS, and CTLA4, which are important for Treg functions. Impairment of Treg functions might well explain the reduction of Junb-deficient Treg cells compared to wild-type Treg cells in a mixed bone marrow transfer experiment, because co-transferred wild-type cells were dominantly repopulated during homeostatic proliferation after transfer. Because Junb-deficient cells were also decreased at the level of total CD4+ T cells compared to wild-type cells, JunB possibly regulates expression of proteins that are required for expansion and maintenance of multiple subsets of CD4+ T cells.
Among cytokines produced by Th17 cells, IL-17A and IL-22 are shown to play a protective role in DSS-induced colitis,30,41 and these cytokines are also abundantly produced by other cell types, including γδT cells and ILC3 cells.28,29 Consistent with the requirement of the transcription factor RORγt for generation and function of these cells,42,43,44 Rorc-deficient mice are more susceptible to DSS-induced colitis.45 On the other hand, the exacerbation of the colitis in Junbfl/flCd4-Cre mice was not due to the reduced expression of IL-17A and IL-22. Recently it was reported that Th17 cells as well as Treg cells suppress expression of IL-23 in CCR2+ monocytes and monocyte-derived dendritic cells.46 Thus Th17 cells may protect from DSS-induced colitis by mechanisms other than production of IL-17A and IL-22, including attenuation of the CCR2+ myeloid cells.
The expression of CD25 is highly induced under Treg-inducing conditions in which CD4+ T cells are activated through TCR in the presence of TGF-β and IL-2.21,22,23 We here demonstrated that JunB was required for production of IL-2 and CD25 expression in CD4+ T cells during Treg differentiation. Given that Junb−/− CD4+ T cells normally differentiated into Treg cells in the presence of exogenous IL-2, large amounts of IL-2 could circumvent the defect in the upregulation of CD25 in Junb−/− CD4+ T cells through low affinity IL-2 receptor at least in vitro. The decrease of Treg cells in tissues of Junbfl/flCd4-Cre mice was likely to be explained by reduced affinity of Junb-deficient CD4+ T cells to the low levels of IL-2 that are supplied in the microenvironment. Consistent with our previous finding that gene expression profile of Th0 cells was not affected by lack of Junb,5 the expression of CD25 was not significantly diminished in Junb−/− CD4+ T cells under Th0-polarizing conditions. Given that the decrease of CD25 in Junb−/− cells was observed only in the presence of TGF-β, JunB presumably enhances TGF-β signaling to induce the expression of CD25. On the other hand, TGF-β-induced expression of Foxp3 is suppressed by JunB under Th17-polarizing conditions,5,6,7 indicating that JunB differently affects TGF-β signaling in a context-dependent manner.
The percentage of CD25+ cells among Foxp3+ cells was markedly reduced in thymus of Junb-deficient mice. Given that IL-2 signaling is required for not only the development of Treg cells7 but also their maintenance after their development,47 the decrease in thymic Treg cells in Junbfl/flCd4-Cre mice was considered to be caused by an impaired expression of CD25 both during and after Treg development. On the other hand, for Foxp3+ cells in the spleen and colon, the expression of CD25 was not diminished in Junbfl/flCd4-Cre mice, suggesting that Junb is dispensable for maintenance of CD25 expression in these peripheral tissues. Thus the reduction of Treg cells in the spleen and colons is likely to be solely caused by impaired CD25 expression during the course of Treg development. Junb appears to be dispensable for maintenance of not only CD25 but also Foxp3 in periphery, because mice lacking Junb in Foxp3+ cells (Junbfl/flFoxp3Cre mice) did not exhibit reduction of Foxp3+ Treg cells in spleen and lymph nodes.7
In induction of Th17 differentiation, JunB plays a non-redundant role by cooperating with the other AP-1 family protein BATF to directly activate Th17-associated genes.5 In sharp contrast, the reduction of Treg cells in Junbfl/flCd4-Cre mice was not so drastic compared to the marked reduction of Th17 cells,5,6,7 implying that another Jun family member might play a redundant role in IL-2 production and CD25 expression during the development of Treg cells. Although the partner protein of JunB in regulation of CD25 expression is unknown, BATF may form a heterodimer with JunB in regulation of Il2ra expression. Consistently, a recent report showed the involvement of BATF in Treg differentiation.48
Given that deletion of Junb blocks the development of Th17 cells, JunB might be a potential therapeutic target to treat Th17-dependent autoimmune diseases, such as multiple sclerosis, rheumatoid arthritis, and psoriasis.49 However, one of the most serious side effects by blockade of JunB functions might be an impaired differentiation of Treg cells in vivo. As the impairment in differentiation of Junb−/− CD4+ T cells to Treg cells appears to be canceled by the addition of exogenous IL-2 at least in vitro, blockade of JunB functions combined with administration of IL-2 might be an ideal strategy to treat various T cell-dependent and -independent diseases.
