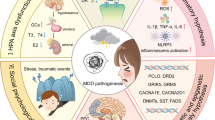
Abstract
Genome-wide association studies (GWAS) have reported substantial genomic loci significantly associated with clinical risk of bipolar disorder (BD), and studies combining techniques of genetics, neuroscience, neuroimaging, and pharmacology are believed to help tackle clinical problems (e.g., identifying novel therapeutic targets). However, translating findings of psychiatric genetics into biological mechanisms underlying BD pathogenesis remains less successful. Biological impacts of majority of BD GWAS risk loci are obscure, and the involvement of many GWAS risk genes in this illness is yet to be investigated. It is thus necessary to review the progress of applying BD GWAS risk genes in the research and intervention of the disorder. A comprehensive literature search found that a number of such risk genes had been investigated in cellular or animal models, even before they were highlighted in BD GWAS. Intriguingly, manipulation of many BD risk genes (e.g., ANK3, CACNA1C, CACNA1B, HOMER1, KCNB1, MCHR1, NCAN, SHANK2 etc.) resulted in altered murine behaviors largely restoring BD clinical manifestations, including mania-like symptoms such as hyperactivity, anxiolytic-like behavior, as well as antidepressant-like behavior, and these abnormalities could be attenuated by mood stabilizers. In addition to recapitulating phenotypic characteristics of BD, some GWAS risk genes further provided clues for the neurobiology of this illness, such as aberrant activation and functional connectivity of brain areas in the limbic system, and modulated dendritic spine morphogenesis as well as synaptic plasticity and transmission. Therefore, BD GWAS risk genes are undoubtedly pivotal resources for modeling this illness, and might be translational therapeutic targets in the future clinical management of BD. We discuss both promising prospects and cautions in utilizing the bulk of useful resources generated by GWAS studies. Systematic integrations of findings from genetic and neuroscience studies are called for to promote our understanding and intervention of BD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Carvalho AF, Firth J, Vieta E. Bipolar disorder. N. Engl J Med. 2020;383:58–66.
Glahn DC, Almasy L, Barguil M, Hare E, Peralta JM, Kent JW Jr, et al. Neurocognitive endophenotypes for bipolar disorder identified in multiplex multigenerational families. Arch Gen Psychiatry. 2010;67:168–77.
Plans L, Barrot C, Nieto E, Rios J, Schulze TG, Papiol S, et al. Association between completed suicide and bipolar disorder: A systematic review of the literature. J Affect Disord. 2019;242:111–22.
Kato T. Molecular neurobiology of bipolar disorder: a disease of ‘mood-stabilizing neurons’? Trends Neurosci. 2008;31:495–503.
Ashok AH, Marques TR, Jauhar S, Nour MM, Goodwin GM, Young AH, et al. The dopamine hypothesis of bipolar affective disorder: the state of the art and implications for treatment. Mol Psychiatry. 2017;22:666–79.
Haggarty SJ, Karmacharya R, Perlis RH. Advances toward precision medicine for bipolar disorder: mechanisms & molecules. Mol Psychiatry. 2021;26:168–85.
Bartoli F, Misiak BOD, Callovini T, Cavaleri D, Cioni RM, Crocamo C, et al. The kynurenine pathway in bipolar disorder: a meta-analysis on the peripheral blood levels of tryptophan and related metabolites. Mol Psychiatry. 2021;26:3419–29.
Harrison PJ, Geddes JR, Tunbridge EM. The emerging neurobiology of bipolar disorder. Trends Neurosci. 2018;41:18–30.
Wang ZW, Jun C, Gao KM, Yang HC, Fang YR. Perspective on etiology and treatment of bipolar disorders in China: clinical implications and future directions. Neurosci Bull. 2019;35:608–12.
Gordovez FJA, McMahon FJ. The genetics of bipolar disorder. Mol Psychiatry. 2020;25:544–59.
Zhang C, Xiao X, Li T, Li M. Translational genomics and beyond in bipolar disorder. Mol Psychiatry. 2021;26:186–202.
McGuffin P, Rijsdijk F, Andrew M, Sham P, Katz R, Cardno A. The heritability of bipolar affective disorder and the genetic relationship to unipolar depression. Arch Gen Psychiatry. 2003;60:497–502.
Kieseppa T, Partonen T, Haukka J, Kaprio J, Lonnqvist J. High concordance of bipolar I disorder in a nationwide sample of twins. Am J Psychiatry. 2004;161:1814–21.
