Abstract
While 1–2% of individuals meet the criteria for a clinical diagnosis of obsessive-compulsive disorder (OCD), many more (~13–38%) experience subclinical obsessive-compulsive symptoms (OCS) during their life. To characterize the genetic underpinnings of OCS and its genetic relationship to OCD, we conducted the largest genome-wide association study (GWAS) meta-analysis of parent- or self-reported OCS to date (N = 33,943 with complete phenotypic and genome-wide data), combining the results from seven large-scale population-based cohorts from Sweden, the Netherlands, England, and Canada (including six twin cohorts and one cohort of unrelated individuals). We found no genome-wide significant associations at the single-nucleotide polymorphism (SNP) or gene-level, but a polygenic risk score (PRS) based on the OCD GWAS previously published by the Psychiatric Genetics Consortium (PGC-OCD) was significantly associated with OCS (Pfixed = 3.06 × 10−5). Also, one curated gene set (Mootha Gluconeogenesis) reached Bonferroni-corrected significance (Ngenes = 28, Beta = 0.79, SE = 0.16, Pbon = 0.008). Expression of genes in this set is high at sites of insulin mediated glucose disposal. Dysregulated insulin signaling in the etiology of OCS has been suggested by a previous study describing a genetic overlap of OCS with insulin signaling-related traits in children and adolescents. We report a SNP heritability of 4.1% (P = 0.0044) in the meta-analyzed GWAS, and heritability estimates based on the twin cohorts of 33–43%. Genetic correlation analysis showed that OCS were most strongly associated with OCD (rG = 0.72, p = 0.0007) among all tested psychiatric disorders (N = 11). Of all 97 tested phenotypes, 24 showed a significant genetic correlation with OCS, and 66 traits showed concordant directions of effect with OCS and OCD. OCS have a significant polygenic contribution and share genetic risk with diagnosed OCD, supporting the hypothesis that OCD represents the extreme end of widely distributed OCS in the population.
Similar content being viewed by others
Introduction
Obsessive-compulsive disorder (OCD) is a common and impairing disorder characterized by persistent, intrusive thoughts and/or repetitive, ritualized behaviors. OCD is a heritable condition with an estimated heritability of 47% in a large twin study [1] and a heritability based on common single nucleotide polymorphisms (SNPs) of 16–28% [2, 3]. However, replicated specific genetic risk factors for OCD have yet to be identified. Several disorders co-occur with OCD such as anxiety disorders, mood disorders, anorexia nervosa (AN), tics, among others [4,5,6] and these disorders share genetic risk with OCD [7, 8].
For many psychiatric disorders, including OCD, it is thought that their genetic risk is continuously distributed in the general population, contributing to varying levels of symptom expression [9]. This is in line with the observation that obsessive-compulsive symptoms (OCS) are relatively common in the population. For example, 13 to 38% [10] of all adult individuals experience OCS, with even higher rates in younger individuals [11, 12], but only 1 to 2% meet the criteria for a clinical diagnosis of OCD13. Nevertheless, these subclinical symptoms can result in substantial distress and interference, even in individuals not meeting diagnostic criteria for the disorder [14].
Studies showing a shared genetic risk between OCS and diagnosed OCD support the hypothesis that clinical OCD may represent the extreme end of a continuous distribution of symptoms [12, 15] and that by considering sub-clinical OCS data we can increase the population available for study [16]. Like clinical OCD, OCS measured as a quantitative trait are heritable, with estimates of 30–74% from twin studies (total heritability; [17,18,19,20,21]) and a SNP-based heritability of 7–16% from genome-wide association studies (GWAS; [12, 15, 22]). Studies of quantitative OCS have also identified genome-wide significant variants. A study by den Braber et al. [22] first reported a genome-wide significant SNP (rs8100480) in MEF2BNB for OCS in the Netherlands Twin Register (NTR; N = 6931), although this was not replicated in a relatively small clinical sample of patients with OCD. When OCS in the NTR sample were meta-analyzed with diagnosed OCD in the Psychiatric Genomics Consortium (PGC) sample (N = 17,992), no genome-wide significant variants were identified [12]. A study by Burton et al. [15] reported a genome-wide significant hit (rs7856850) in PTPRD for OCS in the Spit for Science sample (N = 5018), which was also associated with OCD in a meta-analysis of independent clinical OCD and control samples (N = 11,980). Together these studies suggest that cohort- and community-based samples may be useful for identifying genetic risk for not only OCS but also OCD.
Here we present the results of the largest GWAS meta-analysis of OCS to date, combining the results from various large-scale population-based cohorts from Sweden, the Netherlands, England, and Canada that assessed OCS with a variety of questionnaires. To further characterize our GWAS results, we conducted gene-based and gene-set analyses, as well as genetic correlation analyses with 97 other traits. Further, polygenic risk score (PRS) analysis allowed us to assess the probabilistic susceptibility of OCS using the combined risk measure of variants associated with educational attainment and several psychiatric disorders that often co-occur and genetically correlate with OCD, such as depression (DEP; 15–41% comorbidity rate [23,24,25]), schizophrenia (SCZ; 8–26% comorbidity rate [26]), autism spectrum disorder (ASD; 17% comorbidity rate [27]), and attention-deficit hyperactivity disorder (ADHD; 6–21% comorbidity rate [23]). We further assessed the validity of using OCS in population-based samples as a proxy for clinical OCD diagnosis by comparing the OCS GWAS to the latest GWAS of OCD [2]. We assessed the association of the OCD polygenic risk score (PRS) with OCS in our samples and compared the genetic correlation patterns of OCS and OCD with other traits and disorders.
Methods
Cohorts & obsessive-compulsive symptom measures
Individuals included in this study stem from seven different European-ancestry cohorts, including four cohorts from the Swedish Twin Registry (STR; [28,29,30]), namely CATSS18 [31], CATSS24 [31], STAGE, YATSS, and one each from the Netherlands Twin Register (NTR; [32]), Spit for Science (SfS; [33, 34]), and TwinsUK [35]. The cohorts are predominantly population-based twin cohorts, except SfS, with a mean age between 10 and 57 years (see Table 1). CATSS is a prospective, longitudinal study of all twins born in Sweden since 1992. Here, we used data measured at age 18 (CATSS18), and/or age 24 (CATSS24), selecting only one measurement time point per individual (preferring the measurement at age 24 over age 18 if both measurements were completed as the CATSS24 cohort employed the Obsessive-Compulsive Inventory Revised (OCI-R) which was also used by other cohorts). Data from NTR [12] and SfS [15] were included in previous GWASs. See Supplementary Material for more detailed cohort descriptions.
