Abstract
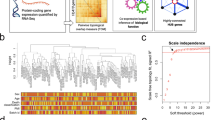
22q11.2 deletion is one of the strongest known genetic risk factors for schizophrenia. Recent whole-genome sequencing of schizophrenia cases and controls with this deletion provided an unprecedented opportunity to identify risk modifying genetic variants and investigate their contribution to the pathogenesis of schizophrenia in 22q11.2 deletion syndrome. Here, we apply a novel analytic framework that integrates gene network and phenotype data to investigate the aggregate effects of rare coding variants and identified modifier genes in this etiologically homogenous cohort (223 schizophrenia cases and 233 controls of European descent). Our analyses revealed significant additive genetic components of rare nonsynonymous variants in 110 modifier genes (adjusted P = 9.4E-04) that overall accounted for 4.6% of the variance in schizophrenia status in this cohort, of which 4.0% was independent of the common polygenic risk for schizophrenia. The modifier genes affected by rare coding variants were enriched with genes involved in synaptic function and developmental disorders. Spatiotemporal transcriptomic analyses identified an enrichment of coexpression between modifier and 22q11.2 genes in cortical brain regions from late infancy to young adulthood. Corresponding gene coexpression modules are enriched with brain-specific protein-protein interactions of SLC25A1, COMT, and PI4KA in the 22q11.2 deletion region. Overall, our study highlights the contribution of rare coding variants to the SCZ risk. They not only complement common variants in disease genetics but also pinpoint brain regions and developmental stages critical to the etiology of syndromic schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
This study is a secondary data analysis of the whole genome sequencing data of SCZ in 22q11.2DS generated by the International 22q11.2DS Brain and Behavior Consortium (IBBC). All summary statistics for the SCZ association of rare coding variants in the IBBC 22q11.2DS cohort are available at http://zdzlab.einsteinmed.edu/1/sz-22q.html. All predicted modifier genes are available in the supplementary material. Due to privacy concerns for our research participants, individual-level genetic data from the IBBC study of SCZ in 22q11.2DS are not publicly available; however, access to anonymized data can be requested from a qualified academic investigator to the IBBC Executive Committee, providing the data transfer is approved by the Institutional Review Board and regulated by a material transfer agreement.
Code availability
IGSP is publicly available at https://zenodo.org/record/1034362#.X-JWQNgzY2w. All other software used in our analyses is open source and described in “Methods”.
References
Edelmann L, Pandita RK, Morrow BE. Low-copy repeats mediate the common 3-Mb deletion in patients with velo-cardio-facial syndrome. Am J Hum Genet. 1999;64:1076–86.
Shaikh TH, Kurahashi H, Saitta SC, O’Hare AM, Hu P, Roe BA, et al. Chromosome 22-specific low copy repeats and the 22q11.2 deletion syndrome: genomic organization and deletion endpoint analysis. Hum Mol Genet. 2000;9:489–501.
Tezenas Du Montcel S, Mendizabai H, Ayme S, Levy A, Philip N. Prevalence of 22q11 microdeletion. J Med Genet. 1996;33:719.
Botto LD, May K, Fernhoff PM, Correa A, Coleman K, Rasmussen SA, et al. A population-based study of the 22q11.2 deletion: phenotype, incidence, and contribution to major birth defects in the population. Pediatrics. 2003;1121:101–7.
Murphy KC, Jones LA, Owen MJ. High rates of schizophrenia in adults with velo-cardio-facial syndrome. Arch Gen Psychiatry. 1999;56:940–5.
Fung WL, McEvilly R, Fong J, Silversides C, Chow E, Bassett A. Elevated prevalence of generalized anxiety disorder in adults with 22q11.2 deletion syndrome. Am J Psychiatry. 2010;167:998.
Bassett AS, Lowther C, Merico D, Costain G, Chow EWC, van Amelsvoort T, et al. Rare genome-wide copy number variation and expression of schizophrenia in 22q11.2 deletion syndrome. Am J Psychiatry. 2017;174:1054–63.
Gur RE, Bassett AS, McDonald-McGinn DM, Bearden CE, Chow E, Emanuel BS, et al. A neurogenetic model for the study of schizophrenia spectrum disorders: the International 22q11.2 Deletion Syndrome Brain Behavior Consortium. Mol Psychiatry. 2017;22:1664–72.
Cleynen I, Engchuan W, Hestand MS, Heung T, Holleman AM, Johnston HR, et al. Genetic contributors to risk of schizophrenia in the presence of a 22q11.2 deletion. Mol Psychiatry. 2020.