Stahl EA, Breen G, Forstner AJ, McQuillin A, Ripke S, Trubetskoy V, et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet. 2019;51:793–803.
Li HJ, Zhang C, Hui L, Zhou DS, Li Y, Zhang CY, et al. Novel risk loci associated with genetic risk for bipolar disorder among Han Chinese individuals: A genome-wide association study and meta-analysis. JAMA Psychiatry. 2021;78:320–30.
Mullins N, Forstner AJ, O’Connell KS, Coombes B, Coleman JRI, Qiao Z, et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet. 2021;53:817–29.
Nunes A, Schnack HG, Ching CRK, Agartz I, Akudjedu TN, Alda M, et al. Using structural MRI to identify bipolar disorders - 13 site machine learning study in 3020 individuals from the ENIGMA Bipolar Disorders Working Group. Mol Psychiatry. 2020;25:2130–43.
Bertocci MA, Hanford L, Manelis A, Iyengar S, Youngstrom EA, Gill MK, et al. Clinical, cortical thickness and neural activity predictors of future affective lability in youth at risk for bipolar disorder: initial discovery and independent sample replication. Mol Psychiatry. 2019;24:1856–67.
Hibar DP, Westlye LT, van Erp TG, Rasmussen J, Leonardo CD, Faskowitz J, et al. Subcortical volumetric abnormalities in bipolar disorder. Mol Psychiatry. 2016;21:1710–6.
Hibar DP, Westlye LT, Doan NT, Jahanshad N, Cheung JW, Ching CRK, et al. Cortical abnormalities in bipolar disorder: an MRI analysis of 6503 individuals from the ENIGMA Bipolar Disorder Working Group. Mol Psychiatry. 2018;23:932–42.
Abe C, Liberg B, Song J, Bergen SE, Petrovic P, Ekman C-J, et al. Longitudinal cortical thickness changes in bipolar disorder and the relation to genetic risk, mania and lithium use. Biol Psychiatry. 2020;87:271–81.
Weinberger DR, Radulescu E. Finding the elusive psychiatric “Lesion” with 21st-century neuroanatomy: A note of caution. Am J Psychiatry. 2016;173:27–33.
Mufford MS, Stein DJ, Dalvie S, Groenewold NA, Thompson PM, Jahanshad N. Neuroimaging genomics in psychiatry-a translational approach. Genome Med. 2017;9:102.
Roberts G, Perry A, Lord A, Frankland A, Leung V, Holmes-Preston E, et al. Structural dysconnectivity of key cognitive and emotional hubs in young people at high genetic risk for bipolar disorder. Mol Psychiatry. 2018;23:413–21.
Pavuluri MN, O’Connor MM, Harral E, Sweeney JA. Affective neural circuitry during facial emotion processing in pediatric bipolar disorder. Biol Psychiatry. 2007;62:158–67.
Brotman MA, Tseng WL, Olsavsky AK, Fromm SJ, Muhrer EJ, Rutenberg JG, et al. Fronto-limbic-striatal dysfunction in pediatric and adult patients with bipolar disorder: impact of face emotion and attentional demands. Psychol Med. 2014;44:1639–51.
Almeida JR, Versace A, Hassel S, Kupfer DJ, Phillips ML. Elevated amygdala activity to sad facial expressions: a state marker of bipolar but not unipolar depression. Biol Psychiatry. 2010;67:414–21.
Wegbreit E, Cushman GK, Puzia ME, Weissman AB, Kim KL, Laird AR, et al. Developmental meta-analyses of the functional neural correlates of bipolar disorder. JAMA Psychiatry. 2014;71:926–35.
Manelis A, Ladouceur CD, Graur S, Monk K, Bonar LK, Hickey MB, et al. Altered amygdala-prefrontal response to facial emotion in offspring of parents with bipolar disorder. Brain. 2015;138:2777–90.
Olsavsky AK, Brotman MA, Rutenberg JG, Muhrer EJ, Deveney CM, Fromm SJ, et al. Amygdala hyperactivation during face emotion processing in unaffected youth at risk for bipolar disorder. J Am Acad Child Adolesc Psychiatry. 2012;51:294–303.