Several questionnaires were used to assess OCS across the cohorts. STR-CATSS18 employed the Brief Obsessive-Compulsive Scale (BOCS; [36]), STR-STAGE used a seven-item OCS instrument [1], while STR-CATSS24, STR-YATSS, and TwinsUK employed the OCI-R [37], excluding the hoarding and neutralizing sub-scales. In NTR, the Padua Inventory Revised was used [38, 39] in the form of the 12-item abbreviated and Dutch translated version [40] excluding the rumination items, leaving 9 items on checking, washing, precision, and intrusive thoughts. In SfS, parent- or self-reported obsessive-compulsive traits within the last 6 months of visiting the Ontario Science Center were assessed using 19 items from the Toronto Obsessive Compulsive Scale (TOCS), a 21-item questionnaire described elsewhere [15, 41, 42]. Two items related to hoarding were removed. To ensure reliable and valid symptom reporting, SfS participants below 12 years of age with self-reported OCS and above 16 years of age with parent-reported OCS were excluded (see Supplementary Table S1 for cohort-specific details on OCS questionnaires).
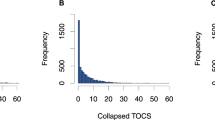
For all cohorts, individuals with one or more missing items were excluded and items were summed and standardized into a Z-score. Distributions of the raw obsessive compulsive item scores are shown in Supplementary Figs. S1–S9. The distributions of the Z-transformed sum scores are shown in Supplementary Figs. S10–S13.
Genome-wide association analysis
All participants were genotyped on SNP-arrays using DNA from saliva or blood. One part of the STR-CATSS samples was genotyped on the PsychChip genotyping array (N = 5683), another part was genotyped on the GSA genotyping array (N = 412). For the GWAS analyses STR-CATSS cohorts (CATSS18, CATSS24) were pooled over each genotyping platform (GSA, PsychChip), forming two separate CATSS datasets (STR-CATSS-GSA and STR-CATSS-PC). Each of the seven datasets (STR-CATSS-GSA, STR-CATSS-PC, STR-YATSS, STR-STAGE, NTR, SfS, and TwinsUK) underwent stringent quality-control (QC), including the removal of non-European ancestry outliers based on PCA and imputation using the Haplotype Reference Consortium (HRC; STR, NTR) or the 1000G (SfS, TwinsUK) reference sets (see the Supplementary Material for more detailed information). Together, all cohorts comprised 33,943 individuals (STR: N = 16,888, NTR: N = 8550, SfS: N = 5171, and TwinsUK: N = 3334) with complete phenotypic and genotypic information (see Table 1 and Supplementary Material for a more detailed description of each cohort).
We used GCTA-fastGWA [43] to perform a mixed-linear-model GWAS within each cohort separately. GCTA-fastGWA controls for population stratification by principal components (PCs) and for relatedness by a SNP-derived genetic relationship matrix (GRM). In STR, NTR, and TwinsUK a sparse family GRM was defined, and each GWAS analysis included the first 10 genetic PCs, sex, age, age squared, and genotyping batches as covariates. In a sparse GRM, all off-diagonal values below 0.05 are set to 0 (default), thereby capturing the same proportion of phenotypic variance as by pedigree-relatedness and accounting for the close relatedness of individuals in the data. In SfS, analyses were performed on unrelated individuals. For sibling pairs, the first enrolled sibling from each family was selected for further analysis (see Table 1 for the number of siblings included). Siblings were removed in this cohort as the high genetic resemblance between them may dominate the genetic variation covered and adjusted for by the GRM, and with only few sibling pairs this adds to the uncertainty of the estimate. Here, the GCTA-fastGWA linear mixed model was performed using a full GRM and sex, age, respondent (parent vs. child reporting), genotyping array type, PCs 1–3, and projected PCs 1–3 (see supplement for details) as covariates.
For each of the seven resulting GWAS summary statistics, variants were filtered on minor allele frequency (MAF) > 1%, and imputation-quality (INFO) score >0.8. For strand ambiguous (A/T and C/G) SNPs, those with a MAF ≥ 0.4 were removed, while the frequencies of those with a MAF < 0.4 were compared to frequencies in the HRC reference [44]. Same strand orientation was assumed if the frequencies matched (i.e., the minor allele was the same in both data sets). All SNPs with matching frequencies were retained while mismatched SNPs (i.e., the minor allele was different in either dataset), assumed to be reported on different strands, were flipped according to the orientation reported in the HRC reference. Following removal of poorly genotyped SNPs, all datasets were aligned to the HRC-reference.
With these harmonized datasets, we conducted an inverse variance weighted meta-analysis utilizing METAL [45], a tool included within the Rapid Imputation for COnsortias PIpeLIne (Ricopili; [46]). To identify any residual population stratification or systematic technical artifacts, we inspected the genomic control factor (Lambda and Lambda1000). The genome-wide significance threshold was set to p < 5 × 10−8.
Gene-based analyses (MAGMA/FUMA)
We performed gene-analysis and gene-set analysis using Multi-marker Analysis of GenoMic Annotation (MAGMA) [47] v1.08 as implemented in Functional Mapping and Annotation of Genome-Wide Association Studies (FUMA; [48]) v1.3.7. To test genetic associations at the gene level for the combined effect of SNPs in or near protein coding genes, we applied default settings (SNP-wise model for gene analysis and competitive model for gene-set analysis). Gene-based p-values were computed by mapping SNPs to their corresponding gene(s) based on their position in the genome. Positional mapping was based on ANNOVAR annotations, and the maximum distance between SNPs and genes was set to 10 kb (default). A multiple regression model was employed while accounting for linkage disequilibrium (LD) between the markers. The 1000 Genomes phase 3 reference panel [49], excluding the MHC region, was used to adjust for gene size and LD across SNPs. Using the result of the gene-based analysis (gene level p-values), competitive gene-set analysis was performed with default parameters: 15,496 gene sets were tested for association. Gene sets were obtained from MsigDB v7.0 (see www.gsea-msigdb.org for details), including ‘Curated gene sets’ consisting of nine data resources including KEGG, Reactome, and BioCarta, and ‘GO terms’ consisting of three categories (biological processes, cellular components, and molecular functions).
Heritability and cross-disorder analyses
Heritability estimates
Heritability estimates of each cohort were extracted from the GCTA association output. GCTA uses the restricted maximum likelihood (REML) approach [50] to estimate heritability in the GRM that is supplied to correct for relatedness in the linear association test. For the twin cohorts (STR, NTR, and TwinsUK), this means that heritability was based on the sparse GRM. For SfS, with no related individuals, the heritability was based on the full GRM. For all heritability estimates, the same covariates as in the GWAS analyses were used. We also calculated the SNP-based heritability of the OCS meta-analysis using LD score regression (LDSC; [51]). LDSC bases its calculation of SNP-based heritability on the estimated slope from the regression of the SNP effect from the GWAS on the LD score.
