Abstract
A body of pre-clinical evidence shows how the gut microbiota influence brain functioning, including brain connectivity. Linking measures of brain connectivity to the gut microbiota can provide important mechanistic insights into the bi-directional gut-brain communication. In this systematic review, we therefore synthesized the available literature assessing this association, evaluating the degree of consistency in microbiota-connectivity associations. Following the PRISMA guidelines, a PubMed search was conducted, including studies published up to September 1, 2022. We identified 16 studies that met the inclusion criteria. Several bacterial genera, including Prevotella, Bacteroides, Ruminococcus, Blautia, and Collinsella were most frequently reported in association with brain connectivity. Additionally, connectivity of the salience (specifically the insula and anterior cingulate cortex), default mode, and frontoparietal networks were most frequently associated with the gut microbiota, both in terms of microbial diversity and composition. There was no discernible pattern in the association between microbiota and brain connectivity. Altogether, based on our synthesis, there is evidence for an association between the gut microbiota and brain connectivity. However, many findings were poorly replicated across studies, and the specificity of the association is yet unclear. The current studies show substantial inter-study heterogeneity in methodology and reporting, limiting the robustness and reproducibility of the findings and emphasizing the need to harmonize methodological approaches. To enhance comparability and replicability, future research should focus on further standardizing processing pipelines and employing data-driven multivariate analysis strategies.
Similar content being viewed by others
Introduction
Neuronal connections are determined by the distance between neurons. Spatially closer neurons have a higher probability of being connected than those further away. As a result, the structural wiring of the brain represents a complex network with clusters of highly connected regions (i.e., structural connectivity) [1] that provide a basis for functional communication between brain areas (i.e., functional connectivity) as measured by the temporal coincidence of neuronal activation patterns of anatomically separated brain regions [2]. Patterns of communication between fixed sets of brain regions form functional networks [3, 4], which can be identified both during active cognitive tasks and during rest.
Functional networks measured during rest (resting-state networks) reflect, among others, processes related to cognitive functions. For example, the salience network (SN) is involved in the detection of behaviorally relevant stimuli [5], the frontoparietal network (FPN) is involved in the coordination of cognitive control [6], and the default mode network (DMN) is linked to basal, stimulus-independent cognitive processes such as information integration and mind-wandering [7]. In addition to cognitive functioning, connectivity may also reflect intrinsic processes, such as emotional and interoceptive awareness [8, 9]. Dysfunction in connectivity networks is observed in a range of psychiatric and neurodevelopmental disorders, including Attention Deficit/Hyperactivity Disorder [10] and Schizophrenia [11], in certain cases even prior to diagnosis [12].
The functional and structural connectivity patterns of the brain are affected by numerous genetic and non-genetic interacting factors. Our genetic makeup has a significant biological effect on both brain structure and function: heritability studies have shown that the additive genetic contribution explains approximately 50% to 93% of the variance in structural connectivity [13] and 20% to 40% of the variance in functional connectivity [14]. Likewise, the brain can change under the influence of environmental factors. Multiple studies have reported changes in connectivity strength following a mindfulness training, both in structural [15] and functional connectivity [16]. Additionally, a systematic review concluded that a lower quality diet was related to decreased structural and functional connectivity of default mode, sensorimotor and attention networks [17].
Such environmental factors, including diet, may exert their influence on the brain through the gut-brain axis (GBA), among others via modulation of the gut microbiota [18]. The GBA refers to the bidirectional communication system connecting the gastrointestinal system with the central nervous system (CNS) through endocrine, immune, and neural/vagal pathways [19]. The gut microbiota, comprising the trillions of microbes (predominantly bacteria) residing in the intestines, can modulate gut-brain communication, for example through the production of neuroactive metabolites, and by affecting the integrity of the gastro-intestinal and blood-brain barriers [19, 20].
A majority of the studies investigating the (microbiota-)gut-brain axis (MGBA) in humans focus on behavioral measures, including clinical diagnoses and questionnaires, providing evidence for a link between the gut microbiota composition and cognitive and emotional functioning [21,22,23]. In recent years the number of studies incorporating neuroimaging into the microbiota-gut-brain investigation has also increased rapidly. Specifically, the acquisition of functional and structural connectivity data is relatively standardized and often done at rest (i.e., without task instructions), making it a suitable research method for many participant populations, including children and patients. Linking such connectivity measures to the gut microbiota can provide important mechanistic insights into the bi-directional gut-brain communication. Therefore, we herein systematically review the available studies associating the gut microbiota with brain connectivity, in an effort to evaluate the degree of consistency in this association.
Method
Search strategy
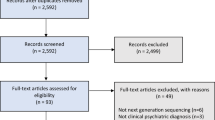
Following PRISMA guidelines, a systematic search on the PubMed database was conducted for reports published up to September 1, 2022. The aim was to capture all human studies that 1) collected a fecal sample to assess the gut microbiota, 2) assessed in vivo functional or structural brain connectivity, and 3) performed statistical analysis on the association between the gut microbiota and brain connectivity. Only peer-reviewed original research studies (i.e., reporting original data, analyses, and findings) published in English were included. Two independent raters (DM, MB) reviewed the titles and abstracts and came to a consensus about study inclusion. After inclusion, the following data was independently extracted by two authors (DM, MB): demographics, sample characteristics, method of gut microbiota estimation, method of brain connectivity estimation, statistical methods, and relevant results. Details on the search strategy and study inclusion are provided in Fig. 1. The PRISMA checklist is available in the Supplementary Materials.
Consistency of the findings
The consistency of the findings was assessed on two levels. First, we looked at the gut microbiota and brain connectivity individually. Second, we assessed the specificity of the microbiota-connectivity association. For the first, the assessment was done by dividing the number of studies reporting a statistically significant association for a diversity index/genus abundance/brain network by the total number of studies assessing it. For the second, the assessment was done by counting the number of studies reporting a statistically significant microbiota-connectivity association and dividing it by the total number of studies that could have identified this association. Only findings assessed in at least three studies were interpreted. A finding was considered consistent if it was reported in at least 50% of the studies (if assessed in four or more studies), with an absolute minimum of two studies (if assessed in three studies).
Quality assessment
Two authors (DM, MB) assessed the risk of bias in the included studies using the National Institutes of Health (NIH) National Health, Lung and Blood Institute Study Quality Assessment Tool for observational Cohort and Cross-sectional Studies [24]. The tool was modified to also be suitable for case-control and before-after studies with no control group (specified in Supplementary Table 3). We used the method section of the STORMS checklist (v1.03) [25] to assess gut microbiota measurements, and an adaptation of the guidelines published by Poldrack et al. [26] to assess brain connectivity measurements (Supplementary Table 4-5). Study quality was rated as ‘Good’ for assessments of 75% or higher; ‘Fair’ for assessments between 50% and 75%; or ‘Poor’ for assessments lower than 50%.