Acuff HE, Versace A, Bertocci MA, Ladouceur CD, Hanford LC, Manelis A, et al. Association of neuroimaging measures of emotion processing and regulation neural circuitries with symptoms of bipolar disorder in offspring at risk for bipolar disorder. JAMA Psychiatry. 2018;75:1241–51.
Meyer-Lindenberg A, Weinberger DR. Intermediate phenotypes and genetic mechanisms of psychiatric disorders. Nat Rev Neurosci. 2006;7:818–27.
Jogia J, Ruberto G, Lelli-Chiesa G, Vassos E, Maieru M, Tatarelli R, et al. The impact of the CACNA1C gene polymorphism on frontolimbic function in bipolar disorder. Mol Psychiatry. 2011;16:1070–1.
Dima D, Jogia J, Collier D, Vassos E, Burdick KE, Frangou S. Independent modulation of engagement and connectivity of the facial network during affect processing by CACNA1C and ANK3 risk genes for bipolar disorder. JAMA Psychiatry. 2013;70:1303–11.
Radua J, Surguladze SA, Marshall N, Walshe M, Bramon E, Collier DA, et al. The impact of CACNA1C allelic variation on effective connectivity during emotional processing in bipolar disorder. Mol Psychiatry. 2013;18:526–7.
Erk S, Meyer-Lindenberg A, Schmierer P, Mohnke S, Grimm O, Garbusow M, et al. Hippocampal and frontolimbic function as intermediate phenotype for psychosis: evidence from healthy relatives and a common risk variant in CACNA1C. Biol Psychiatry. 2014;76:466–75.
Cosgrove D, Mothersill O, Kendall K, Konte B, Harold D, Giegling I, et al. Cognitive characterization of schizophrenia risk variants involved in synaptic transmission: evidence of CACNA1C’s role in working memory. Neuropsychopharmacology. 2017;42:2612–22.
Janiri D, Kotzalidis GD, di Luzio M, Giuseppin G, Simonetti A, Janiri L, et al. Genetic neuroimaging of bipolar disorder: a systematic 2017-2020 update. Psychiatr Genet. 2021;31:50–64.
Lippard ETC, Jensen KP, Wang F, Johnston JAY, Spencer L, Pittman B, et al. Effects of ANK3 variation on gray and white matter in bipolar disorder. Mol Psychiatry. 2017;22:1345–51.
Linke J, Witt SH, King AV, Nieratschker V, Poupon C, Gass A, et al. Genome-wide supported risk variant for bipolar disorder alters anatomical connectivity in the human brain. Neuroimage. 2012;59:3288–96.
Cheng YQ, Xu J, Dong CL, Shen ZL, Zhou C, Li N et al. Age-related atrophy of cortical thickness and genetic effect of ANK3 gene in first episode MDD patients. Neuroimage-Clin. 2020;28:102384.
Cassidy C, Buchy L, Bodnar M, Dell’Elce J, Choudhry Z, Fathalli F, et al. Association of a risk allele of ANK3 with cognitive performance and cortical thickness in patients with first-episode psychosis. J Psychiatr Neurosci. 2014;39:31–9.
Benedetti F, Poletti S, Locatelli C, Mazza E, Lorenzi C, Vitali A, et al. A Homer 1 gene variant influences brain structure and function, lithium effects on white matter, and antidepressant response in bipolar disorder: A multimodal genetic imaging study. Prog Neuropsychopharmacol Biol Psychiatry. 2018;81:88–95.
Koch K, Stegmaier S, Schwarz L, Erb M, Thomas M, Scheffler K, et al. CACNA1C risk variant affects microstructural connectivity of the amygdala. Neuroimage Clin. 2019;22:101774.
Lancaster TM, Foley S, Tansey KE, Linden DE, Caseras X. CACNA1C risk variant is associated with increased amygdala volume. Eur Arch Psychiatry Clin Neurosci. 2016;266:269–75.
Wolf C, Mohr H, Schneider-Axmann T, Reif A, Wobrock T, Scherk H, et al. CACNA1C genotype explains interindividual differences in amygdala volume among patients with schizophrenia. Eur Arch Psychiatry Clin Neurosci. 2014;264:93–102.
Soeiro-de-Souza MG, Lafer B, Moreno RA, Nery FG, Chile T, Chaim K, et al. The CACNA1C risk allele rs1006737 is associated with age-related prefrontal cortical thinning in bipolar I disorder. Transl Psychiatry. 2017;7:e1086.