Cross-trait genetic correlations
With LDSC [51] we calculated genetic correlations between OCS and 97 traits, including psychiatric, substance use, cognition & socio-economic status, personality, neurological, autoimmune, cardiovascular, anthropomorphic, and fertility phenotypes (see Supplementary Table S4 for a list of the source GWASs used). 15 of the included traits pertain to neuroticism and include the neuroticism sum score, worry- and depressive subclusters, as well as individual neuroticism items. The genetic correlation is based on the estimated slope from the regression of the product of Z-scores from the two GWASs of interest on the LD score. It represents the genetic correlation between two traits based on all polygenic effects captured by the included SNPs. Because imputation quality is correlated with LD score, and low imputation quality generally yields lower test statistics, imputation quality is a confounder for LD score regression. We therefore filtered on INFO > 0.9, if INFO was available, and MAF > 0.01. The SNPs from the European HapMap3 [52] were used as a reference. We further compared the genetic correlation patterns between OCS and OCD with all other 96 traits, to identify concordant and discordant patterns. OCD results were based on the publicly available summary statistics from the PGC [2].
Cross-phenotype polygenic risk score analyses
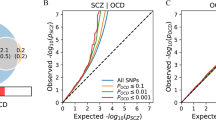
To further explore the genetic relationship between OCS in each of our datasets and other (psychiatric) phenotypes, we calculated polygenic risk-scores (PRS) based on large-scale GWAS summary statistics of OCD (Ncases = 2688, Ncontrols = 7037; [2]), DEP (Ncases = 170,756, Ncontrols = 329,443; excluding 23andMe [53]), SCZ (Ncases = 53,386, Ncontrols = 77,258; [54]), ASD (Ncases = 18,381, Ncontrols = 27,969; [55]), ADHD (Ncases = 19,099, Ncontrols = 34,194; [56]), and educational attainment (EA; Ntotal = 245,621; [57]) using PRSice2 [58] and evaluated their association with OCS in our cohorts (STR-CATSS-GSA, STR-CATSS-PC, STR-STAGE, STR-YATSS, NTR, SfS, and TwinsUK). We pre-selected p-value thresholds based on the best performing thresholds reported in the primary publications (EA: P = 1; ADHD, ASD, OCD, SCZ: P = 0.1; DEP: P = 0.5). The PRS scores were calculated as the weighted sum of the risk allele dosages.
For STR, NTR, and TwinsUK, we employed a generalized estimating equation (GEE) [59] in R to evaluate the relationship between the PRS scores and OCS scores in each cohort. The GEE analysis takes into account the resemblance within clusters, accounting for the relatedness in the datasets. Robust standard errors (sandwich-corrected) and Z-scores are reported. For SfS we applied linear regression, as conducted within the PRSice2 pipeline, to evaluate the relationship between PRS and OCS. Contributions of the PRSs were measured through comparison of the R² of the full model (including the PRS, and covariates) minus the null model (including only covariates). The same covariates that were previously included in the respective GWASs were used. We combined the PRS estimates across all target datasets using an inverse variance weighted meta-analysis using the metagen package in R. Cochran’s Q test [60] and Higgin’s I² [61, 62] were used to examine a possible heterogeneity in PRS estimates across the cohorts. Q is calculated as the weighted sum of the squared differences between individual cohort effects and the pooled effect across cohorts, with the weights being those used in the pooling method. The I² statistic describes the percentage of variation across studies that is due to heterogeneity rather than sample variation and does, unlike Q, not inherently depend on the number of measures included in the meta-analysis. We calculated a fixed effects model to evaluate the association of each PRS with OCS, regardless of observed heterogeneity. We further calculated a random effects model if there was substantial observed heterogeneity across study sites (I² > 0.5 and/or PQ < 0.05).
Comparability of cohorts
To identify if the summary statistics from any of the included cohorts substantially deviated from the rest, we performed leave-one-out (LOO) GWAS meta-analyses and subsequently used those datasets to conduct a set of sensitivity analyses. First, we performed sign-test analyses on the top SNPs (inclusion threshold of p = 0.0001, p = 0.00001, and p = 0.000001) using the replication module of the RICOPILI pipeline. Sign-tests allow for quantification of the number of genomic regions that are independent across the different p-value thresholds and identification of how many genomic regions within the replication study have the same direction of effect as the discovery. The output (in the form of a ratio) provides an estimate of the percentage of genomic regions with the same direction of effect between any two datasets. A sign-test is a binomial test with the null hypothesis = 0.5, with a ratio > 0.5 indicating a positive sign test (convergence), while a ratio < 0.5 indicates divergence. We conducted two sets of sign-tests, one comparing the direction of effect for each pair-wise combination of cohorts, and one comparing each LOO meta-analysis with the respective sample that was left out. While fluctuations in the sign-tests across different p-value thresholds are expected, depending on the true association of each SNP with the phenotype, we mainly aimed to assess whether a specific cohort markedly deviated from the rest.
Following the same procedure as for the cross-trait PRS analyses described above (see previous method-section on cross-trait PRS analyses for details), we conducted LOO PRS analyses to evaluate the relationship between the PRS scores of each LOO GWAS and standardized OCS scores in the left-out cohort. Further, we conducted genetic correlation analyses between each LOO OCD meta-analysis (LOO_NTR, LOO_SfS, LOO_STR, and LOO_TwinsUK) and the same set of 97 phenotypes as described earlier to explore a possible heterogeneity in correlation patterns depending on the included OCS cohorts.
Results
Genome-wide association results
The final OCS GWAS meta-analysis was based on 33,943 individuals with complete phenotypic and genomic data available and included 6,232,765 associations of autosomal SNPs. No significant inflation was observed (λ = 1.027, λ1000 = 1.001, LDSC intercept = 1.0047, see Supplementary Fig. S14 for a QQ-plot). No SNP reached genome-wide significance (p < 5 × 10−8; see Fig. 1 for a Miami-plot including the Manhattan-plot of the GWAS in the upper panel). The SNP with the lowest p-value was rs113538937 (p = 6.36 × 10−8) on chromosome 4 (see Supplementary Fig. S15 for a regional association plot and forest plot). The region tagged by this SNP spans 207.6 kb (LD R² > 0.6) and entails the genes SH3BP2, ADD1, MFSD10, NOP14-AS1, NOP14, GRK4, HTTAS, and HTT. Another 26 independent SNPs with a p-value < 1 × 10−5 were identified (see Supplementary Table S2 for a list of association results).