Results
A comprehensive literature search on PubMed yielded 360 reports, of which a total of 19 publications based on 16 unique studies met the inclusion criteria (Fig. 1). Two of the included studies produced more than one publication ([27,28,29] and [30, 31]). For those studies, the findings reported in the individual publications were pooled.
Population characteristics (Table 1)
There was a wide variation in the target populations. Eight studies were conducted in healthy individuals [27,28,29, 32,33,34,35,36,37] including a study conducted in smokers [32], in newborns [36], and in infants [35]. The other eight studies were performed in a range of disease populations, including bipolar disorder [38], cirrhosis [39], end stage renal disease [30, 31], irritable bowel syndrome [40, 41], major depressive disorder [42], obesity [43] and patients undergoing a laparoscopic/vertical sleeve gastrectomy [44, 45]. Four of those studies were case-controlled [30, 31, 38, 41, 43] and three were longitudinal [40, 44, 45]. All other studies performed associations between gut microbiota composition and brain connectivity based on a single group and timepoint. Further characteristics of the study populations are listed in Table 1.
Methodological characteristics (Table 2)
An overview of the methods and indices used to quantify and analyze the gut microbiota and functional and structural brain connectivity is provided in Fig. 2.
Microbiota quantification
The included studies employed various sequencing workflows to estimate the gut microbiota composition. Three studies performed shotgun metagenomic sequencing [36, 38, 45], while all other studies performed 16s rRNA gene sequencing.
Microbial diversity
Six out of sixteen studies tested the association between alpha diversity/richness and connectivity [27, 28, 30,31,32,33, 35, 36], and one out of sixteen studies assessed beta diversity [35].
Microbial composition
Three studies performed microbiota-based clustering [27, 28, 30, 31, 34], and all but two studies assessed microbial abundance. Additional methodological information regarding sample collection and data processing is presented in Table 2.
Connectivity quantification
Functional connectivity assessment
Fourteen out of sixteen studies assessed functional connectivity, of which twelve used resting-state functional magnetic resonance imaging (rs-fMRI), one used task-based fMRI [33], and one used resting-state functional near-infrared spectroscopy (fNIRS) [36].
Structural connectivity assessment
Four out of fourteen studies assessed structural connectivity using diffusion tensor imaging (DTI) [28, 34, 39, 40]. Additional information about the metrics used to compute connectivity, and the a priori selection of brain regions, networks, or white matter tracts for each study is provided in Table 2.
Statistical analysis on microbiota-connectivity association
The included studies employed various statistical methods to explore the association between microbiota and brain connectivity, including linear regression/ANCOVA [32, 33, 35, 36, 46], (permutation) ANOVA [27, 28, 38, 44], (partial) correlation [27,28,29,30,31, 34, 39, 40, 43, 45] and tripartite network analysis, based on correlations [41]. Three studies used packages for differential abundance testing, including sparse partial least squares discriminant analysis (sPLS-DA) [34, 44], linear discriminant analysis effect size (LefSE) [36], microbiome multivariable association with linear models (MaAsLin2) [36], and differential gene expression analysis based on the negative binomial distribution (DESeq2) [44]. Finally, linked ICA [37], spatial canonical correlation analysis (sCCA) [33] and Parametric Empirical Bayes (PEB) [33] analysis were employed by one study each.
Synthesized results (Table 3)
Results on the microbiota level are discussed first, starting with findings in alpha- and beta diversity, followed by findings in microbial composition (clusters and abundance). Next, results on the brain connectivity level are discussed, starting with findings in functional connectivity, followed by findings in structural connectivity. Finally, the specificity of the association between the microbiota and connectivity is discussed. A detailed summary of the findings per individual study is available in the Supplementary Materials.
Gut microbiome
Microbial diversity
Six out of sixteen studies assessed microbial richness and/or alpha diversity in association with brain connectivity [27,28,29, 32, 33, 35, 36, 44]. Four out of six studies assessed a measure of microbial richness (number of OTUs, Sobs, Ace and/or Chao), all of which report an association with at least one brain connection or network [27,28,29, 32, 35, 36]. All six studies assessed a measure of alpha diversity (Shannon or Simpson), of which four studies reported an association with at least one brain connection or network [27,28,29, 33, 35, 36] (Fig. 3A, Supplementary Table 1). Only one study assessed a measure of beta diversity (weighted UniFrac), and reported an association with functional brain connectivity [32]. As there is only one study assessing this, it does not warrant interpreting at this stage (Supplementary Table 1). A comprehensive overview of the results per study can be found in Table 3.
Graphic summary of the reported associations between microbial diversity and functional connectivity (A), and between microbial abundances and functional connectivity (B). Each connection in the chord diagram reflects a reported association between the abundance of that genus or diversity measure and functional connectivity. Functional connectivity is aggregated to the network level, including intra-/ and internetwork connectivity and connectivity of individual constituents of a particular network, to aid visualization. Of note, the graphical overview displays the absolute number of reported associations, skewing it towards genera and connectivity networks that are studied more frequently (a consequence of the selection bias in the reported studies). ACC anterior cingulate cortex, DMN default mode network, ECN executive control network, FPN frontoparietal network, SMN sensorimotor network, SN salience network.
Microbial composition: clustering
Three out of sixteen studies performed clustering of the microbiome to obtain microbiome clusters/enterotypes. Of those, two studies identified two clusters: one high in Bacteroides and one high in Prevotella [30, 31, 34]. One study identified three clusters: a Bacteroides-high, Prevotella-high, and Ruminococcus-high cluster [27,28,29]. All three studies reported at least one association with connectivity, either functional [27,28,29,30,31] or structural [28, 29, 34] (see Table 3). Given the low number of studies, variability in number of clusters, and uncertainty about what the clusters consist of, it remains difficult to interpret the findings and draw conclusions about potential consistency (Supplementary Table 1).
Microbial composition: abundance
Thirteen out of sixteen studies assessed microbial abundance on the genus level, all of which report at least one significant association with functional and/or structural brain connectivity (for an overview, see Supplementary Table 1). The genera Bacteroides, reported in nine out of eleven studies [32, 33, 36,37,38, 40, 42,43,44], and Prevotella, reported in six out of ten studies [30,31,32, 34, 36,37,38], were most consistently associated with brain connectivity. Additionally, genera within the order Clostridiales were also repeatedly reported in association with brain connectivity. Within this order, the genus Ruminococcus was most consistently reported (in four out of seven studies [30, 31, 33, 37, 44]), followed by Blautia (in three out of five studies [37, 38, 41]). Other studies meeting the criteria for consistency include Collinsella (three out of five studies [30, 36, 38]), Enterococcus [36, 38], and Alistipes [38, 44] (each reported in two out of five studies) and Bifidobacterium (two out of three studies [36, 37]). These genera were associated with regions distributed throughout the brain, both on a region-to-region and network level. A comprehensive overview of the results per study can be found in Table 3.