During the revision of our manuscript, Koizumi et al. reported a role for JunB in Treg cells.50 The authors showed that JunB participates in maintenance of suppressive function of Treg cells by directly regulating transcription of Treg-related genes, and mice deficient in JunB in Treg cells (Junbfl/flFoxp3Cre mice) developed severe autoimmune diseases. They also demonstrated that the expression of Treg effector molecules is downregulated in Junb-deficient Treg cells, which is consistent with our present study.
Methods
Reagents
The following antibodies used in the present study were obtained from the indicated sources: anti-CD3 (BioLegend, 100302, clone 145-2C11), anti-CD28 (BioLegend, 102102, clone 37.51), anti-IFN-γ (BioLegend, 505812, clone XMG1.2), anti-IL-4 (TONBO biosciences, 40-7041, clone 11B11), anti-IL-2 (BioLegend, 503706, clone JES6-1A12 or BD PharmingenTM, 561061, clone JES6-5H4), anti-CD45.2 (BioLegend, 109824, clone 104), anti-CD45.1 (TONBO, 50-0453, clone A20), anti-CD4 (TONBO, 20-0042, clone RM4-5), anti-TCRβ (TONBO, 30-5961, clone H57-597), anti-TCRγδ (BioLegend, 118105, clone GL3), anti-CD25 (TONBO, 50-0251, clone PC61.5), anti-ICOS (BioLegend, 117405, clone 7E.17G9), anti-CD103 (BioLegend, 121405, clone 2E7), anti-CTLA-4 (BioLegend, 106309, clone UC10-4B9), anti-GITR (BioLegend, 126311, clone DTA-1), anti-IL-17A (BioLegend, 506903, clone TC11-18H10), anti-Foxp3 (eBioscience, 11-5773-82, clone FJK-16s), anti-RORγt (BD Bioscience, 562607, clone Q31-378), anti-JunB (Santa Cruz, N-17, sc-46), and rabbit control IgG (GeneTex, GTX35035). Human TGF-β1 (Peprotech, 100-21), murine IL-2 (Peprotech, 212-12) were purchased from the indicated sources. FVD506 (eBioscience, 65-0866-14), phorbol-12-myristate-13-acetate (PMA) (SIGMA, P8139), ionomycin (SIGMA, I9657), and GoigiPlug (BD Bioscience, 555029) were obtained from the indicated sources.
Mice
Junbfl/fl mice are described previously and backcrossed onto C57BL/6J mice at least 15 generations.5 Mice deficient in Junb specifically in CD4+ T cells (Junbfl/flCd4-Cre mice), intestinal epithelial cells (Junbfl/flVil1-Cre), and myeloid cells (Junbfl/flLyz2Cre) were generated by crossing of Junbfl/fl mice with Cd4-Cre, Vil1-Cre mice, and Lyz2Cre mice, respectively. All Cre driver mice were purchased from the Jackson Lab. C57BL/6-SJL (CD45.1) mice were from Sankyo Lab. All animals were housed and maintained under specific pathogen-free conditions in the animal facility at Toho University School of Medicine. The experimental protocol was approved by the Toho University Administrative Panel for Animal Care (18-44-288 and 19-51-413) and Recombinant DNA (18-54-299, 19-55-299, 18-55-271, and 19-51-410). All efforts were made to minimize the number of the animals and their suffering.
Dextran sulfate sodium (DSS)-induced colitis
For all of the DSS-induced colitis experiments, we used littermate mice that had been co-housed since their birth. Mice were orally given 2% DSS (MW 36,000–50,000, MP Biomedicals Inc.) ad libitum in drinking water for 5 days, followed by regular drinking water without DSS. Body weight was measured every 24 h during the experiment. Mice were sacrificed at the indicated day, and the colon length was measured.