The y-axes represent -log10 p-values for the association of SNPs/genes with OCS. The x-axis represents chromosomes 1 to 22. In the upper plot, the p-value threshold for genome-wide significance (p = 5 × 10−8) is represented by the horizontal red line, suggestive significance (p = 1 × 10−5) by the blue line. In the lower panel, Bonferroni-corrected gene-wide significance (p = 2.708 × 10−6) is represented by the horizontal red line, suggestive gene-wide significance (p = 1 × 10−3) is indicated by the blue horizontal line.
Gene and gene-set analyses
Gene-based tests were conducted to test whether any protein-coding gene carries a load of common variation associated with OCS. Using MAGMA v1.08 within FUMA v1.3.7, input SNPs were mapped to 18,464 protein coding genes. No gene reached the Bonferroni-corrected significance threshold (p = 0.05/18,464 = 2.71 × 10−6) in the gene-based test (see Fig. 1 for a Miami-plot including the Manhattan-plot of the gene-based test in the lower panel, and Supplementary Fig. S14 for a QQ-plot). Further, 15,496 gene sets (Curated gene sets: 5500, GO terms: 9996) from MsigDB v7.0 were tested for association. One curated gene set (Mootha Gluconeogenesis) reached Bonferroni-corrected significance (Ngenes = 28, Beta = 0.7872, SE = 0.1611, Pbon = 0.008).
Heritability and cross-disorder analyses
Heritability estimates
For the twin cohorts, the additive genetic variance of OCS, estimated based on the sparse genetic relatedness matrix from the association tests in GCTA, ranged between 0.33 (NTR) and 0.43 (TwinsUK), with estimates for the STR cohorts in between: STR-CATSS: 0.35 (GSA chip sub-sample), STR-CATSS: 0.41 (PsychChip sub-sample), STR-YATSS: 0.39, STR-STAGE: 0.39. Note that these heritability estimates based on the twin cohorts are largely driven by the twin resemblance (~0.5 between DZ twins and siblings, 1.0 for MZ twins, and 0 between unrelated individuals). The heritability for SfS, only including unrelated individuals, was 0.083 (SE = 0.053, P = 0.0516). The SNP-based heritability estimate of the GWAS meta-analysis using LDSC resulted in a total observed scale h² of 0.041 (SE = 0.0144, Z = 2.85, P = 0.0044).
Cross-disorder genetic correlations
We estimated genetic correlations using LDSC between the current OCS GWAS and OCD as well as 97 other phenotypes, including psychiatric, personality, psychological, substance-use, neurological, cognition, socioeconomic status, autoimmune, cardio-vascular, anthropomorphic, and fertility traits. Of these, 24 exceeded the FDR-corrected significance threshold (Fig. 2). As expected, OCS were most strongly associated with OCD out of all psychiatric disorders, followed by anxiety, DEP, major depressive disorder (MDD), SCZ, and AN. Significant positive genetic correlations were also observed for all 15 neuroticism phenotypes (14 sub-items and the neuroticism total score). Higher correlations emerged for all worry-related items, and slightly lower correlations for all depressive-related items. A significant positive correlation was further observed for “cigarettes per day” and “tiredness”, while “subjective well-being” yielded a significant negative correlation (see Fig. 2, and Supplementary Table S4). We further compared the genetic correlation patterns of OCS and OCD with all other 97 traits, to identify concordant and discordant patterns. There was a clear relationship between the two correlation-patterns of OCS and OCD (see Fig. 3), with 66 traits showing concordant directions of correlation (i.e., both correlations above 0 or both below 0), and 30 traits showing a discordant direction of correlation (i.e., one correlation above 0 and one below 0). Especially strong concordance was observed for the psychiatric and personality traits. Of the significant correlations with OCS, only ‘neuroticism loneliness’ (non-significant for OCD) and ‘cigarettes per day’ (significant for OCD) showed a different direction of effect with OCD (both positive with OCS, but negative with OCD).
Genetic correlations (rg) between OCS and a broad range (N = 97) of other phenotypes, assembled into 11 groups (psychiatric, substance, cognition/socioeconomic status (SES), personality, psychological, neurological, autoimmune, cardiovascular, anthropomorphic, fertility, and other). Error bars represent 95% confidence intervals; red circles indicate significant association after FDR correction for multiple testing.
Comparison between genetic correlation estimates (rG) of OCS and OCD for 97 other phenotypes, color-coded according to 11 groups (psychiatric, substance, cognition/socioeconomic status (SES), personality, psychological, neurological, autoimmune, cardiovascular, anthropomorphic, fertility, and other). On the x-axis the genetic correlation estimates for OCD are displayed, on the y-axis for OCS. Green quadrants indicate concordance, red quadrants discordance in the direction of genetic correlations between OCS and OCD.
Polygenic risk score analyses
We calculated a range of PRSs, based on publicly available summary statistics of OCD, DEP, SCZ, ASD, ADHD, and EA and examined their relationship with OCS in each cohort. GEE (STR, NTR, TwinsUK) and linear regression analyses (SfS) revealed significant (Bonferroni-corrected p < 0.05/7 = 0.0071) associations between OCS and PRS based on OCD, DEP, SCZ, and EA, but not consistently across all target datasets (see Supplementary Table S4). In the meta-analysis, summarizing the PRS results across all target cohorts, the OCD (Pfixed = 3.06 × 10−5) and SCZ (Prandom = 3.69 × 10−6) PRS showed significant associations with OCS. The DEP PRS also showed a significant association with OCS, however, there was significant heterogeneity across the four cohorts (PQ = 0.0020), and the random effects model failed to reach significance (Prandom = 0.0801). The other traits’ PRSs (ASD, ADHD, EA) were not significantly associated with OCS in the PRS meta-analysis (All meta-analyzed results are shown in Table 2).
Comparability between cohorts
No genome-wide significant heterogeneity was observed in the OCS GWAS meta-analysis (see Supplementary Fig. S16 for Manhattan plot and QQplot). The LOO PRS meta-analysis showed a significant positive association between the LOO PRS and OCS (Pfixed = 5.03 × 10−7) (see Table 2 for the PRS meta-analysis results and Supplementary Table S4 for individual results for each target cohort).
Given the low power for the main GWAS analysis, the power for LOO GWAS analyses and signtests between partial analyses is even lower, making it difficult to draw any definitive conclusions from the results. While no individual study markedly stands out from the rest, results of the sign tests analyses fluctuate with estimates ranging from 0.28 to 0.68 (and between 0 and 1 for P = 1 × 10−6). See Supplementary Figs. S17–S20 for Manhattan-plots and QQ-plots of the LOO GWASs and Supplementary Tables S5 and S6 for sign-test results.