Brain connectivity
Functional connectivity
Fourteen out of sixteen studies assessed the association between functional connectivity and the gut microbiota, either employing a seed-based (i.e., region-to-region or region-to-voxel) or ICA-based (i.e. resting-state networks) approach. All functional connections, seed-based and ICA-based were aggregated to a network level to improve interpretability (Supplementary Table 2). Salience network connectivity was consistently reported in association with the gut microbiome (nine out of ten studies). Particularly the connectivity of the anterior cingulate cortex (ACC) [27,28,29,30,31, 33, 35, 37, 38] and insula [27,28,29, 32, 33, 35, 38, 41] (each reported in six out of eight studies) were frequently reported. Additionally, amygdala connectivity (three out of six studies [30, 31, 35, 38]) and ICA-based salience network connectivity (two out of three studies [37, 38]) met the criterion of consistency on the brain connectivity level. The default mode network (DMN) was reported in association with the gut microbiota in seven out of nine studies: six out of six studies reported associations between ICA-based DMN-connectivity and the gut microbiota [27,28,29,30,31, 36,37,38, 42], and associations with the precuneus were reported in three out of six studies [27,28,29, 38, 44]. Finally, the frontoparietal network (FPN) was consistently associated with the gut microbiota (reported in seven out of nine studies). Particularly, ICA-based FPN-connectivity (four out of four studies [27,28,29, 36,37,38]) and connectivity of the dorsolateral prefrontal cortex (three out of six studies [27,28,29, 38, 42]) and inferior parietal lobe (two out of four studies [30, 31, 35]) were reported with higher frequency. Finally, regions within the sensorimotor network were reported in four out of seven studies (no specific regions within this network) [27,28,29, 35, 38, 41] and the superior temporal gyrus was associated with the gut microbiota in three out of six studies [27,28,29, 38, 40]. These connectivity networks were exhibiting associations with a wide range of diversity indices and genera abundances. A comprehensive overview of the findings per study can be found in Table 3 and Supplementary Table 2.
Structural connectivity
Three out of sixteen studies assessed structural connectivity, all of which reported at least one association with gut microbiota diversity or composition [28, 29, 34, 39]. However, the small number of studies and the variability in how structural connectivity was quantified and labeled, precludes meaningful interpretation at this time (Table 3, Supplementary Table 1).
Specificity of the microbiota-connectivity associations
Despite observing recurring patterns in findings on the gut microbiota and brain connectivity level, patterns are not evident in the association between microbiota and connectivity. None of the microbiota-connectivity associations were reported in at least fifty percent of the studies, thus not meeting the consistency criterion. Instead, diversity indices and microbial abundances were associated with a widespread set of brain regions and networks without any discernible pattern emerging. That is, the association between the gut microbiota and brain connectivity was non-specific (Fig. 3, Supplementary Tables 1 and 2).
Quality assessment
A majority of the studies were rated as ‘Fair’ (n = 9), four studies were rated as ‘Poor’ and only three studies were rated as ‘Good’. The primary source of bias was related to incomplete reporting of the methods, including a lack of detail in the description of the recruitment procedure, microbiota data handling, and description of the used statistical methods. Additionally, about a third of the studies did not correct for the effects of key confounders (i.e., age and sex). The quality assessment and explanatory notes per study are shown in Supplementary Table 6.
Discussion
Summary
A qualitative systematic synthesis of the available study findings shows associations between the gut microbiota and brain connectivity. On the microbiota level, a majority of the studies reported an association between microbial richness or diversity and connectivity. In terms of genus abundance, the genera Bacteroides, Prevotella, Ruminococcus, Blautia, and Collinsella were reported with the highest consistency, followed by the genera Enterococcus, Alistipes, and Bifidobacterium. For functional brain connectivity, the highest level of consistency was found for the DMN, FPN, and salience network (particularly the insula and ACC). There were too few studies assessing microbial clusters or structural connectivity to draw definite conclusions at this time. Moreover, although some microbial genera or brain networks were reported with higher frequency, there was no specificity in the association between the gut microbiome and brain connectivity, and a majority of the findings, both in microbiota and brain, were inconsistent and poorly or not replicated across studies. Since some of the included studies did not provide information on the direction of the associations, and as the interpretation of the direction depends on the specific brain connection/network involved, we decided not to consider the directionality in our synthesis. Consequently, the conclusion is limited to the observation of a potential association between the gut microbiota and brain connectivity.
Finally, the studies differed in their target population, focusing on a range of different diseases. Considering the large methodological differences between studies as well as a lack of direct comparisons between the case and control participants, is presently not possible to draw conclusions about differences in the microbiota-connectivity associations between cases and controls or between different disease populations.
Below we will first discuss the findings in relation to neurocognitive functioning and possible functional mechanisms. Finally, we will address study aspects that can explain inconsistencies in the findings, and which need to be considered to further advance the field.
Neurocognitive functions associated with microbiota
The current findings suggest an involvement of brain networks involved in emotion-related cognition and executive functioning. The association between the gut microbiota and emotion-related functioning is a recurring topic in gut-brain research. Several of the included studies found associations between the microbiota composition and levels of anxiety, depression, or negative affect, measured using questionnaires [30, 31, 34]. Moreover, several of the brain structures associated with the gut microbiota have a putative emotion-related function. For example, the insula, whose connectivity was associated with alpha diversity [32, 33, 35] and with the abundance of the genera Roseburia and Bacteroides [28, 32, 41], is involved in socio-emotional processing, and insular brain damage can result in apathy and anxiety [47]. Moreover, the insula is involved in interoception, both emotional and visceral [8, 9]. The amygdala, whose connectivity was associated with alpha diversity and Roseburia abundance [31, 35], is one of the brain’s main emotion-processing structures, which is implicated in, among others, autism and anxiety [48]. Finally, the DMN, whose connectivity was associated with microbial diversity and the abundance of a wide range of bacterial genera (Fig. 3), is involved in mind-wandering and conceptualization of emotions [49]. Moreover, DMN connectivity is altered in depression, a disorder characterized by impairments in emotion regulation [50]. Interestingly, Kelsey and colleagues [36] reported that stronger intranetwork connectivity of the homologous-interhemispheric network in newborns mediated the association between higher alpha diversity and behavioral temperament, behavior that is predictive of anxiety and depression in adulthood [51].