Induction of experimental autoimmune encephalomyelitis (EAE)
EAE was induced as described previously.5 Briefly, mice were subcutaneously immunized with the synthetic MOG35–55 peptide (200 μg/mouse, Scrum Inc.) emulsified in complete Freund’s adjuvant supplemented with Mycobacterium tuberculosis H37RA (500 μg per mouse, Difco). Pertussis toxin (500 ng per mouse, Calbiochem) was intraperitoneally injected to mice on day 0 and day 2. Severity of the disease was monitored over time, and scored as follows: 0, no clinical sign; 1, tail limpness; 2, hind limb weakness; 3, hind limb paralysis; 4, fore limb weakness.
Anti-CD3 antibody-induced intestinal inflammation model
The anti-CD3 agonistic antibody (clone 145-2C11) was made in house and intraperitoneally injected to mice at 0, 48, and 96 h (20 μg per mouse). Body weight was measured every 24 h.
Histological analysis
The colon was fixed in formaldehyde (10%) overnight at room temperature, and then embedded in paraffin blocks. The paraffin-embedded section was stained with hematoxylin and eosin. Pictures were obtained using the BZ-X700 All-in-one microscope (KEYENCE).
Preparation of lamina propria cells from the small intestine and colon
Lamina propria cells were prepared from mouse small intestine and colon as described previously.5 Briefly, after removal of mucosa and epithelia by incubating in the presence of EDTA (1 mM), the intestine was cut into small fragments and digested with collagenase (1 mg/ml, Wako). Cells were filtrated using nylon mesh, and suspended in a 40% Percoll solution (GE Healthcare) and placed on a 80% Percoll solution. After centrifugation for 20 min at 880 × g at room temperature, cells at interface of 40 and 80% Percoll were harvested and analyzed by flow cytometry.
In vitro culture of CD4+ T cells
Naive CD4+ T cells were purified from mouse splenocytes using EasySepTM Mouse Naive CD4 + T Cell Isolation kit according to the manufacturer’s instruction (STEMCELL). Isolated naive CD4+ T cells were cultured in the presence of plate-bound anti-CD3 (5 μg/ml), anti-CD28 (2.5 μg/ml), anti-IFN-γ (5 μg/ml), and anti-IL-4 (5 μg/ml) for Th0-polarizing conditions. For induction of Treg cells, recombinant human TGF-β1 (5 ng/ml) was added to the culture of Th0 conditions. As indicated, cells were cultured in the presence of recombinant murine IL-2 (50 ng/ml) or anti-IL-2 (20 μg/ml). For microarray gene expression experiments, naive CD4+ T cells (CD4+CD62LhighCD44lowCD25− cells) were sorted using FACSAria III cell sorter (BD Biosciences), and cultured for 3 days in the presence of anti-CD3 (1 μg/ml), anti-CD28 (0.5 μg/ml), anti-IFN-γ (5 μg/ml), anti-IL-4 (5 μg/ml), and TGF-β1 (5 ng/ml) without addition of IL-2.
Flow cytometry analysis
For cell surface staining, cells were stained with FVD506 and fluorochrome-conjugated antibodies against cell surface proteins (anti-CD45.1, anti-CD45.2, anti-CD4, anti-TCRβ, anti-CD103, and anti-GITR) in flow cytometry staining buffer (eBioscience). For intracellular cytokine staining, cells were stimulated for 4 h with PMA (50 ng/ml) and ionomycin (500 ng/ml) in the presence of GolgiPlug, and then fixed with intracellular fixation buffer (eBioscience). After permeabilization, cells were stained with anti-IL-17A, anti-IFN-γ, or anti-IL-2. For intracellular staining, cells were fixed with 1 × Foxp3 Fixation/Permeabilization solution (eBioscience), and then stained with anti-Foxp3, anti-CTLA4, or anti-ICOS. Cells were analyzed using LSR Fortessa X-20 cell analyzer (BD Bioscience) and FlowJo (BD Biosciences).