Chang H, Hoshina N, Zhang C, Ma Y, Cao H, Wang Y, et al. The protocadherin 17 gene affects cognition, personality, amygdala structure and function, synapse development and risk of major mood disorders. Mol Psychiatry. 2018;23:400–12.
Heinrich A, Lourdusamy A, Tzschoppe J, Vollstadt-Klein S, Buhler M, Steiner S, et al. The risk variant in ODZ4 for bipolar disorder impacts on amygdala activation during reward processing. Bipolar Disord. 2013;15:440–5.
Wessa M, Linke J, Witt SH, Nieratschker V, Esslinger C, Kirsch P, et al. The CACNA1C risk variant for bipolar disorder influences limbic activity. Mol Psychiatry. 2010;15:1126–7.
Bigos KL, Mattay VS, Callicott JH, Straub RE, Vakkalanka R, Kolachana B, et al. Genetic variation in CACNA1C affects brain circuitries related to mental illness. Arch Gen Psychiatry. 2010;67:939–45.
Erk S, Meyer-Lindenberg A, Schnell K, Opitz von Boberfeld C, Esslinger C, Kirsch P, et al. Brain function in carriers of a genome-wide supported bipolar disorder variant. Arch Gen Psychiatry. 2010;67:803–11.
Li M, Luo XJ, Rietschel M, Lewis CM, Mattheisen M, Muller-Myhsok B, et al. Allelic differences between Europeans and Chinese for CREB1 SNPs and their implications in gene expression regulation, hippocampal structure and function, and bipolar disorder susceptibility. Mol Psychiatry. 2014;19:452–61.
Bipolar Disorder Schizophrenia Working Group of the Psychiatric Genomics Consortium. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell. 2018;173:1705–15.
Ruderfer DM, Fanous AH, Ripke S, McQuillin A, Amdur RL. Schizophrenia Working Group of the Psychiatric Genomics C et al. Polygenic dissection of diagnosis and clinical dimensions of bipolar disorder and schizophrenia. Mol Psychiatry. 2014;19:1017–24.
Takeuchi H, Kimura R, Tomita H, Taki Y, Kikuchi Y, Ono C et al. Polygenic risk score for bipolar disorder associates with divergent thinking and brain structures in the prefrontal cortex. Hum Brain Mapp. 2021;42:6028–37.
Power RA, Steinberg S, Bjornsdottir G, Rietveld CA, Abdellaoui A, Nivard MM, et al. Polygenic risk scores for schizophrenia and bipolar disorder predict creativity. Nat Neurosci. 2015;18:953–5.
Le-Niculescu H, Roseberry K, Gill SS, Levey DF, Phalen PL, Mullen J, et al. Precision medicine for mood disorders: objective assessment, risk prediction, pharmacogenomics, and repurposed drugs. Mol Psychiatry. 2021;26:2776–804.
Dunayevich E, Keck PE Jr. Prevalence and description of psychotic features in bipolar mania. Curr Psychiatry Rep. 2000;2:286–90.
Allardyce J, Leonenko G, Hamshere M, Pardinas AF, Forty L, Knott S, et al. Association between schizophrenia-related polygenic liability and the occurrence and level of mood-incongruent psychotic symptoms in bipolar disorder. JAMA Psychiatry. 2018;75:28–35.
Cheng R, Juo SH, Loth JE, Nee J, Iossifov I, Blumenthal R, et al. Genome-wide linkage scan in a large bipolar disorder sample from the National Institute of Mental Health genetics initiative suggests putative loci for bipolar disorder, psychosis, suicide, and panic disorder. Mol Psychiatry. 2006;11:252–60.
Goes FS, Zandi PP, Miao K, McMahon FJ, Steele J, Willour VL, et al. Mood-incongruent psychotic features in bipolar disorder: familial aggregation and suggestive linkage to 2p11-q14 and 13q21-33. Am J Psychiatry. 2007;164:236–47.
Buoli M, Caldiroli A, Cumerlato Melter C, Serati M, de Nijs J, Altamura AC. Biological aspects and candidate biomarkers for psychotic bipolar disorder: A systematic review. Psychiatry Clin Neurosci. 2016;70:227–44.
Goes FS, Hamshere ML, Seifuddin F, Pirooznia M, Belmonte-Mahon P, Breuer R, et al. Genome-wide association of mood-incongruent psychotic bipolar disorder. Transl Psychiatry. 2012;2:e180.