Another set of findings suggests that the gut microbiota may be associated with executive functioning. Cai and colleagues [27] reported that higher internetwork connectivity between the FPN and visual networks and lower internetwork connectivity between the dorsal attention, and visual networks mediated the association between higher alpha diversity and better working memory and attention. In addition, the association between Prevotella-, Bacteroides-, and Ruminococcus-high clusters and response inhibition was mediated by connectivity of the orbitofrontal cortex (part of the FPN), a region involved in (emotional) decision making [52]. Specifically, individuals in the Prevotella and Ruminococcus-high clusters exhibited stronger orbitofrontal connectivity, which was associated with poorer response inhibition. Consistent with these findings, several functional networks associated with gut microbiota diversity are involved in executive functioning. For example, the FPN (or its individual constituents) was associated with the abundance of, among others, the genera Prevotella, Bacteroides, and Blautia [36, 37, 46]. This network is mainly involved in executive control, encapsulating processes related to response inhibition, attention, and working memory [6, 53].
Altogether, the associations between the gut microbiota and the brain, described in this review, indirectly link the gut microbiota with processes related to emotion and executive functioning through brain connectivity patterns. However, in the absence of direct, empirically tested, associations with cognition and behavior, this should be interpreted with caution. These speculations should be validated using, for example, larger-scale mediation analyses to disentangle the intermediate role of brain connectivity in the association between the gut microbiota and behavior.
Microbial function and functional pathways
Up to now, studies mostly report taxonomic findings, which are not appropriate to deduce functional pathways. Nevertheless, we can still leverage the findings from previous studies to speculate about potential mechanisms, which can be used as hypothesis-generating ideas as the field moves towards more function-focused microbiome research. Based on the taxonomic findings, one possible communication pathway would be through the production of short-chain fatty acids (SCFAs; e.g., acetate, propionate, and butyrate). For example, species within the genera Prevotella and Bacteroides, whose abundances on a genus level were associated with connectivity of a widely distributed set of brain regions and networks (Fig. 3), are described as main propionate and acetate-producing species [54]. Moreover, species in the genus Blautia, whose abundance on a genus level was associated with the connectivity of sensorimotor regions as well as network connectivity of the DMN and executive control network, are described to have propionate-producing properties [54]. Finally, species in the genus Roseburia, whose abundance on a genus level was associated with insular, amygdala, and DMN connectivity, are among the main butyrate-producing species. Among others, SCFAs possess the ability to modulate immune activation (reviewed in [55]). In line with this, the association between Roseburia abundance and functional connectivity of the DMN and amygdala, reported by Wang and colleagues [30] and Zheng and colleagues [31], was mediated by levels of the pro-inflammatory cytokines interleukin-6 and tumor necrosis factor alpha, suggesting a role for the immune signaling pathway in the microbiota-brain connectivity communication, possibly through the production of butyrate. The involvement of this pathway may go beyond the gut, as there is preliminary evidence for a role of the immune signaling pathway in the association between functional connectivity and the oral microbiome as well [56]. Other ways through which gut-synthesized SCFAs may modulate gut-brain communication, and hereby brain connectivity, could be by positively or negatively affecting the integrity of the intestinal and blood-brain barriers [55, 57, 58], or by traveling directly through the blood-brain barrier to the CNS [59,60,61], although – similar to the above-suggested role of the SCFAs – evidence directly supporting this is yet lacking.
The gut microbiota may also affect gut-brain communication through the production of neurotransmitters and their precursors [46]. For example, species within the genera Bacteroides and Bifidobacterium and species within genera Ruminococcus and Blautia, whose abundances were associated with connectivity in a widespread set of brain regions and networks, are major modulators of GABA and serotonin availability in the gut [62,63,64]. There are several proposed pathways through which gut-derived neuroactive compounds can affect the brain. For example, there is evidence showing that both GABA and serotonin have immunomodulatory properties [65, 66], and both GABA and serotonin receptors have been located on vagal afferents to the CNS, proposing a role for the vagal signaling pathway [67, 68]. However, in the absence of direct evidence, the involvement of these pathways remains hypothetical.
Altogether, based on the taxonomic findings we can speculate about potential mechanisms. However, most taxonomic findings are reported on a genus level due to the constraints of 16s rRNA sequencing. A certain genus, and even a single species within a genus, can contribute to multiple metabolite pathways. As such, our speculations should be verified through pathway analysis, preferably using higher resolution metagenomics data coupled with, when possible, bacterial culture-functional studies.
Recommendations for future studies
The current review identified several links between microbial genera and brain connectivity, but there is low specificity in the associations between the gut and brain. Additionally, despite identifying some recurring patterns, a majority of the findings, both in microbiota and brain, were inconsistent and poorly replicated across studies. This may suggest a complex multifaceted relationship between microbial composition and brain connectivity, but it is likely that inconsistencies are at least partially methodology-driven. The limitations identified in this review underscore the need to harmonize the methodological approaches currently applied to microbiome research and (functional) brain connectivity analysis. In the following section, we will discuss important study aspects that may explain inconsistencies in the findings, and which need to be considered to further advance the field.
Methodological comparability
The number of studies investigating the microbiota-gut-brain axis is rapidly growing, and in recent years there have been major technological advances in the field. Nevertheless, there is no golden standard on how to collect, process, and analyze microbiome data. As a result, there is high inter-study variability in laboratory processing (e.g., variable region of the 16S rRNA gene and sequencing platform), pre-processing (e.g., taxonomic database, processing pipeline and prevalence filtering, and data transformation) and statistical analysis approach. The number of observed taxa and statistical outcomes can change considerably depending on the collection, storage, bioinformatic pipeline, and statistical test used [69,70,71,72]. Therefore, to make reliable between-study comparisons it is essential to harmonize the methodological approach. To achieve this, researchers should follow standardized reporting guidelines, such as the STORMS checklist [25], on how the microbiome data was processed, follow standardized processing pipelines where possible, apply consistent and appropriate statistical approaches for analysis (e.g., accounting for the zero-inflated and compositional nature of the data), and report findings (including effect-sizes) in a complete and transparent manner.
While reducing the multiple comparison problem, additional sources of inter-study variability come from a priori selection of microbial genera and brain regions (e.g., seed-based resting-state analyses). Some of these genera – for example Prevotella and Bacteroides – or brain networks – for example the DMN – are of common interest in studies focusing on the gut-brain communication, increasing the likelihood that the effects are observed or amplified as a result of selection bias. Moreover, a priori selection, both in taxonomy and brain regions, hinders meaningful comparison between study findings. At the current stage of the field, using a data-driven approach to study the association between the gut microbiota and brain connectivity will help with the identification of patterns of associations in the data without biasing the findings towards pre-existing assumptions. However, data-driven approaches have a higher risk of identifying spurious associations, and findings may be more challenging to interpret. Therefore, once consistent findings have been established using data-driven approaches, the field could move towards a hypothesis-driven framework that offers clearer research questions and facilitates the interpretation of the findings.