Reverse transcription-quantitative PCR (RT-qPCR)
Total RNA was extracted from the colon using Sepasol®-RNA Super G (Nakalai tesque) or TRI reagent® (Molecular Research Center, Inc.). RNA was reverse-transcribed using ReverTra Ace kit (TOYOBO). cDNA was used as a template for quantitative PCR using FAST SYBRTM Green Master Mix (Applied biosystems) on 7500 Fast (Applied biosystems). Expression of the housekeeping gene Hprt was used as internal control for normalizing expression levels of target genes. Sequences of the primers used in the study are listed in Supplementary Table 3.
Transcriptome analysis using microarray
After administration of DSS (2%) for 5 days, the control (Junbfl/fl) and Junbfl/flCd4-Cre mice were sacrificed, and total RNA was prepared from the colon using miRNeasy Micro Kit according to the manufacture’s instruction (Qiagen). For analysis of in vitro cultured cells, naive CD4+ T cells from control and Junbfl/flCd4-Cre mice were cultured for 3 days under Treg-inducing conditions in the absence of exogenous IL-2, and total RNA was extracted using the same kit. Total RNA was amplified and labeled with Cyanine 3 (Cy3) using Agilent Low Input Quick Amp Labeling Kit, one-color following the manufacturer’s instructions (Agilent Technologies). Briefly, after total RNA was reverse-transcribed using a poly(dT)-T7 promoter primer, the resultant cDNA was used as a template for in vitro transcription to generate Cy3-labeled cRNA. The fluorescent cRNA was fragmented, and hybridized at 65 °C for 17 h to an Agilent SurePrint G3 Mouse GE v2 8 × 60 K Microarray (Design ID: 074809). After washing, the microarray was scanned using an Agilent DNA microarray scanner. Intensity values of each scanned feature were quantified using Agilent feature extraction software version 11.5.1.1, which performs background subtractions. Normalization was carried out using Agilent GeneSpring software version 14.8 (per chip:normalization to 75 percentile shift), and deposited to the National Center for Biotechnology Information as GEO accession numbers GSE120814 and GSE131292. Clustering and depiction of a heat map was performed using R.51
Analysis of bacterial DNA in Feces
DNA was extracted from feces using QIAampⓇ PowerFecalⓇ DNA Kit (QIAGEN) according to the manufacturer’s instruction. Relative content of the specific bacteria was analyzed by qPCR using primer pairs listed in Supplementary Table 3, and normalized by the amount of total bacteria. Deep sequencing and statistical analysis were performed as described previously.52 Briefly, the library targeting the 16S rRNA V3 and V4 regions was prepared using the primer set consisting of 341 F (5′-CCTACGGGNGGCWGCAG-3′) and 806 R (5′-GACTACHVGGGTATCTAATCC-3′) according to the manufacturer’s protocol (http://jp.support.illumina.com/content/dam/illumina-support/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf) for sequencing on Illumina MiSeq (Illumina, San Diego, CA, USA). The library was sequenced with 2 × 300-bp paired-end reads using MiSeq v3 reagent kits (Illumina, San Diego, CA, USA), resulting in a total of 1,611,524 reads with a mean of 134,294 ± 12,918 sequences per sample. Sequence data were deposited to DDBJ under the accession number DRA007424. The sequencing data were processed using CLC Genomics Workbench 11.0.1 and CLC Microbial Genomics Module 3.5 (Qiagen, Redwood City, CA USA). After the overlapping paired-end reads were merged and trimmed, and chimeric reads were filtered using default parameters, the remaining reads were clustered into operational taxonomy units (OTUs) with 97% identity using the Greengenes database (version 13_5) as the reference.53 A total of 9344 bacterial OTUs were detected. Alpha diversity was calculated as the number of observed OTUs, Shannon entropy.54 Beta diversity was measured as an unweighted UniFrac distance based on the OTU table,55 and the relative abundance of the taxa was calculated from an unrarefied OTU table.
Chromatin immunoprecipitation (ChIP)
ChIP assays were essentially performed as previously described.5 Briefly, cells were fixed with formaldehyde (1.0%) for 10 min at room temperature, and subjected to sonication for fragmentation of DNA. Fragmented chromatin was immunoprecipitated using 2 μg of anti-JunB or rabbit control IgG. After de-crosslinking, precipitated DNA was analyzed by quantitative PCR using FAST SYBRTM Green Master Mix (Applied biosystems) on 7500 Fast (Applied biosystems). The amount of the precipitated DNA was normalized by that of input DNA. Sequences of the primers used in the study are listed in Supplementary Table 3.