Meier S, Mattheisen M, Vassos E, Strohmaier J, Treutlein J, Josef F, et al. Genome-wide significant association between a ‘negative mood delusions’ dimension in bipolar disorder and genetic variation on chromosome 3q26.1. Transl Psychiatry. 2012;2:e165.
Salvatore P, Baldessarini RJ, Khalsa HK, Tohen M. Prodromal features in first-psychotic episodes of major affective and schizoaffective disorders. J Affect Disord. 2021;295:1251–8.
Ambati A, Hillary R, Leu-Semenescu S, Ollila HM, Lin L, During EH et al. Kleine-Levin syndrome is associated with birth difficulties and genetic variants in the TRANK1 gene loci. Proc Natl Acad Sci. 2021;118:e2005753118.
Etain B, Milhiet V, Bellivier F, Leboyer M. Genetics of circadian rhythms and mood spectrum disorders. Eur Neuropsychopharmacol. 2011;21:S676–82.
McCarthy MJ, Nievergelt CM, Kelsoe JR, Welsh DK. A survey of genomic studies supports association of circadian clock genes with bipolar disorder spectrum illnesses and lithium response. PLoS One. 2012;7:e32091.
Soria V, Martinez-Amoros E, Escaramis G, Valero J, Perez-Egea R, Garcia C, et al. Differential association of circadian genes with mood disorders: CRY1 and NPAS2 are associated with unipolar major depression and CLOCK and VIP with bipolar disorder. Neuropsychopharmacology. 2010;35:1279–89.
Pagani L, St Clair PA, Teshiba TM, Service SK, Fears SC, Araya C, et al. Genetic contributions to circadian activity rhythm and sleep pattern phenotypes in pedigrees segregating for severe bipolar disorder. Proc Natl Acad Sci. 2016;113:E754–61.
Lewis KJS, Richards A, Karlsson R, Leonenko G, Jones SE, Jones HJ, et al. Comparison of genetic liability for sleep traits among individuals with bipolar disorder I or II and control participants. JAMA Psychiatry. 2020;77:303–10.
Harrison PJ, Colbourne L, Harrison CH. The neuropathology of bipolar disorder: systematic review and meta-analysis. Mol Psychiatry 2020;25:1787–808.
Cosgrove VE, Kelsoe JR, Suppes T. Toward a valid animal model of bipolar disorder: how the Research Domain Criteria help bridge the clinical-basic science divide. Biol Psychiatry. 2016;79:62–70.
van Enkhuizen J, Geyer MA, Minassian A, Perry W, Henry BL, Young JW. Investigating the underlying mechanisms of aberrant behaviors in bipolar disorder from patients to models: Rodent and human studies. Neurosci Biobehav Rev. 2015;58:4–18.
Cichon S, Muhleisen TW, Degenhardt FA, Mattheisen M, Miro X, Strohmaier J, et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am J Hum Genet. 2011;88:372–81.
Miro X, Meier S, Dreisow ML, Frank J, Strohmaier J, Breuer R, et al. Studies in humans and mice implicate neurocan in the etiology of mania. Am J Psychiatry. 2012;169:982–90.
Pappas AL, Bey AL, Wang X, Rossi M, Kim YH, Yan H, et al. Deficiency of Shank2 causes mania-like behavior that responds to mood stabilizers. JCI Insight. 2017;2:e92052.
Schmeisser MJ, Ey E, Wegener S, Bockmann J, Stempel AV, Kuebler A, et al. Autistic-like behaviours and hyperactivity in mice lacking ProSAP1/Shank2. Nature. 2012;486:256–60.
Won H, Lee HR, Gee HY, Mah W, Kim JI, Lee J, et al. Autistic-like social behaviour in Shank2-mutant mice improved by restoring NMDA receptor function. Nature. 2012;486:261–5.
Kim R, Kim J, Chung C, Ha S, Lee S, Lee E, et al. Cell-type-specific Shank2 deletion in mice leads to differential synaptic and behavioral phenotypes. J Neurosci. 2018;38:4076–92.
Modi ME, Brooks JM, Guilmette ER, Beyna M, Graf R, Reim D, et al. Hyperactivity and hypermotivation associated with increased striatal mGluR1 signaling in a Shank2 rat model of autism. Front Mol Neurosci. 2018;11:107.