Data integration and multivariate approaches
Data analysis should better acknowledge the inherent complexities of both the gut microbiota and brain connectivity data, as this will lead to better integration of the two domains. Both the gut microbiota and the brain are complex systems, characterized by intricate relationships and interconnections [73, 74]. At the current time, most studies investigating the association between the gut microbiota and brain connectivity do so using simple bivariate association analyses, even though this is likely an oversimplification of the existing association. As a result, information concealed in the relationship within and between multiple variables (i.e., interaction effects) is lost [75]. Although several studies opted for a multivariate approach for one of the two domains (i.e., differential abundance testing for the microbiome, and ICA or sparse canonical correlation analysis for brain connectivity), only one study used an integrative multivariate approach, integrating microbiota and connectivity data [37]. The authors identified several multivariate associations between microbial clusters and functional network connectivity.
Future studies should make an effort to integrate data from the gut microbiota and brain connectivity. Possible approaches are the Linked Independent Component Analysis (as applied in ref. [37]) or co-inertia analysis (as applied in ref. [38]), which allow for simultaneous factorization of data from different domains. Moreover, the systems biology approach focuses on multivariate interactions in biological systems rather than exploring each modality in parallel [76]. There are already different systems biology software programs available [77]. However, they are mainly utilized to integrate the so-called ‘omics’ techniques and, so far, there is little contribution of macroscale neurobiological measures. Nevertheless, the field of human connectomics was already proposed as an extension of systems biology to macroscale neuroscience [78,79,80] and could provide insight into the complex microbiota-connectivity interactions.
Another advantage of integrating more (-omics) domains would be improved mechanistic insight into the human gut-brain axis. There have been numerous studies on possible mechanisms underlying gut-brain communication, both preclinical and in humans (reviewed in e.g. ref. [81]). Usually, mechanisms are studied on a molecular level, without assessing how such processes would affect the brain on a larger scale. To date, only Wang and colleagues [30] and Zheng and colleagues [31] explored the mediation effect of immune activation as an underlying mechanism in the gut-brain connectivity association. Although we can speculate about underlying mechanisms based on taxonomic findings, it would be more informative to directly integrate mechanistic data (potentially) related to brain connectivity. For example, metagenomics can complement 16s rRNA sequencing to provide information about the functional potential of the present microbial taxa, metabolomics could provide information about bacterial metabolites that are present in the gut (e.g., SCFAs), and inflammatory markers in the blood could be investigated to provide information about immune activation.
Confounders
There is currently no agreement on which factors that affect the gut should be considered and corrected for in statistical analyses. This is also reflected in the way studies correct for confounders: approximately two-thirds consider age and sex as key confounders. Recent antibiotic use is used as an exclusion criterion by three-fourths of the studies, whereas only one study corrects for smoking. Surprisingly, none of the studies correct for ethnicity [82], medication use [83], or diet [17, 84], even though several studies have demonstrated their effects on the gut microbiome and/or brain. To better understand how these factors affect the association between microbiota and brain connectivity, we recommend that future studies conduct sensitivity analyses. This will – besides establishing the robustness of the findings – help to identify the (often environmental) factors that influence the association between the gut microbiota and brain connectivity and potentially inform the development of future treatment strategies.
Study characteristics and group comparisons
Currently, studies assessing the association between the gut microbiota and brain connectivity include a wide range of target populations, from healthy individuals to various disease populations. While these studies could provide valuable information about disease pathophysiology, the field lacks a reference to which associations in disease populations can be compared. Investigating the microbiota-connectivity associations in large population cohorts can provide such references and aid in the interpretation of study findings. Moreover, with a case-control study design, it is possible to test differences in the microbiota-connectivity association between groups. However, only one study has performed such analyses [41], with other studies only focusing on the microbiota-connectivity association in cases. Performing such comparative analyses will help in clarifying whether there are differences in the associations between different disease populations, and improve our ability to draw meaningful conclusions about group differences.
Once such methodological issues are resolved, it would be valuable to have longitudinal microbiota-connectivity association studies, for example in developing children or elderly individuals, to see how microbial development covaries with neurodevelopment, whether microbiota-connectivity associations vary over time and how this could be related to neurodevelopmental or neurodegenerative disorders.
Association not causation
Most studies discussed here were observational, from which it is not possible to infer causality nor directionality. To advance research on the human MGBA, specifically for the development of gut microbiota-targeting approaches in the treatment of brain-related disorders, it is important to establish causality and directionality. Assessing directionality and causality in human trials remains a costly challenge, in which randomized controlled trials in combination with appropriate statistical models such as mediation or Mendelian randomization can assist [85, 86]. For example, results from mediation analyses point towards gut-to-brain signaling, showing that immune activation (one of the MGBA signaling pathways) partially mediated the association between microbial abundance and functional brain connectivity [27, 36]. Such studies should assess if and how an intervention affects the gut microbiota or brain connectivity, which microbes are causally involved, if intervention-induced changes in the microbiota result in altered brain connectivity, and – importantly – whether this is also reflected in altered cognition and behavior.
Conclusion
In this systematic review, we identified several genera as well as brain regions and networks that were repeatedly associated in microbiota-brain connectivity analyses, showing the potential of employing brain connectivity measures to gain insight into gut-brain communication. At the same time, there is still limited evidence for specificity in the microbiota-brain connectivity association. Current methodological limitations, including high inter-study variability in methodology and target population, small sample sizes, the a priori selection of microbial genera and brain regions of interest, and the statistical approaches used, introduce bias and thus contribute to the inconsistent findings. We know the gut-brain communication is multidimensional, and therefore a more systematic and harmonized methodology, which acknowledges this complexity, is key to further unraveling how the gut and brain communicate. To enhance comparability and replicability, future research should focus on further standardizing processing pipelines and employ data-driven multivariate analysis approaches. Moreover, interventional studies can help to clarify the causality and directionality of the reported findings.
References
Sporns O, Tononi G, Kötter R. The human connectome: a structural description of the human brain. PLoS Comput Biol. 2005;1:245–51.
Suárez LE, Markello RD, Betzel RF, Misic B. Linking structure and function in macroscale brain networks. Trends Cogn Sci. 2020;24:302–15.