We further calculated genetic correlations between each LOO OCS meta-analysis and the same 97 phenotypes (described above) to compare the individual influence of each cohort on the overall correlation estimates. When not considering the individual sub-items of neuroticism, the LOO GWAS excluding SfS showed 13 significant correlations, the GWAS without TwinsUK showed eight significant correlations, the GWAS without STR showed two significant correlations, while the GWAS excluding NTR did not significantly correlate with any of the traits (see Supplementary Fig. S21). As sample sizes for the LOO GWAS meta-analyses varied (LOO STR: N = 17,055; LOO NTR: N = 25,393; LOO SfS: N = 28,772; LOO TwinsUK: N = 30,609), it was expected that the power to detect significant correlations for each LOO GWAS differs. For almost all genetic correlations, each LOO GWAS meta-analysis showed the same direction of effect. For all correlations with psychiatric disorders and neuroticism phenotypes, the LOO analysis excluding NTR showed slightly higher estimates (but also larger confidence intervals).
Discussion
This is the first meta-analysis aiming to identify the genetic underpinnings of OCS in the general population. Although we could not replicate previous findings, two SNPs previously associated with OCS reached suggestive significance in this meta-analysis: the SNP found by den Braber et al. (rs8100480, p = 2.56 × 10−8; [22]) had a p-value of 0.0155 in the current study, while the SNP reported in Burton et al. (rs7856850, p = 2.48 × 10−8; [15]) had a p-value of 0.00029. The SNP with the lowest p-value (p = 6.36 × 10−8) in the current meta-analysis was rs113538937 on chromosome 4, tagging eight genes which have previously been associated with alcohol use (HTT), smoking (GRK4, NOP14, NOP14-AS1, ADD1, SH3BP2), worry (HTT), and measures of socioeconomic status and education (HTT, GRK4, NOP14, NOP14-AS, ADD1). In the gene-based tests, no gene achieved genome-wide significance. One gene-set reached Bonferroni-corrected significance. Expression of genes in this set is high at sites of insulin-mediated glucose disposal [63]. Dysregulated insulin signaling in the etiology of OCS has been suggested by a previous study describing a genetic overlap of OCS with an insulin signaling-related trait in children and adolescents [64]. Epidemiological studies similarly support the role of dysregulated insulin signaling in OCD and OCS. Specifically, patients diagnosed with OCD were found to have a significantly higher risk of developing type 2 diabetes compared to population controls [65].
Our results show that OCS share genetic risk with OCD. Polygenic risk for OCD was associated with OCS; and OCS and OCD case/control status showed a substantial genetic correlation (rG = 0.72, p = 0.0007). This is comparable to estimates reported in recent studies (rG = 0.61, p = 0.017 [12]; rG = 0.83, p = 0.07; [15]). Notably, OCS did show the highest correlation coefficient with OCD (rG = 0.72), but the strength of this genetic correlation was statistically not significantly different from that with anxiety (rG = 0.62), DEP (rG = 0.51), SCZ (rG = 0.34), or AN (rG = 0.30), as confidence intervals overlapped. While we did not exclude participants with any of these diagnoses, the most likely explanation is that these common comorbidities of OCD share genetic underpinnings with OCS. We also observed a high concordance in direction and strength of the correlation patterns of OCS and OCD with other phenotypes. The concordance between the OCD and OCS GWASs indicate that the genetic variation captured by our symptom based GWAS in the general population reflects that of the disorder OCD. However, given the present sample size, these values must also be interpreted with caution. A fact that is further illustrated by our PRS analyses: scores calculated based on the most recent SCZ GWAS more accurately predicted OCS than scores calculated based on the most recent OCD GWAS, which is considerably underpowered. This suggests that larger cohorts are needed for the accurate estimation of these associations.
SNP-based heritability for OCS in the current sample of 4.1% was significant, but lower than previous studies (7–16%; [12, 15, 22]), also standing in stark contrast with heritability estimates observed in twin studies (37–41%; [66]). This may be explained by the heterogeneity across the eight cohorts. Specifically, these samples differed in their ascertainment (twins vs. community-based samples), instruments employed, as well as age range of participants. While no specific pattern emerged in our compatibility analyses, the sign tests indicated some level of heterogeneity. Further evidence for heterogeneity was found in the leave-one-out analysis (Supplementary Fig. S3), supporting the view that there were some differences in cohorts with respect to the genetic correlation with specific variables. For leave-STR-out, the correlations with IQ and some related phenotypes were lower, whereas for leave-NTR-out, the correlations with neuroticism were increased. For most phenotypes, however, the leave-one-out genetic correlations did not yield strong or significant change as evidenced from the confidence intervals. In the PRS analyses, results deviated mostly for the SfS sample, which is the youngest and only non-twin sample included. It also uses the only questionnaire that is coded from strengths to weaknesses (from −3 meaning far less often than average to +3 meaning far more often than average), which could have led to further differences between the cohorts.
In line with previous research reports, we found a lower SNP-based heritability for OCS [12, 15] than for clinical OCD (0.28–0.37) [67, 68]. The reason for the disparity in SNP heritability between traits and diagnosis is unclear but could be explained by the fact that clinical diagnosed OCD represents the extreme of the OCS distribution in the general population. The conversion to liability scale SNP-h2 [50] may also have contributed to uncertainty in the estimates, as the uncertainty of a population prevalence close to zero may improperly inflate the correction factor from observed to liability scale. Previous studies of related psychiatric disorders, such as SCZ, or MDD, revealed that the genetic risk is higher for more severe and chronic cases [69, 70]. In addition to this consideration of impairment, differences including informant (parent/self vs. clinician), type of measurement (categorical vs. quantitative), and timing (cross-sectional vs. lifetime symptoms) could contribute to the divergence in SNP-based heritability estimates of OCS and OCD.