Enzyme-linked immunosorbent assay (ELISA)
Protein concentrations of IL-2 in the culture supernatant were measured using Mouse IL-2 Uncoated ELISA kit according to the manufacturer’s instruction (ThermoFisher).
Mixed bone marrow transfer experiment
Bone marrow cells from wild-type (CD45.1) and Junbfl/flCd4-Cre (CD45.2) mice (each 5 × 106 cells) were mixed, and transferred to a recipient wild-type mouse (CD45.1) that had been irradiated at 8.5 Gy. Eight weeks later, the bone marrow chimera mice were sacrificed, and thymocytes, splenocytes, and colonic lamina propria cells were analyzed by flow cytometry.
Induction of Treg cell expansion by injection of IL-2-anti-IL-2 mAb complex
Recombinant murine IL-2 (1 μg, Peprotech, 212-12) and anti-IL-2 mAb (5 μg, BioXCell, BE0043) were mixed, and incubated for 30 min at 37 °C. The resultant IL-2-anti-IL-2 immune complexes were i.p. injected to Junbfl/flCd4-Cre mice. In control experiments, purified rat IgG2a κ isotype antibody (5 μg, BD Bioscience, 553927) was injected. For analysis of expansion of Treg cells in tissues, the IL-2-anti-IL-2 Ab complexes were injected on days 0, 1, and 2, and mice were sacrificed on day 5.
Statistical analysis
Statistical analysis was performed with two-tailed unpaired Student’s t test unless specified otherwise. p values of < 0.05 were considered to be significant.
References
Schorpp-Kistner, M., Wang, Z. Q., Angel, P. & Wagner, E. F. JunB is essential for mammalian placentation. EMBO J. 18, 934–948 (1999).
Passegue, E., Jochum, W., Schorpp-Kistner, M., Mohle-Steinlein, U. & Wagner, E. F. Chronic myeloid leukemia with increased granulocyte progenitors in mice lacking junB expression in the myeloid lineage. Cell 104, 21–32 (2001).
Kenner, L. et al. Mice lacking JunB are osteopenic due to cell-autonomous osteoblast and osteoclast defects. J. Cell Biol. 164, 613–623 (2004).
Zenz, R. et al. Psoriasis-like skin disease and arthritis caused by inducible epidermal deletion of Jun proteins. Nature 437, 369–375 (2005).
Yamazaki, S. et al. The AP-1 transcription factor JunB is required for Th17 cell differentiation. Sci. Rep. 7, 17402 (2017).
Hasan, Z. et al. JunB is essential for IL-23-dependent pathogenicity of Th17 cells. Nat. Commun. 8, 15628 (2017).
Carr, T. M., Wheaton, J. D., Houtz, G. M. & Ciofani, M. JunB promotes Th17 cell identity and restrains alternative CD4( + ) T-cell programs during inflammation. Nat. Commun. 8, 301 (2017).
Abraham, C. & Cho, J. H. Inflammatory bowel disease. New Engl. J. Med. 361, 2066–2078 (2009).
Kaser, A., Zeissig, S. & Blumberg, R. S. Inflammatory bowel disease. Annu. Rev. Immunol. 28, 573–621 (2010).
Maloy, K. J. & Powrie, F. Intestinal homeostasis and its breakdown in inflammatory bowel disease. Nature 474, 298–306 (2011).
Okayasu, I. et al. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology 98, 694–702 (1990).
Chassaing, B., Aitken, J. D., Malleshappa, M. & Vijay-Kumar, M. Dextran sulfate sodium (DSS)-induced colitis in mice. Curr. Protoc. Immunol. 104, https://doi.org/10.1002/0471142735.im1525s104 (2014).
Dieleman, L. A. et al. Dextran sulfate sodium-induced colitis occurs in severe combined immunodeficient mice. Gastroenterology 107, 1643–1652 (1994).
Krieglstein, C. F. et al. Collagen-binding integrin alpha1beta1 regulates intestinal inflammation in experimental colitis. J. Clin. Investig. 110, 1773–1782 (2002).
Kitagawa, Y. & Sakaguchi, S. Molecular control of regulatory T cell development and function. Curr. Opin. Immunol. 49, 64–70 (2017).