Damoiseaux JS, Rombouts SARB, Barkhof F, Scheltens P, Stam CJ, Smith SM, et al. Consistent resting-state networks across healthy subjects. Proc Natl Acad Sci. 2006;103:13848–53.
Smitha KA, Akhil Raja K, Arun KM, Rajesh PG, Thomas B, Kapilamoorthy TR, et al. Resting state fMRI: a review on methods in resting state connectivity analysis and resting state networks. Neuroradiol J. 2017;30:305.
Seeley WW. The salience network: a neural system for perceiving and responding to homeostatic demands. J Neurosci. 2019;39:9878–82.
Marek S, Dosenbach NUF. The frontoparietal network: function, electrophysiology, and importance of individual precision mapping. Dialogues Clin Neurosci. 2018;20:133–40.
Smallwood J, Bernhardt BC, Leech R, Bzdok D, Jefferies E, Margulies DS. The default mode network in cognition: a topographical perspective. Nat Rev Neurosci. 2021;22:503–13.
Smith R, Alkozei A, Bao J, Smith C, Lane RD, Killgore WDS. Resting state functional connectivity correlates of emotional awareness. Neuroimage. 2017;159:99–106.
Longarzo M, Quarantelli M, Aiello M, Romano M, Del Prete A, Cimminiello C, et al. The influence of interoceptive awareness on functional connectivity in patients with irritable bowel syndrome. Brain Imaging Behav. 2017;11:1117–28.
Gao Y, Shuai D, Bu X, Hu X, Tang S, Zhang L, et al. Impairments of large-scale functional networks in attention-deficit/hyperactivity disorder: a meta-analysis of resting-state functional connectivity. Psychol Med. 2019;49:2475–85.
Dong D, Wang Y, Chang X, Luo C, Yao D. Dysfunction of large-scale brain networks in schizophrenia: a meta-analysis of resting-state functional connectivity. Schizophr Bull. 2018;44:168–81.
Collin G, Nieto-Castanon A, Shenton ME, Pasternak O, Kelly S, Keshavan MS, et al. Brain functional connectivity data enhance prediction of clinical outcome in youth at risk for psychosis. NeuroImage Clin. 2020;26:300–11.
Kochunov P, Jahanshad N, Marcus D, Winkler A, Sprooten E, Nichols TE, et al. Heritability of fractional anisotropy in human white matter: a comparison of Human Connectome Project and ENIGMA-DTI data. Neuroimage. 2015;111:300–11.
Adhikari BM, Jahanshad N, Shukla D, Glahn DC, Blangero J, Fox PT, et al. Comparison of heritability estimates on resting state fMRI connectivity phenotypes using the ENIGMA analysis pipeline. Hum Brain Mapp. 2018;39:4893–902.
Sharp PB, Sutton BP, Paul EJ, Sherepa N, Hillman CH, Cohen NJ, et al. Mindfulness training induces structural connectome changes in insula networks. Sci Rep. 2018 81. 2018;8:1–10.
Sezer I, Pizzagalli DA, Sacchet MD. Resting-state fMRI functional connectivity and mindfulness in clinical and non-clinical contexts: a review and synthesis. Neurosci Biobehav Rev. 2022;135:104583.
Jensen DEA, Leoni V, Klein-Flügge MC, Ebmeier KP, Suri S. Associations of dietary markers with brain volume and connectivity: a systematic review of MRI studies. Ageing Res Rev. 2021;70:101360.
Berding K, Vlckova K, Marx W, Schellekens H, Stanton C, Clarke G, et al. Diet and the microbiota-gut-brain axis: sowing the seeds of good mental health. Adv Nutr. 2021;12:1239–85.
Cryan JF, O’riordan KJ, Cowan CSM, Sandhu KV, Bastiaanssen TFS, Boehme M, et al. The microbiota-gut-brain axis. Physiol Rev. 2019;99:1877–2013.
Takiishi T, Fenero CIM, Câmara NOS. Intestinal barrier and gut microbiota: Shaping our immune responses throughout life. Tissue barriers. 2017;5:e1373208.
Taylor AM, Thompson SV, Edwards CG, Musaad SMA, Khan NA, Holscher HD. Associations among diet, the gastrointestinal microbiota, and negative emotional states in adults. Nutr Neurosci. 2020;23:983–92.
Fattorusso A, Di Genova L, Dell’isola GB, Mencaroni E, Esposito S. Autism spectrum disorders and the gut microbiota. Nutrients. 2019;11:521.
Huang TT, Lai JB, Du YL, Xu Y, Ruan LM, Hu SH. Current understanding of gut microbiota in mood disorders: an update of human studies. Front Genet. 2019;10:1–12.
National Institutes of Health. Quality assessment tool for observational cohort and cross-sectional studies. 2013. Available from: https://www.nhlbi.nih.gov/health-topics/study-quality-assessment-tools.
Mirzayi C, Renson A, Furlanello C, Sansone SA, Zohra F, Elsafoury S, et al. Reporting guidelines for human microbiome research: the STORMS checklist. Nat Med. 2021;27:1885–92.
Poldrack RA, Fletcher PC, Henson RN, Worsley KJ, Brett M, Nichols TE. Guidelines for reporting an fMRI study. Neuroimage. 2008;40:409–14.
Cai H, Wang C, Qian Y, Zhang S, Zhang C, Zhao W, et al. Large-scale functional network connectivity mediate the associations of gut microbiota with sleep quality and executive functions. Hum Brain Mapp. 2021;42:3088–101.
Zhu J, Wang C, Qian Y, Cai H, Zhang S, Zhang C, et al. Multimodal neuroimaging fusion biomarkers mediate the association between gut microbiota and cognition. Prog Neuro Psychopharmacol Biol Psychiatry. 2022;113:110468.
Zhang S, Xu X, Li Q, Chen J, Liu S, Zhao W, et al. Brain network topology and structural–functional connectivity coupling mediate the association between gut microbiota and cognition. Front Neurosci. 2022;16:1–17.
Wang YF, Zheng LJ, Liu Y, Ye YB, Luo S, Lu GM et al. The gut microbiota-inflammation-brain axis in end-stage renal disease: perspectives from default mode network. Theranostics. 2019. https://doi.org/10.7150/thno.35387.
Zheng LJ, Lin L, Zhong J, Zhang Z, Ye YB, Zhang XY, et al. Gut dysbiosis-influence on amygdala-based functional activity in patients with end stage renal disease: a preliminary study. Brain Imaging Behav. 2020;14:2731–44.
Curtis K, Stewart CJ, Robinson M, Molfese DL, Gosnell SN, Kosten TR, et al. Insular resting state functional connectivity is associated with gut microbiota diversity. Eur J Neurosci. 2019;50:2446–52.