The present study had several limitations. First, despite being the largest meta-analysis of OCS including, to our knowledge, all samples currently available worldwide, the sample size is still relatively small for estimating heritability and detecting specific significant genetic markers. The meta-analysis also currently lacks the integration of non-European samples. The present results thus call for replication in, and extension to larger and more diverse cohorts. Second, previous research has suggested that OCS dimensions (e.g. contamination, checking, harm or symmetry) may be etiologically heterogeneous [12, 65, 66, 71]. As such, future studies might aim to identify the genetic underpinnings of specific OCS dimensions. This could also tackle some of the heterogeneity issues that may have caused imperfect overlap between cohorts, lower SNP-h2, and reduced genetic correlation with clinical OCD. Third, no information was available on the presence of common clinical comorbidities in all samples. This precluded detailed analysis of the identified genetic overlap. Future studies in larger cohorts should also investigate in more detail how OCS relate to other phenotypes, for instance addiction and personality disorders. Fourth, the observation that patients often establish non-random relationships with persons affected by the same or another mental disorder [72], might extend to people with OCS and contribute to the observed genetic correlations of OCS with anxiety, DEP, SCZ and AN. However, the LD-score method does not investigate the impact of assortative mating [73]. Therefore, assessing the degree to which this phenomenon may have influenced the genetic correlation estimates was beyond the scope of the present study. Future investigations of larger data sets for OCS and other psychiatric disorders are needed to refine the analysis of shared and specific genetic risk as well as communalities and specificities of the respective disorders. Finally, for many of the samples included in our study the distribution of quantitative OCS scores followed a right-tailed distribution, with most of the subjects reporting low OC symptom scores (see Supplementary Material for more detail). Although we captured substantial variability that would not have been present in more conventional case-control analyses, less variability amongst individuals with lower scores may have limited our power to identify positive genetic associations. The right-tailed distribution is due to the fact that most OCS scales are designed to screen for disorder in a clinical setting, rather than capture population variation in symptoms. The one exception in our study was the Spit for Science cohort, where the TOCS measure was explicitly designed for capturing variation in non-clinical samples [41, 42]. Despite these limitations, we argue for the inclusion of data based on conventional screening measures in order to achieve the necessary sample sizes for genomic studies, as they have been widely adopted and still capture greater variability in OCS compared with studies based on dichotomized case/control samples.
To summarize, OCS have a significant polygenic contribution and share genetic risk with diagnosed OCD, supporting the hypothesis that OCD represents the extreme end of widely distributed OCS in the population.
Data availability
The meta-analyzed summary statistics are available via the Psychiatric Genomics Consortium Download page (https://www.med.unc.edu/pgc/download-results/).
References
Mataix-Cols D, Boman M, Monzani B, Rück C, Serlachius E, Längström N, et al. Population-based, multigenerational family clustering study of obsessive-compulsive disorder. JAMA Psychiatry. 2013;70:709–17.
International Obsessive Compulsive Disorder Foundation Genetics Collaborative (IOCDF-GC) and OCD Collaborative Genetics Association Studies (OCGAS). Revealing the complex genetic architecture of obsessive–compulsive disorder using meta-analysis. Mol Psychiatry. 2017;23:1181–8.
Strom NI, Yu D, Gerring ZF, Halvorsen MW, Abdellaoui A, Rodriguez-Fontenla C, et al. Genome-wide association study identifies new locus associated with OCD. medRxiv. 2021 https://doi.org/10.1101/2021.10.13.21261078.
Brakoulias V, Starcevic V, Belloch A, Brown C, Ferrao YA, Fontenelle LF, et al. Comorbidity, age of onset and suicidality in obsessive-compulsive disorder (OCD): An international collaboration. Compr Psychiatry. 2017;76:79–86.
Mandelli L, Draghetti S, Albert U, De Ronchi D, Atti AR. Rates of comorbid obsessive compulsive disorder in eating disorders: A meta-analysis of the literature. J Affect Disord. 2020;277:927–39.
Pauls DL, Abramovitch A, Rauch SL, Geller DA. Obsessive-compulsive disorder: An integrative genetic and neurobiological perspective. Nat Rev Neurosci. 2014;15:410–24.
Lee PH, Anttila V, Won H, Feng YCA, Rosenthal J, Zhu Z, et al. Genomic Relationships, Novel Loci, and Pleiotropic Mechanisms across Eight Psychiatric Disorders. Cell. 2019;179:1469–82.e11.
Yilmaz Z, Halvorsen M, Bryois J, Yu D, Thornton LM, Zerwas S, et al. Examination of the shared genetic basis of anorexia nervosa and obsessive–compulsive disorder. Mol Psychiatry. 2020;25:2036–46.
Strom NI, Soda T, Mathews CA, Davis LK. A dimensional perspective on the genetics of obsessive-compulsive disorder. Transl Psychiatry. 2021;11:1–11.
Fullana MA, Vilagut G, Rojas-Farreras S, Mataix-Cols D, de Graaf R, Demyttenaere K, et al. Obsessive-compulsive symptom dimensions in the general population: Results from an epidemiological study in six European countries. J Affect Disord. 2010;124:291–9.
Barzilay R, Patrick A, Calkins ME, Moore TM, Wolf DH, Benton TD, et al. Obsessive-Compulsive Symptomatology in Community Youth: Typical Development or a Red Flag for Psychopathology? J Am Acad Child Adolesc Psychiatry. 2019;58:277–86.e4.
Smit DJA, Cath D, Zilhão NR, Ip HF, Denys D, den Braber A, et al. Genetic metaanalysis of obsessive–compulsive disorder and self-report compulsive symptoms. Am J Med Genet Part B Neuropsychiatr Genet. 2020;183:208–16.
Fawcett EJ, Power H, Fawcett JM. Women Are at Greater Risk of OCD ThanMen: A Meta-Analytic Review of OCD Prevalence Worldwide. J Clin Psychiatry. 2020:81;19r13085. https://doi.org/10.4088/JCP.19r13085.
Fullana MA, Mataix-Cols D, Caspi A, Harrington H, Grisham JR, Moffitt TE, et al. Obsessions and compulsions in the community: prevalence, interference, help-seeking, developmental stability, and co-occurring psychiatric conditions. Am J Psychiatry. 2009;166:329–36.
Burton CL, Lemire M, Xiao B, Corfield EC, Erdman L, Bralten J, et al. Genome-wide association study of pediatric obsessive-compulsive traits: shared genetic risk between traits and disorder. Transl Psychiatry. 2021;11:1–10.
Mataix-Cols D, Conceição do Rosario-Campos MD, Leckman JF. A Multidimensional Model of Obsessive-Compulsive Disorder. Am J Psychiatry. 2005;162:228–38.
Burton CL, Park LS, Corfield EC, Forget-Dubois N, Dupuis A, Sinopoli VM, et al. Heritability of obsessive–compulsive trait dimensions in youth from the general population. Transl Psychiatry. 2018;8:191.
Hudziak JJ, Van Beijsterveldt CEM, Althoff RR, Stanger C, Rettew DC, Nelson EC, et al. Genetic and environmental contributions to the Child Behavior Checklist Obsessive-Compulsive Scale: a cross-cultural twin study. Arch Gen psychiatry. 2004;61:608–16.
van Grootheest DS, Cath DC, Beekman AT, Boomsma DI. Twin Studies on Obsessive–Compulsive Disorder: A Review. Twin Res Hum Genet. 2005;8:450–8.
Zilhao NR, Smit DJ, Boomsma DI, Cath DC. Cross-disorder genetic analysis of tic disorders, obsessive-compulsive, and hoarding symptoms. Front Psychiatry. 2016:7. https://doi.org/10.3389/fpsyt.2016.00120.
