Abstract
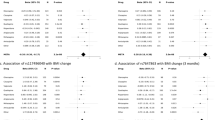
Antipsychotic-induced weight gain (AIWG) is a common side effect of antipsychotic medication and may contribute to diabetes and coronary heart disease. To expand the unclear genetic mechanism underlying AIWG, we conducted a two-stage genome-wide association study in Han Chinese patients with schizophrenia. The study included a discovery cohort of 1936 patients and a validation cohort of 534 patients, with an additional 630 multi-ancestry patients from the CATIE study for external validation. We applied Mendelian randomization (MR) analysis to investigate the relationship between AIWG and antipsychotic-induced lipid changes. Our results identified two novel genome-wide significant loci associated with AIWG: rs10422861 in PEPD (P = 1.373 × 10–9) and rs3824417 in PTPRD (P = 3.348 × 10–9) in Chinese Han samples. The association of rs10422861 was validated in the European samples. Fine-mapping and functional annotation revealed that PEPD and PTPRD are potentially causal genes for AIWG, with their proteins being prospective therapeutic targets. Colocalization analysis suggested that AIWG and type 2 diabetes (T2D) shared a causal variant in PEPD. Polygenic risk scores (PRSs) for AIWG and T2D significantly predicted AIWG in multi-ancestry samples. Furthermore, MR revealed a risky causal effect of genetically predicted changes in low-density lipoprotein cholesterol (P = 7.58 × 10−4) and triglycerides (P = 2.06 × 10−3) caused by acute-phase of antipsychotic treatment on AIWG, which had not been previously reported. Our model, incorporating antipsychotic-induced lipid changes, PRSs, and clinical predictors, significantly predicted BMI percentage change after 6-month antipsychotic treatment (AUC = 0.79, R2 = 0.332). Our results highlight that the mechanism of AIWG involves lipid pathway dysfunction and may share a genetic basis with T2D through PEPD. Overall, this study provides new insights into the pathogenesis of AIWG and contributes to personalized treatment of schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
NIMH REPOSITORY & GENOMICS RESOURCE approved the individual data from the CATIE study to the present study (https://www.nimhgenetics.org/, Requested ID: 63084551a4921). GWAS summary data on the association between SNPs and East Asian T2D was retrieved from the website https://blog.nus.edu.sg/agen/summary-statistics/t2d-2020. Summary statistics for the 912,253 SNP-protein expression pairs in the ROSMAP pQTL dataset are based on data obtained from the AD Knowledge Portal (https://adknowledgeportal.org), which can be found at https://www.synapse.org/#!Synapse:syn23191787. The other datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Lin J-J, Liang F-W, Li C-Y, Lu T-H. Leading causes of death among decedents with mention of schizophrenia on the death certificates in the United States. Schizophr Res. 2018;197:116–23.
Hjorthøj C, Stürup AE, McGrath JJ, Nordentoft M. Years of potential life lost and life expectancy in schizophrenia: a systematic review and meta-analysis. Lancet Psychiatry. 2017;4:295–301.
Nielsen RE, Banner J, Jensen SE. Cardiovascular disease in patients with severe mental illness. Nat Rev Cardiol. 2021;18:136–45.
Pillinger T, McCutcheon RA, Vano L, Mizuno Y, Arumuham A, Hindley G, et al. Comparative effects of 18 antipsychotics on metabolic function in patients with schizophrenia, predictors of metabolic dysregulation, and association with psychopathology: a systematic review and network meta-analysis. Lancet Psychiatry. 2020;7:64–77.
Willett WC, Manson JE, Stampfer MJ, Colditz GA, Rosner B, Speizer FE, et al. Weight, weight change, and coronary heart disease in women. Risk within the ‘normal’ weight range. JAMA. 1995;273:461–5.
Grajales D, Ferreira V, Valverde ÁM. Second-generation antipsychotics and dysregulation of glucose metabolism: beyond weight gain. Cells. 2019;8:1336.
Gebhardt S, Theisen FM, Haberhausen M, Heinzel-Gutenbrunner M, Wehmeier PM, Krieg J-C, et al. Body weight gain induced by atypical antipsychotics: an extension of the monozygotic twin and sib pair study. J Clin Pharm Ther. 2010;35:207–11.