Sumitomo A, Horike K, Hirai K, Butcher N, Boot E, Sakurai T, et al. A mouse model of 22q11.2 deletions: molecular and behavioral signatures of Parkinson’s disease and schizophrenia. Sci Adv. 2018;4:eaar6637.
Davies RW, Fiksinski AM, Breetvelt EJ, Williams NM, Hooper SR, Monfeuga T, et al. Using common genetic variation to examine phenotypic expression and risk prediction in 22q11.2 deletion syndrome. Nat Med. 2020;26:1912–8.
Merico D, Zarrei M, Costain G, Ogura L, Alipanahi B, Gazzellone MJ, et al. Whole-genome sequencing suggests schizophrenia risk mechanisms in humans with 22q11.2 deletion syndrome. G3 (Bethesda). 2015;5:2453–61.
Michaelovsky E, Carmel M, Frisch A, Salmon-Divon M, Pasmanik-Chor M, Weizman A, et al. Risk gene-set and pathways in 22q11.2 deletion-related schizophrenia: a genealogical molecular approach. Transl Psychiatry. 2019;9:15.
Lin JR, Zhang Q, Cai Y, Morrow BE, Zhang ZD. Integrated rare variant-based risk gene prioritization in disease case-control sequencing studies. PLoS Genet. 2017;13:e1007142.
Rentzsch P, Witten D, Cooper GM, Shendure J, Kircher M. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 2019;47:D886–94.
Wang Q, Charych EI, Pulito VL, Lee JB, Graziane NM, Crozier RA, et al. The psychiatric disease risk factors DISC1 and TNIK interact to regulate synapse composition and function. Mol Psychiatry. 2011;16:1006–23.
Kirov G, Rujescu D, Ingason A, Collier DA, O’Donovan MC, Owen MJ. Neurexin 1 (NRXN1) deletions in schizophrenia. Schizophr Bull. 2009;35:851–4.
Bhat S, Dao DT, Terrillion CE, Arad M, Smith RJ, Soldatov NM, et al. CACNA1C (Cav1.2) in the pathophysiology of psychiatric disease. Prog Neurobiol. 2012;99:1–14.
Fernandes BS, Molendijk ML, Kohler CA, Soares JC, Leite CM, Machado-Vieira R, et al. Peripheral brain-derived neurotrophic factor (BDNF) as a biomarker in bipolar disorder: a meta-analysis of 52 studies. BMC Med. 2015;13:289.
Kishi T, Yoshimura R, Ikuta T, Iwata N. Brain-derived neurotrophic factor and major depressive disorder: evidence from meta-analyses. Front Psychiatry. 2017;8:308.
Shibata T, Yamagata H, Uchida S, Otsuki K, Hobara T, Higuchi F, et al. The alteration of hypoxia inducible factor-1 (HIF-1) and its target genes in mood disorder patients. Prog Neuropsychopharmacol Biol Psychiatry. 2013;43:222–9.
Brosch K, Stein F, Schmitt S, Pfarr JK, Ringwald KG, Thomas-Odenthal F, et al. Reduced hippocampal gray matter volume is a common feature of patients with major depression, bipolar disorder, and schizophrenia spectrum disorders. Mol Psychiatry. 2022;27:4234–43.
Bassett AS, Costain G, Fung WL, Russell KJ, Pierce L, Kapadia R, et al. Clinically detectable copy number variations in a Canadian catchment population of schizophrenia. J Psychiatr Res. 2010;44:1005–9.
Li M, Santpere G, Imamura Kawasawa Y, Evgrafov OV, Gulden FO, Pochareddy S, et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science. 2018;362:eaat7615.
Willsey AJ, Sanders SJ, Li M, Dong S, Tebbenkamp AT, Muhle RA, et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell. 2013;155:997–1007.
Forsyth JK, Nachun D, Gandal MJ, Geschwind DH, Anderson AE, Coppola G, et al. Synaptic and gene regulatory mechanisms in schizophrenia, autism, and 22q11.2 copy number variant-mediated risk for neuropsychiatric disorders. Biol Psychiatry. 2020;87:150–63.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Singh T, Poterba T, Curtis D, Akil H, Al Eissa M, Barchas JD, et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature. 2022;604:509–16.
Iyegbe CO, O’Reilly PF. Genetic origins of schizophrenia find common ground. Nature. 2022;604:433–5.
Lin GN, Corominas R, Lemmens I, Yang X, Tavernier J, Hill DE, et al. Spatiotemporal 16p11.2 protein network implicates cortical late mid-fetal brain development and KCTD13-Cul3-RhoA pathway in psychiatric diseases. Neuron. 2015;85:742–54.