Mathews CA, Delucchi K, Cath DC, Willemsen G, Boomsma DI. Partitioning the etiology of hoarding and obsessive-compulsive symptoms. Psychological Med. 2014;44:2867–76.
den Braber A, Zilhão NR, Fedko IO, Hottenga JJ, Pool R, Smit DJA, et al. Obsessive-compulsive symptoms in a large population-based twin-family sample are predicted by clinically based polygenic scores and by genome-wide SNPs. Transl psychiatry. 2016;6:e731.
Ruscio AM, Stein DJ, Chiu WT, Kessler RC. The Epidemiology of Obsessive-Compulsive Disorder in the National Comorbidity Survey Replication. Mol Psychiatry. 2010;15:53–63.
Lochner C, Fineberg NA, Zohar J, Van Ameringen M, Juven-Wetzler A, Altamura AC, et al. Comorbidity in obsessive-compulsive disorder (OCD): A report from the International College of Obsessive-Compulsive Spectrum Disorders (ICOCS). Compr Psychiatry. 2014;55:1513–9.
Tükel R, Polat A, Özdemir Ö, Aksüt D, Türksoy N. Comorbid conditions in obsessive-compulsive disorder. Compr Psychiatry. 2002;43:204–9.
Pallanti S, Grassi G, Sarrecchia ED, Cantisani A, Pellegrini M. Obsessive-compulsive disorder comorbidity: Clinical assessment and therapeutic implications. Front Psychiatry. 2011;2. https://doi.org/10.3389/fpsyt.2011.00070.
van Steensel FJA, Bögels SM, Perrin S. Anxiety Disorders in Children and Adolescents with Autistic Spectrum Disorders: A Meta-Analysis. Clin Child Fam Psychol Rev. 2011;14:302–17.
Zagai U, Lichtenstein P, Pedersen NL, Magnusson PKE. The Swedish Twin Registry: Content and Management as a Research Infrastructure. Twin Res Hum Genet. 2019;22:672–80.
Lichtenstein P, Sullivan PF, Cnattingius S, Gatz M, Johansson S, Carlstrom E, et al. The Swedish Twin Registry in the third millennium: an update. Twin Res Hum Genet. 2006;9:875–82.
Magnusson PKE, Almqvist C, Rahman I, Ganna A, Viktorin A, Walum H, et al. The Swedish twin registry: Establishment of a biobank and other recent developments. Twin Res Hum Genet. 2013;16:317–29.
Anckarsäter H, Lundström S, Kollberg L, Kerekes N, Palm C, Carlström E, et al. The Child and Adolescent Twin Study in Sweden (CATSS). Twin Res Hum Genet. 2011;14:495–508.
Boomsma, Vink DI, Beijsterveldt JM, van TCEM, Geus, de EJC, et al. Netherlands Twin Register: A Focus on Longitudinal Research. Twin Res Hum Genet. 2002;5:401–6.
Crosbie J, Arnold P, Paterson A, Swanson J, Dupuis A, Li X, et al. Response Inhibition and ADHD Traits: Correlates and Heritability in a Community Sample. J Abnorm Child Psychol. 2013;41:497.
Ameis SH, Lerch JP, Taylor MJ, Lee W, Viviano JD, Pipitone J, et al. A diffusion tensor imaging study in children with ADHD, autism spectrum disorder, OCD, and matched controls: Distinct and non-distinct white matter disruption and dimensional brain-behavior relationships. Am J Psychiatry. 2016;173:1213–22.
Spector TD, Williams FMK. The UK Adult Twin Registry (TwinsUK). Twin Res Hum Genet. 2006;9:899–906.
Bejerot S, Edman G, Anckarsäter H, Berglund G, Gillberg C, Hofvander B, et al. The Brief Obsessive–Compulsive Scale (BOCS): A self-report scale for OCD and obsessive– compulsive related disorders. Nord J Psychiatry. 2014;68:549.
Foa EB, Huppert JD, Leiberg S, Langner R, Kichic R, Hajcak G, et al. The obsessive-compulsive inventory: Development and validation of a short version. Psychological Assess. 2002;14:485–96.
Burns GL, Keortge SG, Formea GM, Sternberger LG. Revision of the Padua Inventory of obsessive compulsive disorder symptoms: distinctions between worry, obsessions, and compulsions. Behav Res Ther. 1996;34:163–73.
Van Grootheest DS, Cath D, Hottenga JJ, Beekman AT, Boomsma DI. Genetic factors underlie stability of obsessive-compulsive symptoms. Twin Res Hum Genet. 2009;12:411–9.
Cath DC, Van Grootheest DS, Willemsen G, Van Oppen P, Boomsma DI. Environmental factors in obsessive-compulsive behavior: Evidence from discordant and concordant monozygotic twins. Behav Genet. 2008;38:108–20.
Lambe LJ, Burton CL, Anagnostou E, Kelley E, Nicolson R, Georgiades S, et al. Clinical validation of the parent-report Toronto Obsessive–Compulsive Scale (TOCS): A pediatric open-source rating scale. JCPP Adv. 2021;1:e12056.
Park LS, Burton CL, Dupuis A, Shan J, Storch EA, Crosbie J, et al. The Toronto Obsessive-Compulsive Scale: Psychometrics of a Dimensional Measure of Obsessive-Compulsive Traits. J Am Acad Child Adolesc Psychiatry. 2016;55:310–18.e4.
Yang J, Lee SH, Goddard ME, Visscher PM. GCTA: A tool for genome-wide complex trait analysis. Am J Hum Genet. 2011;88:76–82.
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279–83.
Willer CJ, Li Y, Abecasis GR. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Lam M, Awasthi S, Watson HJ, Goldstein J, Panagiotaropoulou G, Trubetskoy V, et al. RICOPILI: Rapid Imputation for COnsortias PIpeLIne. Bioinformatics. 2020;36:930–3.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: Generalized Gene-Set Analysis of GWAS Data. PLoS Comput Biol. 2015;11:1–19.
Watanabe K, Taskesen E, Van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Comm. 2017;8:1–10.
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature. 2015;526:68–74.
Lee SH, Wray NR, Goddard ME, Visscher PM. Estimating missing heritability for disease from genome-wide association studies. Am J Hum Genet. 2011;88:294–305.
Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Altshuler DM, Gibbs RA, Peltonen L, Schaffner SF, Yu F, Dermitzakis E, et al. Integrating common and rare genetic variation in diverse human populations. Nature. 2010;467:52–58.
Howard DM, Adams MJ, Clarke TK, Hafferty JD, Gibson J, Shirali M, et al. Genomewide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat Neurosci. 2019;22:343–52.