Lett TaP, Wallace TJM, Chowdhury NI, Tiwari AK, Kennedy JL, Müller DJ. Pharmacogenetics of antipsychotic-induced weight gain: review and clinical implications. Mol Psychiatry. 2012;17:242–66.
Malhotra AK, Correll CU, Chowdhury NI, Müller DJ, Gregersen PK, Lee AT, et al. Association between common variants near the melanocortin 4 receptor gene and severe antipsychotic drug-induced weight gain. Arch Gen Psychiatry. 2012;69:904–12.
Yu H, Wang L, Lv L, Ma C, Du B, Lu T, et al. Genome-wide association study suggested the PTPRD polymorphisms were associated with weight gain effects of atypical antipsychotic medications. Schizophr Bull. 2016;42:814–23.
Brandl EJ, Tiwari AK, Zai CC, Nurmi EL, Chowdhury NI, Arenovich T, et al. Genome-wide association study on antipsychotic-induced weight gain in the CATIE sample. Pharmacogenom J. 2016;16:352–6.
Maciukiewicz M, Tiwari AK, Zai CC, Gorbovskaya I, Laughlin CP, Nurmi EL, et al. Genome-wide association study on antipsychotic-induced weight gain in Europeans and African-Americans. Schizophr Res. 2019;212:204–12.
ter Hark SE, Jamain S, Schijven D, Lin BD, Bakker MK, Boland-Auge A, et al. A new genetic locus for antipsychotic-induced weight gain: a genome-wide study of first-episode psychosis patients using amisulpride (from the OPTiMiSE cohort). J Psychopharmacol. 2020;34:524–31.
Sjaarda J, Delacrétaz A, Dubath C, Laaboub N, Piras M, Grosu C, et al. Identification of four novel loci associated with psychotropic drug-induced weight gain in a Swiss psychiatric longitudinal study: a GWAS analysis. Mol Psychiatry. 2023;28:2320–7.
Mallard TT, Grotzinger AD, Smoller JW. Examining the shared etiology of psychopathology with genome-wide association studies. Physiol Rev. 2023;103:1645–65.
Lau A, So H-C. Turning genome-wide association study findings into opportunities for drug repositioning. Comput Struct Biotechnol J. 2020;18:1639–50.
Segura ÀG, Martínez-Pinteño A, Gassó P, Rodríguez N, Bioque M, Cuesta MJ, et al. Metabolic polygenic risk scores effect on antipsychotic-induced metabolic dysregulation: a longitudinal study in a first episode psychosis cohort. Schizophr Res. 2022;244:101–10.
Johnson D, Wilke MAP, Lyle SM, Kowalec K, Jorgensen A, Wright GEB, et al. A systematic review and analysis of the use of polygenic scores in pharmacogenomics. Clin Pharm Ther. 2022;111:919–30.
Yu H, Yan H, Wang L, Li J, Tan L, Deng W, et al. Five novel loci associated with antipsychotic treatment response in patients with schizophrenia: a genome-wide association study. Lancet Psychiatry. 2018;5:327–38.
Yue W-H, Wang H-F, Sun L-D, Tang F-L, Liu Z-H, Zhang H-X, et al. Genome-wide association study identifies a susceptibility locus for schizophrenia in Han Chinese at 11p11.2. Nat Genet. 2011;43:1228–31.
Stroup TS, McEvoy JP, Swartz MS, Byerly MJ, Glick ID, Canive JM, et al. The National Institute of Mental Health Clinical Antipsychotic Trials of Intervention Effectiveness (CATIE) project: schizophrenia trial design and protocol development. Schizophr Bull. 2003;29:15–31.
Lieberman JA, Stroup TS, McEvoy JP, Swartz MS, Rosenheck RA, Perkins DO, et al. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N Engl J Med. 2005;353:1209–23.