Li, M. O. & Rudensky, A. Y. T cell receptor signalling in the control of regulatory T cell differentiation and function. Nat. Rev. Immunol. 16, 220–233 (2016).
Tanoue, T., Atarashi, K. & Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 16, 295–309 (2016).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011).
Tang, C. et al. Inhibition of dectin-1 signaling ameliorates colitis by inducing lactobacillus-mediated regulatory T cell expansion in the intestine. Cell Host Microbe 18, 183–197 (2015).
Malek, T. R. The biology of interleukin-2. Annu. Rev. Immunol. 26, 453–479 (2008).
Kim, H. P., Kelly, J. & Leonard, W. J. The basis for IL-2-induced IL-2 receptor alpha chain gene regulation: importance of two widely separated IL-2 response elements. Immunity 15, 159–172 (2001).
Kim, H. P. & Leonard, W. J. The basis for TCR-mediated regulation of the IL-2 receptor alpha chain gene: role of widely separated regulatory elements. EMBO J. 21, 3051–3059 (2002).
Kim, H. P., Kim, B. G., Letterio, J. & Leonard, W. J. Smad-dependent cooperative regulation of interleukin 2 receptor alpha chain gene expression by T cell receptor and transforming growth factor-beta. J. Biol. Chem. 280, 34042–34047 (2005).
Marson, A. et al. Foxp3 occupancy and regulation of key target genes during T-cell stimulation. Nature 445, 931–935 (2007).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Merger, M. et al. Defining the roles of perforin, Fas/FasL, and tumour necrosis factor alpha in T cell induced mucosal damage in the mouse intestine. Gut 51, 155–163 (2002).
Chatenoud, L. & Bluestone, J. A. CD3-specific antibodies: a portal to the treatment of autoimmunity. Nat. Rev. Immunol. 7, 622–632 (2007).
Roark, C. L., Simonian, P. L., Fontenot, A. P., Born, W. K. & O’Brien, R. L. gammadelta T cells: an important source of IL-17. Curr. Opin. Immunol. 20, 353–357 (2008).
Killig, M., Glatzer, T. & Romagnani, C. Recognition strategies of group 3 innate lymphoid cells. Front. Immunol. 5, 142 (2014).
Zenewicz, L. A. et al. Innate and adaptive interleukin-22 protects mice from inflammatory bowel disease. Immunity 29, 947–957 (2008).
Zheng, Y. et al. Interleukin-22 mediates early host defense against attaching and effacing bacterial pathogens. Nat. Med. 14, 282–289 (2008).
Yang, B. H. et al. Foxp3( + ) T cells expressing RORgammat represent a stable regulatory T-cell effector lineage with enhanced suppressive capacity during intestinal inflammation. Mucosal Immunol. 9, 444–457 (2016).
Wing, K. et al. CTLA-4 control over Foxp3 + regulatory T cell function. Science 322, 271–275 (2008).
Strauch, U. G. et al. Influence of intestinal bacteria on induction of regulatory T cells: lessons from a transfer model of colitis. Gut 54, 1546–1552 (2005).
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008).
Waldmann, T. A. The multi-subunit interleukin-2 receptor. Annu. Rev. Biochem. 58, 875–911 (1989).
Sugimoto, N. et al. Foxp3-dependent and -independent molecules specific for CD25 + CD4 + natural regulatory T cells revealed by DNA microarray analysis. Int. Immunol. 18, 1197–1209 (2006).
Lio, C. W. & Hsieh, C. S. A two-step process for thymic regulatory T cell development. Immunity 28, 100–111 (2008).
Webster, K. E. et al. In vivo expansion of T reg cells with IL-2-mAb complexes: induction of resistance to EAE and long-term acceptance of islet allografts without immunosuppression. J. Exp. Med. 206, 751–760 (2009).
Bauche, D. et al. LAG3( + ) regulatory t cells restrain interleukin-23-producing CX3CR1( + ) gut-resident macrophages during group 3 innate lymphoid cell-driven colitis. Immunity 49, 342–352 e345 (2018).
Ogawa, A., Andoh, A., Araki, Y., Bamba, T. & Fujiyama, Y. Neutralization of interleukin-17 aggravates dextran sulfate sodium-induced colitis in mice. Clin. Immunol. 110, 55–62 (2004).