Trubetskoy V, Pardiñas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Grove J, Ripke S, Als TD, Mattheisen M, Børglum AD, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431–44.
Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E, et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet. 2019;51:63–75.
Lee JJ, Wedow R, Okbay A, Kong E, Maghzian O, Zacher M, et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet. 2018;50:1112–21.
Choi SW, O’Reilly PF. PRSice-2: Polygenic Risk Score software for biobank-scale data. GigaScience. 2019;8:1–6.
Carey VJ. gee: Generalized Estimation Equation Solver. R package version 4.13-20. 2019 https://CRANR-project.org/package=gee.
Cochran WG. The comparison of percentages in matched samples. Biometrika. 1950;37:256–66.
Higgins JPT, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–60.
Higgins JPT, Thompson SG. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–58.
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–73.
Bralten J, Widomska J, Witte WD, Yu D, Mathews CA, Scharf JM, et al. Shared genetic etiology between obsessive-compulsive disorder, obsessive-compulsive symptoms in the population, and insulin signaling. Transl Psychiatry. 2020;10:121.
Isomura K, Brander G, Chang Z, Kuja-Halkola R, Rück C, Hellner C, et al. Metabolic and Cardiovascular Complications in Obsessive-Compulsive Disorder: A Total Population, Sibling Comparison Study With Long-Term Follow-up. Biol Psychiatry. 2018;84:324–31.
Taylor S, Jang KL, Asmundson GJG. Etiology of obsessions and compulsions: a behavioral-genetic analysis. J Abnorm Psychol. 2010;119:672–82.
Davis LK, Yu D, Keenan CL, Gamazon ER, Konkashbaev AI, Derks EM, et al. Partitioning the Heritability of Tourette Syndrome and Obsessive Compulsive Disorder Reveals Differences in Genetic Architecture. PLoS Genet. 2013;9. https://doi.org/10.1371/journal.pgen.1003864.
Mahjani B, Klei L, Mattheisen M, Halvorsen MW, Reichenberg A, Roeder K, et al. The Genetic Architecture of Obsessive-Compulsive Disorder: Contribution of Liability to OCD From Alleles Across the Frequency Spectrum. Am J Psychiatry. 2021 https://doi.org/10.1176/APPI.AJP.2021.21010101.
Meier SM, Agerbo E, Maier R, Pedersen CB, Lang M, Grove J, et al. High loading of polygenic risk in cases with chronic schizophrenia. Mol Psychiatry. 2016;21:969–74.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne E, Abdellaoui A. Genomewide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Iervolino AC, Rijsdijk FV, Cherkas L, Fullana MA, Mataix-Cols D. A multivariate twin study of obsessive-compulsive symptom dimensions. Arch Gen Psychiatry. 2011;68:637–44.
Nordsletten AE, Larsson H, Crowley JJ, Almqvist C, Lichtenstein P, Mataix-Cols D. Patterns of Nonrandom Mating Within and Across 11 Major Psychiatric Disorders. JAMA Psychiatry. 2016;73:354–61.
Bulik-Sullivan B, Finucane HK, Anttila V, Gusev A, Day FR, Loh PR, et al. An atlas of genetic correlations across human diseases and traits. Nat Genet. 2015;47:1236–41.
Acknowledgements
We acknowledge The Swedish Twin Registry (STR) for data access. The STR is managed by Karolinska Institutet and receives funding through the Swedish Research Council under the grant no 2017-00641. The computations/data handling were enabled by resources provided by the Swedish National Infrastructure for Computing (SNIC) at Uppmax partially funded by the Swedish Research Council through grant agreement no. 2018-05973. The Netherlands Twin Register (NTR) warmly thanks all twin families for their participation. NTR is supported by multiple grants from the Netherlands Organizations for Scientific Research (NWO) and Medical Research (ZonMW): Netherlands Twin Registry Repository (NWO 480-15-001/674); the Biobank-based integrative omics study (BIOS) funded by BBMRI-NL (NWO projects 184.021.007 and 184.033.111); the European Science Council (ERC) Genetics of Mental Illness (ERC Advanced, 230374, PI Boomsma); the Royal Netherlands Academy of Science Professor Award (PAH/6635) to DIB; Rutgers University Cell and DNA Repository (NIMH U24 MH06845706), the Avera Institute, Sioux Falls, South Dakota (USA) and the National Institutes of Health (NIH R01 HD042157-01A1). Part of the genotyping was funded by the Genetic Association Information Network (GAIN) of the Foundation for the National Institutes of Health and Grand Opportunity grants 1RC2 MH089951). TwinsUK is funded by the Wellcome Trust, Medical Research Council, Versus Arthritis, European Union Horizon 2020, Chronic Disease Research Foundation (CDRF), Zoe Ltd, the National Institute for Health and Care Research (NIHR) Clinical Research Network (CRN) and Biomedical Research Centre based at Guy’s and St Thomas’ NHS Foundation Trust in partnership with King’s College London. Spit for Science was supported by the Canadian Institutes of Health Research (RJS, MOP‐93696 and PDA, MOP‐106573). PDA is supported by the Alberta Innovates Translational Health Chair in Child and Youth Mental Health.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
PDA, DJAS, SMM, DMC, DIB, MM, and NIS contributed to the conception of the overall study design; CLB, LA, RP, ML, JJC, JJH, VZI, HL, PL, PM, CR, RJS, HMW, JC, PDA, DIB, SMM, DMC, and DC contributed to the data collection of the individual datasets and/or provided code; NIS, DJAS, TS, and CI conducted all primary data analyses; NIS, DJAS, SMM, and CLB drafted the manuscript; all authors provided critical edits and discussions and approved the submitted version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
DM-C receives royalties for contributing articles to UpToDate, Wolters Kluwer Health, outside of the submitted work. RJS has consulted to E. Lilly, Highland Therapeutics and eHave. HL reports receiving grants from Shire Pharmaceuticals; personal fees from and serving as a speaker for Medice, Shire/Takeda Pharmaceuticals and Evolan Pharma AB; and sponsorship for a conference on attention-deficit/hyperactivity disorder from Shire/Takeda Pharmaceuticals and Evolan Pharma AB, all outside the submitted work. HL is editorin-chief of JCPP Advances. PA receives research funding from Biohaven Pharmaceuticals, unrelated to the submitted work. All other authors report no potential conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Strom, N.I., Burton, C.L., Iyegbe, C. et al. Genome-Wide Association Study of Obsessive-Compulsive Symptoms including 33,943 individuals from the general population. Mol Psychiatry 29, 2714–2723 (2024). https://doi.org/10.1038/s41380-024-02489-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-024-02489-6