Chen J, Lippold D, Frank J, Rayner W, Meyer-Lindenberg A, Schwarz E. Gimpute: an efficient genetic data imputation pipeline. Bioinformatics. 2019;35:1433–5.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Bulik-Sullivan BK, Loh P-R, Finucane HK, Ripke S, Yang J, Patterson N, et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
Lu Z, Zhang Y, Sun Y, Liao Y, Kang Z, Feng X, et al. The positive association between antipsychotic-induced weight gain and therapeutic response: new biotypes of schizophrenia. Psychiatry Res. 2023;324:115226.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Schilder BM, Humphrey J, Raj T. echolocatoR: an automated end-to-end statistical and functional genomic fine-mapping pipeline. Bioinformatics. 2021;38:536–9.
Wakefield J. A Bayesian measure of the probability of false discovery in genetic epidemiology studies. Am J Hum Genet. 2007;81:208–27.
Maller JB, McVean G, Byrnes J, Vukcevic D, Palin K, Su Z.Wellcome Trust Case Control Consortium et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat Genet. 2012;44:1294–301.
Wang G, Sarkar A, Carbonetto P, Stephens M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J R Stat Soc Ser B Stat Methodol. 2020;82:1273–1300.
Kichaev G, Roytman M, Johnson R, Eskin E, Lindström S, Kraft P, et al. Improved methods for multi-trait fine mapping of pleiotropic risk loci. Bioinformatics. 2017;33:248–55.
Weissbrod O, Hormozdiari F, Benner C, Cui R, Ulirsch J, Gazal S, et al. Functionally informed fine-mapping and polygenic localization of complex trait heritability. Nat Genet. 2020;52:1355–63.
Guo L-K, Su Y, Zhang Y-Y-N, Yu H, Lu Z, Li W-Q, et al. Prediction of treatment response to antipsychotic drugs for precision medicine approach to schizophrenia: randomized trials and multiomics analysis. Mil Med Res. 2023;10:24.
Kamat MA, Blackshaw JA, Young R, Surendran P, Burgess S, Danesh J, et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics. 2019;35:4851–3.
Wallace C. Eliciting priors and relaxing the single causal variant assumption in colocalisation analyses. PLoS Genet. 2020;16:e1008720.
Cheng W, Luo N, Zhang Y, Zhang X, Tan H, Zhang D, et al. DNA methylation and resting brain function mediate the association between childhood urbanicity and better speed of processing. Cereb Cortex. 2021;31:4709–18.
Euesden J, Lewis CM, O’Reilly PF. PRSice: polygenic risk score software. Bioinformatics. 2015;31:1466–8.
Spracklen CN, Horikoshi M, Kim YJ, Lin K, Bragg F, Moon S, et al. Identification of type 2 diabetes loci in 433,540 East Asian individuals. Nature. 2020;582:240–5.
Burgess S, Timpson NJ, Ebrahim S, Davey Smith G. Mendelian randomization: where are we now and where are we going? Int J Epidemiol. 2015;44:379–88.
Permutt T, Hebel JR. Simultaneous-equation estimation in a clinical trial of the effect of smoking on birth weight. Biometrics. 1989;45:619–22.
Liu X, Tong X, Zou Y, Lin X, Zhao H, Tian L, et al. Mendelian randomization analyses support causal relationships between blood metabolites and the gut microbiome. Nat Genet. 2022;54:52–61.
Wu D-M. Alternative tests of independence between stochastic regressors and disturbances. Econometrica. 1973;41:733–50.
Fitzgerald I, Sahm LJ, Byrne A, O’Connell J, Ensor J, Dhubhlaing CN, et al. Predicting antipsychotic-induced weight gain in first episode psychosis—a field-wide systematic review and meta-analysis of non-genetic prognostic factors. Eur Psychiatry. 2023;66:e42.
Burschinski A, Schneider‐Thoma J, Chiocchia V, Schestag K, Wang D, Siafis S, et al. Metabolic side effects in persons with schizophrenia during mid‐ to long‐term treatment with antipsychotics: a network meta‐analysis of randomized controlled trials. World Psychiatry. 2023;22:116–28.
Steyerberg EW, Vergouwe Y. Towards better clinical prediction models: seven steps for development and an ABCD for validation. Eur Heart J. 2014;35:1925–31.