Jaubert PJ, Golub MS, Lo YY, Germann SL, Dehoff MH, Worley PF, et al. Complex, multimodal behavioral profile of the Homer1 knockout mouse. Genes Brain Behav. 2007;6:141–54.
Szumlinski KK, Lominac KD, Kleschen MJ, Oleson EB, Dehoff MH, Schwartz MK, et al. Behavioral and neurochemical phenotyping of Homer1 mutant mice: possible relevance to schizophrenia. Genes Brain Behav. 2005;4:273–88.
Wagner KV, Hartmann J, Labermaier C, Hausl AS, Zhao G, Harbich D, et al. Homer1/mGluR5 activity moderates vulnerability to chronic social stress. Neuropsychopharmacology. 2015;40:1222–33.
Yu Z, Lin D, Zhong Y, Luo B, Liu S, Fei E, et al. Transmembrane protein 108 involves in adult neurogenesis in the hippocampal dentate gyrus. Cell Biosci. 2019;9:9.
Jiao HF, Sun XD, Bates R, Xiong L, Zhang L, Liu F, et al. Transmembrane protein 108 is required for glutamatergic transmission in dentate gyrus. Proc Natl Acad Sci. 2017;114:1177–82.
Huang Y, Huang J, Zhou QX, Yang CX, Yang CP, Mei WY, et al. ZFP804A mutant mice display sex-dependent schizophrenia-like behaviors. Mol Psychiatry. 2021;26:2514–32.
Hawkins NA, Misra SN, Jurado M, Kang SK, Vierra NC, Nguyen K, et al. Epilepsy and neurobehavioral abnormalities in mice with a dominant-negative KCNB1 pathogenic variant. Neurobiol Dis. 2021;147:105141.
Speca DJ, Ogata G, Mandikian D, Bishop HI, Wiler SW, Eum K, et al. Deletion of the Kv2.1 delayed rectifier potassium channel leads to neuronal and behavioral hyperexcitability. Genes Brain Behav. 2014;13:394–408.
Leussis MP, Berry-Scott EM, Saito M, Jhuang H, de Haan G, Alkan O, et al. The ANK3 bipolar disorder gene regulates psychiatric-related behaviors that are modulated by lithium and stress. Biol Psychiatry. 2013;73:683–90.
Zhu S, Cordner ZA, Xiong J, Chiu CT, Artola A, Zuo Y, et al. Genetic disruption of ankyrin-G in adult mouse forebrain causes cortical synapse alteration and behavior reminiscent of bipolar disorder. Proc Natl Acad Sci. 2017;114:10479–84.
Shen W, Wang QW, Liu YN, Marchetto MC, Linker S, Lu SY, et al. Synaptotagmin-7 is a key factor for bipolar-like behavioral abnormalities in mice. Proc Natl Acad Sci. 2020;117:4392–9.
Wang QW, Lu SY, Liu YN, Chen Y, Wei H, Shen W, et al. Synaptotagmin-7 deficiency induces mania-like behavioral abnormalities through attenuating GluN2B activity. Proc Natl Acad Sci. 2020;117:31438–47.
Wang QW, Wang YH, Wang B, Chen Y, Lu SY, Yao J. Synaptotagmin-7-mediated activation of spontaneous NMDAR currents is disrupted in bipolar disorder susceptibility variants. PLoS Biol. 2021;19:e3001323.
Lex C, Bazner E, Meyer TD. Does stress play a significant role in bipolar disorder? A meta-analysis. J Affect Disord. 2017;208:298–308.
O’Tuathaigh CMP, Fumagalli F, Desbonnet L, Perez-Branguli F, Moloney G, Loftus S, et al. Epistatic and independent effects on schizophrenia-related phenotypes following co-disruption of the risk factors Neuregulin-1 x DISC1. Schizophr Bull. 2017;43:214–25.
Zheng Y, Shen W, Zhang J, Yang B, Liu YN, Qi HH, et al. CRISPR interference-based specific and efficient gene inactivation in the brain. Nat Neurosci. 2018;21:447–54.
Lee Y, Zhang Y, Kim S, Han K. Excitatory and inhibitory synaptic dysfunction in mania: an emerging hypothesis from animal model studies. Exp Mol Med. 2018;50:12.
Schloesser RJ, Martinowich K, Manji HK. Mood-stabilizing drugs: mechanisms of action. Trends Neurosci. 2012;35:36–46.