Ivanov, I. I. et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17 + T helper cells. Cell 126, 1121–1133 (2006).
Sanos, S. L. et al. RORgammat and commensal microflora are required for the differentiation of mucosal interleukin 22-producing NKp46 + cells. Nat. Immunol. 10, 83–91 (2009).
Shibata, K. et al. Notch-Hes1 pathway is required for the development of IL-17-producing gammadelta T cells. Blood 118, 586–593 (2011).
Lochner, M. et al. Microbiota-induced tertiary lymphoid tissues aggravate inflammatory disease in the absence of RORgamma t and LTi cells. J. Exp. Med. 208, 125–134 (2011).
Mao, K. et al. Innate and adaptive lymphocytes sequentially shape the gut microbiota and lipid metabolism. Nature 554, 255–259 (2018).
Chinen, T. et al. An essential role for the IL-2 receptor in Treg cell function. Nat. Immunol. 17, 1322–1333 (2016).
Hayatsu, N. et al. Analyses of a mutant Foxp3 allele reveal BATF as a critical transcription factor in the differentiation and accumulation of tissue regulatory T cells. Immunity 47, 268–283 e269 (2017).
Maddur, M. S., Miossec, P., Kaveri, S. V. & Bayry, J. Th17 cells: biology, pathogenesis of autoimmune and inflammatory diseases, and therapeutic strategies. Am. J. Pathol. 181, 8–18 (2012).
Koizumi, S. I. et al. JunB regulates homeostasis and suppressive functions of effector regulatory T cells. Nat. Commun. 9, 5344 (2018).
Team, R. C. A language and Environment for Statistical Computing. (R Foundation for Statistical Computing, Vienna, Austria, 2017).
Fukui, Y., Aoki, K., Ishii, Y. & Tateda, K. The palatine tonsil bacteriome, but not the mycobiome, is altered in HIV infection. BMC Microbiol 18, 127 (2018).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ. Microbiol. 72, 5069–5072 (2006).
Shannon, C. E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011).
Acknowledgements
This work was supported by grants from a Grant-in Aid for Scientific Research (C) 16K08747, 19K07516 (to S.Y.), (B) 17H04069 (to H.N.), 15H03103 (to S.E.), Young Scientists (B) 17K15626 (to T.N.) and Challenging Exploratory Research 17K19533 (to H.N.) from Japan Society for the Promotion of Science (JSPS), Scientific Research on Innovative Areas 26110003 (to H.N.), the Japan Agency for Medical Research and Development (AMED) through AMED-CREST with a grant number JP19gm1210002 (to H.N.), Private University Research Branding project from the MEXT (Ministry of Education, Culture, Sports, Science and Technology), Japan (to H.N.), the Research Promotion Grant from Toho University Graduate School of Medicine No.17-02 (to S.Y.), the Project Research Grant No. 17-34 and No. 18-14 of Toho University School of Medicine (to T.K.), and GSK Japan Research Grant 2018 (to T.K.).
Author information
Authors and Affiliations
Contributions
T.K., S.Y. and H.N designed the experiments; T.K., S.Y., K.A. and A.S. performed the experiments; T.K., S.Y., Y.F., K.A., T.N., T.M., S. Katagiri, A.S., S. Kimura., K.T., H.K. and H.N. analyzed data; H.Y. provided antibody; H.S. and S.E. provided mice; T.K., S.Y. and H.N. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katagiri, T., Yamazaki, S., Fukui, Y. et al. JunB plays a crucial role in development of regulatory T cells by promoting IL-2 signaling. Mucosal Immunol 12, 1104–1117 (2019). https://doi.org/10.1038/s41385-019-0182-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-019-0182-0
This article is cited by
-
Single-cell chromatin accessibility and transcriptomic characterization of Behcet’s disease
Communications Biology (2023)
-
Identification of differentially expressed genes and signaling pathways with Candida infection by bioinformatics analysis
European Journal of Medical Research (2022)
-
IκBζ controls IL-17-triggered gene expression program in intestinal epithelial cells that restricts colonization of SFB and prevents Th17-associated pathologies
Mucosal Immunology (2022)
-
Prediction of disease flare by biomarkers after discontinuing biologics in patients with rheumatoid arthritis achieving stringent remission
Scientific Reports (2021)