Alba AC, Agoritsas T, Walsh M, Hanna S, Iorio A, Devereaux PJ, et al. Discrimination and calibration of clinical prediction models: users’ guides to the medical literature. JAMA. 2017;318:1377–84.
Wingo AP, Liu Y, Gerasimov ES, Gockley J, Logsdon BA, Duong DM, et al. Integrating human brain proteomes with genome-wide association data implicates new proteins in Alzheimer’s disease pathogenesis. Nat Genet. 2021;53:143–6.
Verbanck M, Chen C-Y, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018;50:693–8.
Chen Y-W, Diamante G, Ding J, Nghiem TX, Yang J, Ha S-M, et al. PharmOmics: a species- and tissue-specific drug signature database and gene-network-based drug repositioning tool. iScience. 2022;25:104052.
Yuan J, Li T, Yin X-B, Guo L, Jiang X, Jin W, et al. Characterization of prolidase activity using capillary electrophoresis with tris(2,2’-bipyridyl)ruthenium(II) electrochemiluminescence detection and application to evaluate collagen degradation in diabetes mellitus. Anal Chem. 2006;78:2934–8.
García-Flores LA, Green CL. Of Mice and Men: Impacts of calorie restriction on metabolomics of the cerebellum. J Gerontol A Biol Sci Med Sci. 2021;76:547–51.
Zhao B, Li T, Fan Z, Yang Y, Shu J, Yang X, et al. Heart-brain connections: phenotypic and genetic insights from magnetic resonance images. Science. 2023;380:abn6598.
Chen Z, Yu H, Shi X, Warren CR, Lotta LA, Friesen M, et al. Functional screening of candidate causal genes for insulin resistance in human preadipocytes and adipocytes. Circ Res. 2020;126:330–46.
Hua Y, Nair S. Proteases in cardiometabolic diseases: pathophysiology, molecular mechanisms and clinical applications. Biochim Biophys Acta. 2015;1852:195–208.
Mishra I, Xie WR, Bournat JC, He Y, Wang C, Silva ES, et al. Protein tyrosine phosphatase receptor δ serves as the orexigenic asprosin receptor. Cell Metab. 2022;34:549–63.e8.
Feng B, Liu H, Mishra I, Duerrschmid C, Gao P, Xu P, et al. Asprosin promotes feeding through SK channel-dependent activation of AgRP neurons. Sci Adv. 2023;9:eabq6718.
Chen M, Dong Y, Tian L, Zhou J, Zhu E, Yuan H, et al. Metastasis suppressor 1 interacts with protein tyrosine phosphatase receptor-δ to regulate adipogenesis. FASEB J. 2023;37:e22857.
Truong TTT, Panizzutti B, Kim JH, Walder K. Repurposing drugs via network analysis: opportunities for psychiatric disorders. Pharmaceutics. 2022;14:1464.
Fitzgerald I, O’Connell J, Keating D, Hynes C, McWilliams S, Crowley EK. Metformin in the management of antipsychotic-induced weight gain in adults with psychosis: development of the first evidence-based guideline using GRADE methodology. Evid Based Ment Health. 2022;25:15–22.
Maida A, Lamont BJ, Cao X, Drucker DJ. Metformin regulates the incretin receptor axis via a pathway dependent on peroxisome proliferator-activated receptor-α in mice. Diabetologia. 2011;54:339–49.
Liu X, Lian J, Hu C-H, Deng C. Betahistine co-treatment ameliorates dyslipidemia induced by chronic olanzapine treatment in rats through modulation of hepatic AMPKα-SREBP-1 and PPARα-dependent pathways. Pharm Res. 2015;100:36–46.
Caricilli AM, Penteado E, de Abreu LL, Quaresma PGF, Santos AC, Guadagnini D, et al. Topiramate treatment improves hypothalamic insulin and leptin signaling and action and reduces obesity in mice. Endocrinology. 2012;153:4401–11.
Pellegrinelli V, Rodriguez-Cuenca S, Rouault C, Figueroa-Juarez E, Schilbert H, Virtue S, et al. Dysregulation of macrophage PEPD in obesity determines adipose tissue fibro-inflammation and insulin resistance. Nat Metab. 2022;4:476–94.