Hall CV, Harrison BJ, Iyer KK, Savage HS, Zakrzewski M, Simms LA, et al. Microbiota links to neural dynamics supporting threat processing. Hum Brain Mapp. 2022;43:733–49.
Tillisch K, Mayer EA, Gupta A, Gill Z, Brazeilles R, Le Nevé B, et al. Brain structure and response to emotional stimuli as related to gut microbial profiles in healthy women. Psychosom Med. 2017;79:905–13.
Gao W, Salzwedel AP, Carlson AL, Xia K, Azcarate-Peril MA, Styner MA, et al. Gut microbiome and brain functional connectivity in infants-a preliminary study focusing on the amygdala. Psychopharmacology. 2019;236:1641–51.
Kelsey CM, Prescott S, McCulloch JA, Trinchieri G, Valladares TL, Dreisbach C, et al. Gut microbiota composition is associated with newborn functional brain connectivity and behavioral temperament. Brain Behav Immun. 2021;91:472–86.
Kohn N, Szopinska-Tokov J, Llera Arenas A, Beckmann CF, Arias-Vasquez A, Aarts E. Multivariate associative patterns between the gut microbiota and large-scale brain network connectivity. Gut Microbes. 2021;13:2006586.
Li Z, Lai J, Zhang P, Ding J, Jiang J, Liu C, et al. Multi-omics analyses of serum metabolome, gut microbiome and brain function reveal dysregulated microbiota-gut-brain axis in bipolar depression. Mol Psychiatry. 2022;27:1–13.
Ahluwalia V, Betrapally NS, Hylemon PB, White MB, Gillevet PM, Unser AB, et al. Impaired gut-liver-brain axis in patients with cirrhosis. Sci Rep. 2016;6:1–11.
Jacobs JP, Gupta A, Bhatt RR, Brawer J, Gao K, Tillisch K, et al. Cognitive behavioral therapy for irritable bowel syndrome induces bidirectional alterations in the brain-gut-microbiome axis associated with gastrointestinal symptom improvement. Microbiome. 2021;9:1–14.
Labus JS, Osadchiy V, Hsiao EY, Tap J, Derrien M, Gupta A, et al. Evidence for an association of gut microbial Clostridia with brain functional connectivity and gastrointestinal sensorimotor function in patients with irritable bowel syndrome, based on tripartite network analysis. Microbiome. 2019;7:1–15.
Strandwitz P, Kim KH, Terekhova D, Liu JK, Sharma A, Levering J, et al. GABA-modulating bacteria of the human gut microbiota. Nat Microbiol. 2018;4:396–403.
Dong TS, Guan M, Mayer EA, Stains J, Liu C, Vora P, et al. Obesity is associated with a distinct brain-gut microbiome signature that connects Prevotella and Bacteroides to the brain’s reward center. Gut Microbes. 2022;14:1–17.
Dong TS, Gupta A, Jacobs JP, Lagishetty V, Gallagher E, Bhatt RR, et al. Improvement in uncontrolled eating behavior after laparoscopic sleeve gastrectomy is associated with alterations in the brain–gut–microbiome axis in obese women. Nutrients. 2020;12:1–16.
Hong J, Bo T, Xi L, Xu X, He N, Zhan Y, et al. Reversal of functional brain activity related to gut microbiome and hormones after VSG surgery in patients with obesity. J Clin Endocrinol Metab. 2021;106:3619–33.
Strandwitz P. Neurotransmitter modulation by the gut microbiota. Brain Res. 2018;1693:128–33.
Uddin LQ, Nomi JS, Hébert-Seropian B, Ghaziri J, Boucher O. Structure and function of the human insula. J Clin Neurophysiol. 2017;34:300–6.
Janak PH, Tye KM. From circuits to behaviour in the amygdala. Nature. 2015;517:284–92.
Satpute AB, Lindquist KA. The default mode network’s role in discrete emotion. Trends Cogn Sci. 2019;23:851–64.
Yan C-G, Chen X, Li L, Castellanos FX, Bai T-J, Bo Q-J, et al. Reduced default mode network functional connectivity in patients with recurrent major depressive disorder. Proc Natl Acad Sci USA. 2019;116:9078–83.
Tang A, Crawford H, Morales S, Degnan KA, Pine DS, Fox NA. Infant behavioral inhibition predicts personality and social outcomes three decades later. Proc Natl Acad Sci. 2020;117:9800–7.
Rudebeck PH, Rich EL. Current biology orbitofrontal cortex. Curr Biol. 2018;28:1075–95.
Niendam TA, Laird AR, Ray KL, Dean YM, Glahn DC, Carter CS. Meta-analytic evidence for a superordinate cognitive control network subserving diverse executive functions. Cogn Affect Behav Neurosci. 2012;12:241–68.
Louis P, Flint HJ. Formation of propionate and butyrate by the human colonic microbiota. Environ Microbiol. 2017;19:29–41.
Venegas DP, De La Fuente MK, Landskron G, González MJ, Quera R, Dijkstra G, et al. Short chain fatty acids (SCFAs)mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front Immunol. 2019;10:277.
Lin D, Hutchison KE, Portillo S, Vegara V, Ellingson JM, Liu J, et al. Association between the oral microbiome and brain resting state connectivity in smokers. Neuroimage. 2019;200:121–31.
Silva YP, Bernardi A, Frozza RL. The role of short-chain fatty acids from gut microbiota in gut-brain communication. Front Endocrinol. 2020;11:1–14.
Hoyles L, Snelling T, Umlai UK, Nicholson JK, Carding SR, Glen RC, et al. Microbiome–host systems interactions: Protective effects of propionate upon the blood–brain barrier. Microbiome. 2018;6:1–13.
Frost G, Sleeth ML, Sahuri-Arisoylu M, Lizarbe B, Cerdan S, Brody L, et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat Commun. 2014;5:1–11.
Yoo DY, Kim W, Nam SM, Kim DW, Chung JY, Choi SY, et al. Synergistic effects of sodium butyrate, a histone deacetylase inhibitor, on increase of neurogenesis induced by pyridoxine and increase of neural proliferation in the mouse dentate gyrus. Neurochem Res. 2011;36:1850–7.
Erny D, De Angelis ALH, Jaitin D, Wieghofer P, Staszewski O, David E, et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat Neurosci. 2015;18:965–77.
Otaru N, Ye K, Mujezinovic D, Berchtold L, Constancias F, Cornejo FA, et al. GABA production by human intestinal Bacteroides spp.: prevalence, regulation, and role in acid stress tolerance. Front Microbiol. 2021;12:1–14.
Barrett E, Ross RP, O’Toole PW, Fitzgerald GF, Stanton C. γ-Aminobutyric acid production by culturable bacteria from the human intestine. J Appl Microbiol. 2012;113:411–7.