Egan MF, Goldberg TE, Kolachana BS, Callicott JH, Mazzanti CM, Straub RE, et al. Effect of COMT Val108/158 Met genotype on frontal lobe function and risk for schizophrenia. Proc Natl Acad Sci USA. 2001;98:6917–22.
Williams HJ, Owen MJ, O’Donovan MC. Is COMT a susceptibility gene for schizophrenia? Schizophr Bull. 2007;33:635–41.
Gokhale A, Hartwig C, Freeman AAH, Bassell JL, Zlatic SA, Sapp Savas C, et al. Systems analysis of the 22q11.2 microdeletion syndrome converges on a mitochondrial interactome necessary for synapse function and behavior. J Neurosci. 2019;39:3561–81.
Li J, Ryan SK, Deboer E, Cook K, Fitzgerald S, Lachman HM, et al. Mitochondrial deficits in human iPSC-derived neurons from patients with 22q11.2 deletion syndrome and schizophrenia. Transl Psychiatry. 2019;9:302.
Genovese G, Fromer M, Stahl EA, Ruderfer DM, Chambert K, Landen M, et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat Neurosci. 2016;19:1433–41.
Flannick J, Mercader JM, Fuchsberger C, Udler MS, Mahajan A, Wessel J, et al. Exome sequencing of 20,791 cases of type 2 diabetes and 24,440 controls. Nature. 2019;570:71–6.
Euesden J, Lewis CM, O’Reilly PF. PRSice: polygenic risk score software. Bioinformatics. 2015;31:1466–8.
Choi SW, O’Reilly PF. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience. 2019;8:giz082.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011;27:1739–40.
Zhang X, Lan Y, Xu J, Quan F, Zhao E, Deng C, et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 2019;47:D721–8.
BrainSpan Atlas of the Developing Human Brain [Internet]. Available at http://brainspan.org. 2011.
Consortium F, the RP, Clst, Forrest AR, Kawaji H, Rehli M, et al. A promoter-level mammalian expression atlas. Nature. 2014;507:462–70.
Wang D, Liu S, Warrell J, Won H, Shi X, Navarro FCP, et al. Comprehensive functional genomic resource and integrative model for the human brain. Science. 2018;362:6420.
Hammal F, de Langen P, Bergon A, Lopez F, Ballester B. ReMap 2022: a database of Human, Mouse, Drosophila and Arabidopsis regulatory regions from an integrative analysis of DNA-binding sequencing experiments. Nucleic Acids Res. 2022;50:D316–25.
Meuleman W, Muratov A, Rynes E, Halow J, Lee K, Bates D, et al. Index and biological spectrum of human DNase I hypersensitive sites. Nature. 2020;584:244–51.
Lee S, Emond MJ, Bamshad MJ, Barnes KC, Rieder MJ, Nickerson DA, et al. Optimal unified approach for rare-variant association testing with application to small-sample case-control whole-exome sequencing studies. Am J Hum Genet. 2012;91:224–37.
Subramanian A, Narayan R, Corsello SM, Peck DD, Natoli TE, Lu X, et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell. 2017;171:1437–52.e1417.
Zinkstok JR, Boot E, Bassett AS, Hiroi N, Butcher NJ, Vingerhoets C, et al. Neurobiological perspective of 22q11.2 deletion syndrome. Lancet Psychiatry. 2019;6:951–60.
Hu Y, Vinayagam A, Nand A, Comjean A, Chung V, Hao T, et al. Molecular Interaction Search Tool (MIST): an integrated resource for mining gene and protein interaction data. Nucleic Acids Res. 2018;46:D567–74.
Acknowledgements
This work was supported by funding to ZDZ from NIH Grants U01 MH101720, R01 AG057909, U19 AG056278, R01 AG061521, R01 AG057706, RF1 AG057341, R01 AG061155, P01 AG047200, P01 AG017242, P30 AG038072, and a Career Scientist Award from the Irma T. Hirschl Trust.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, JR., Zhao, Y., Jabalameli, M.R. et al. Rare coding variants as risk modifiers of the 22q11.2 deletion implicate postnatal cortical development in syndromic schizophrenia. Mol Psychiatry 28, 2071–2080 (2023). https://doi.org/10.1038/s41380-023-02009-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-023-02009-y
This article is cited by
-
Psychosis and autism spectrum disorder: a special issue of Molecular Psychiatry
Molecular Psychiatry (2023)