Kennedy SA, Jarboui M-A, Srihari S, Raso C, Bryan K, Dernayka L, et al. Extensive rewiring of the EGFR network in colorectal cancer cells expressing transforming levels of KRASG13D. Nat Commun. 2020;11:499.
Harrington M, Pond-Tor S, Boney CM. Role of epidermal growth factor and ErbB2 receptors in 3T3-L1 adipogenesis. Obesity. 2007;15:563–71.
Uhl GR, Martinez MJ. PTPRD: neurobiology, genetics, and initial pharmacology of a pleiotropic contributor to brain phenotypes. Ann NY Acad Sci. 2019;1451:112–29.
Fan L, You Y, Fan Y, Shen C, Xue Y. Association between ApoA1 gene polymorphisms and antipsychotic drug-induced dyslipidemia in schizophrenia. Neuropsychiatr Dis Treat. 2021;17:1289–97.
Hong C-J, Chen T-T, Bai YM, Liou Y-J, Tsai S-J. Impact of apolipoprotein A5 (APOA5) polymorphisms on serum triglyceride levels in schizophrenic patients under long-term atypical antipsychotic treatment. World J Biol Psychiatry. 2012;13:22–9.
Huang QQ, Sallah N, Dunca D, Trivedi B, Hunt KA, Hodgson S, et al. Transferability of genetic loci and polygenic scores for cardiometabolic traits in British Pakistani and Bangladeshi individuals. Nat Commun. 2022;13:4664.
Hu Y, Graff M, Haessler J, Buyske S, Bien SA, Tao R, et al. Minority-centric meta-analyses of blood lipid levels identify novel loci in the Population Architecture using Genomics and Epidemiology (PAGE) study. PLoS Genet. 2020;16:e1008684.
Sinnott-Armstrong N, Tanigawa Y, Amar D, Mars N, Benner C, Aguirre M, et al. Genetics of 35 blood and urine biomarkers in the UK Biobank. Nat Genet. 2021;53:185–94.
Wojcik GL, Graff M, Nishimura KK, Tao R, Haessler J, Gignoux CR, et al. Genetic analyses of diverse populations improves discovery for complex traits. Nature. 2019;570:514–8.
Acknowledgements
This study was supported by the National Natural Science Foundation of China (81825009, 82330042, 82301687, 81901358); National Key R&D Program of China (2023YFE0119400, 2021YFF1201103); Fundamental Research Funds for the Central Universities (Peking University Medicine Fund for world’s leading discipline or discipline cluster development, BMU2022DJXK007); Chinese Academy of Medical Sciences Research Unit (2019-I2M-5-006, 2021-I2M-C&T-B-099), Collaborative Research Fund of Chinese Institute for Brain Research Beijing (grant 2020-NKX-XM-12); Major science and technology projects of Henan Province (201300310200); Natural Science Foundation of Shandong Province (ZR2019BH001 and ZR2021YQ55), and Young Taishan Scholars of Shandong Province (tsqn201909146). We thank all subjects who participated in our study. We also thank the assistance from the Chinese Antipsychotics Pharmacogenomics Consortium and the research assistant from Peking University Sixth Hospital (Xuan Lu).
Author information
Authors and Affiliations
Contributions
WY, YL, and HYu conceived and designed the project. YL and HYu contributed to the analysis and interpretation of data and wrote the manuscript. YL, HYu, and YZ did the statistical analyses and prepared the tables and figures. ZL, YaoS, LG, JG, ZK, and XF verified the results. WY, YZ, and DZ edited the manuscript and provided supervision. TL, YY, WL, LL, and HYa contributed to the data acquisition. YuS, GW, and ZS made the further interpretation of data. All authors contributed to drafting the work or critically revising it for important intellectual content and made substantial contributions to the concept and design of the study and data acquisition, analysis, and interpretation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, Y., Yu, H., Zhang, Y. et al. Genome-wide association study implicates lipid pathway dysfunction in antipsychotic-induced weight gain: multi-ancestry validation. Mol Psychiatry (2024). https://doi.org/10.1038/s41380-024-02447-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41380-024-02447-2