Yano JM, Yu K, Donaldson GP, Shastri GG, Ann P, Ma L, et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell. 2015;161:264–76.
Bjurstöm H, Wang JY, Ericsson I, Bengtsson M, Liu Y, Kumar-Mendu S, et al. GABA, a natural immunomodulator of T lymphocytes. J Neuroimmunol. 2008;205:44–50.
Cloëz-Tayarani I, Changeux J-P. Nicotine and serotonin in immune regulation and inflammatory processes: a perspective. J Leukoc Biol. 2007;81:599–606.
Breit S, Kupferberg A, Rogler G, Hasler G. Vagus nerve as modulator of the brain-gut axis in psychiatric and inflammatory disorders. Front Psychiatry. 2018;9:1–15.
Cawthon CR, de La Serre CB. Gut bacteria interaction with vagal afferents. Brain Res. 2018;1693:134–9.
Boers SA, Jansen R, Hays JP. Understanding and overcoming the pitfalls and biases of next-generation sequencing (NGS) methods for use in the routine clinical microbiological diagnostic laboratory. Eur J Clin Microbiol Infect Dis. 2019;38:1059–70.
Nearing JT, Douglas GM, Hayes MG, MacDonald J, Desai DK, Allward N, et al. Microbiome differential abundance methods produce different results across 38 datasets. Nat Commun. 2022;13:1–16.
Teng F, Darveekaran Nair SS, Zhu P, Li S, Huang S, Li X, et al. Impact of DNA extraction method and targeted 16S-rRNA hypervariable region on oral microbiota profiling. Sci Rep. 2018;8:1–12.
Szopinska-Tokov J, Bloemendaal M, Boekhorst J, Hermes GDA, Ederveen T, Vlaming P et al. A comparison of bioinformatics pipelines for compositional analysis of the human gut microbiome (submitted). Preprint at https://www.biorxiv.org/content/10.1101/2023.02.13.528280v1. 2023.
Thursby E, Juge N. Introduction to the human gut microbiota. Biochem J. 2017;474:1823–36.
Kelly C, Biswal BB, Craddock RC, Castellanos FX, Milham MP. Characterizing variation in the functional connectome: promise and pitfalls. Trends Cogn Sci. 2012;16:181–8.
Lahat D, Adali T, Jutten C. Multimodal data fusion: an overview of methods, challenges, and prospects. Proc IEEE. 2015;103:1449–77.
Tavassoly I, Goldfarb J, Iyengar R. Systems biology primer: the basic methods and approaches. Essays Biochem. 2018;62:487–500.
De Souza HSP, Fiocchi C, Iliopoulos D. The IBD interactome: an integrated view of aetiology, pathogenesis and therapy. Nat Rev Gastroenterol Hepatol. 2017;14:739–49.
Bigler ED. Systems biology, neuroimaging, neuropsychology, neuroconnectivity and traumatic brain injury. Front Syst Neurosci. 2016;10:1–23.
Holzinger A, Haibe-Kains B, Jurisica I. Why imaging data alone is not enough: AI-based integration of imaging, omics, and clinical data. Eur J Nucl Med Mol Imaging. 2019;46:2722–30.
Sporns O. The human connectome: origins and challenges. Neuroimage. 2013;80:53–61.
Martin CR, Osadchiy V, Kalani A, Mayer EA. The brain-gut-microbiome axis. Cell Mol Gastroenterol Hepatol. 2018;6:133–48.
Deschasaux M, Bouter KE, Prodan A, Levin E, Groen AK, Herrema H, et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat Med. 2018;24:1526–31.
Imhann F, Vich Vila A, Bonder MJ, Lopez Manosalva AG, Koonen DPY, Fu J, et al. The influence of proton pump inhibitors and other commonly used medication on the gut microbiota. Gut Microbes. 2017;8:1–8.
Singh RK, Chang HW, Yan D, Lee KM, Ucmak D, Wong K, et al. Influence of diet on the gut microbiome and implications for human health. J Transl Med. 2017;15:1–17.
Lv BM, Quan Y, Zhang HY. Causal inference in microbiome medicine: principles and applications. Trends Microbiol. 2021;29:736–46.
Chaudhari SN, McCurry MD, Devlin AS. Chains of evidence from correlations to causal molecules in microbiome-linked diseases. Nat Chem Biol. 2021;17:1046–56.
Wang WL, Xu SY, Ren ZG, Tao L, Jiang JW, Zheng SS. Application of metagenomics in the human gut microbiome. World J Gastroenterol. 2015;21:803.
Hugerth LW, Andersson AF. Analysing microbial community composition through amplicon sequencing: from sampling to hypothesis testing. Front Microbiol. 2017;8:1561.
Emsell L, Van Hecke W, Tournier JD. Introduction to diffusion tensor imaging. In: Diffusion tensor imaging: a practical handbook. New York: Springer; 2016.
Bijsterbosch J, Smith SM, Beckmann CF. Introduction to resting state fMRI functional connectivity. In: Oxford Neuroimaging Primers. Oxford: Oxford University Press; 2017.
Beckmann CF, DeLuca M, Devlin JT, Smith SM. Investigations into resting-state connectivity using independent component analysis. Philos Trans R Soc B Biol Sci. 2005;360:1001–13.
Hevey D. Network analysis: a brief overview and tutorial. Heal Psychol Behav Med. 2018;6:301–28.
Acknowledgements
The research leading to these results received funding from the European Community’s Horizon 2020 research and innovation program through the Eat2beNICE (grant agreement no. 728018), CoCA (grant agreement no. 667302) and CANDY (grant agreement no. 847818) projects. Esther Aarts received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement no. 852189).
Author information
Authors and Affiliations
Contributions
DM conceptualized the study. DM and MB designed the study and performed the systematic literature search and data extraction. DM wrote the initial draft and visualized the findings. AAV and MB supervised the project. AAV, AE, DM and MB contributed to the manuscript’s revision and interpretation of the findings. All authors approved the final submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mulder, D., Aarts, E., Arias Vasquez, A. et al. A systematic review exploring the association between the human gut microbiota and brain connectivity in health and disease. Mol Psychiatry 28, 5037–5061 (2023). https://doi.org/10.1038/s41380-023-02146-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-023-02146-4
This article is cited by
-
Cellular interplay to 3D in vitro microphysiological disease model: cell patterning microbiota–gut–brain axis
Bio-Design and Manufacturing (2024)
-
Nutraceutical blends predict enhanced health via microbiota reshaping improving cytokines and life quality: a Brazilian double-blind randomized trial
Scientific Reports (2024)