Kim HJ, Thayer SA. Lithium increases synapse formation between hippocampal neurons by depleting phosphoinositides. Mol Pharm. 2009;75:1021–30.
Logan RW, Ozburn AR, Arey RN, Ketchesin KD, Winquist A, Crain A, et al. Valproate reverses mania-like behaviors in mice via preferential targeting of HDAC2. Mol Psychiatry. 2021;26:4066–84.
Tobe BTD, Crain AM, Winquist AM, Calabrese B, Makihara H, Zhao WN, et al. Probing the lithium-response pathway in hiPSCs implicates the phosphoregulatory set-point for a cytoskeletal modulator in bipolar pathogenesis. Proc Natl Acad Sci. 2017;114:E4462–E71.
Nasrallah HA. Neurodevelopmental aspects of bipolar affective disorder. Biol Psychiatry. 1991;29:1–2.
O’Shea KS, McInnis MG. Neurodevelopmental origins of bipolar disorder: iPSC models. Mol Cell Neurosci. 2016;73:63–83.
Konopaske GT, Lange N, Coyle JT, Benes FM. Prefrontal cortical dendritic spine pathology in schizophrenia and bipolar disorder. JAMA Psychiatry. 2014;71:1323–31.
Penzes P, Cahill ME, Jones KA, VanLeeuwen JE, Woolfrey KM. Dendritic spine pathology in neuropsychiatric disorders. Nat Neurosci. 2011;14:285–93.
Forrest MP, Parnell E, Penzes P. Dendritic structural plasticity and neuropsychiatric disease. Nat Rev Neurosci. 2018;19:215–34.
Penzes P, Jones KA. Dendritic spine dynamics-a key role for kalirin-7. Trends Neurosci. 2008;31:419–27.
Yoon S, Piguel NH, Khalatyan N, Dionisio LE, Savas JN, Penzes P. Homer1 promotes dendritic spine growth through ankyrin-G and its loss reshapes the synaptic proteome. Mol Psychiatry. 2021;26:1775–89.
McGuier NS, Padula AE, Mulholland PJ, Chandler LJ. Homer2 deletion alters dendritic spine morphology but not alcohol-associated adaptations in GluN2B-containing N-methyl-D-aspartate receptors in the nucleus accumbens. Front Pharm. 2015;6:28.
Spratt PWE, Ben-Shalom R, Keeshen CM, Burke KJ Jr, Clarkson RL, Sanders SJ, et al. The autism-associated gene Scn2a contributes to dendritic excitability and synaptic function in the prefrontal cortex. Neuron. 2019;103:673–85 e5.
Shin W, Kweon H, Kang R, Kim D, Kim K, Kang M, et al. Scn2a haploinsufficiency in mice suppresses hippocampal neuronal excitability, excitatory synaptic drive, and long-term potentiation, and spatial learning and memory. Front Mol Neurosci. 2019;12:145.
Teng LL, Lu GL, Chiou LC, Lin WS, Cheng YY, Hsueh TE, et al. Serotonin receptor HTR6-mediated mTORC1 signaling regulates dietary restriction-induced memory enhancement. PLoS Biol. 2019;17:e2007097.
Chen C, Meng Q, Xia Y, Ding C, Wang L, Dai R et al. The transcription factor POU3F2 regulates a gene coexpression network in brain tissue from patients with psychiatric disorders. Sci Transl Med. 2018;10:eaat8178.
Meng Q, Wang L, Dai R, Wang J, Ren Z, Liu S, et al. Integrative analyses prioritize GNL3 as a risk gene for bipolar disorder. Mol Psychiatry. 2020;25:2672–84.
Yu H, Yan H, Li J, Li Z, Zhang X, Ma Y, et al. Common variants on 2p16.1, 6p22.1 and 10q24.32 are associated with schizophrenia in Han Chinese population. Mol Psychiatry. 2017;22:954–60.
Ding C, Zhang C, Kopp R, Kuney L, Meng Q, Wang L, et al. Transcription factor POU3F2 regulates TRIM8 expression contributing to cellular functions implicated in schizophrenia. Mol Psychiatry. 2021;26:3444–60.
Durak O, de Anda FC, Singh KK, Leussis MP, Petryshen TL, Sklar P, et al. Ankyrin-G regulates neurogenesis and Wnt signaling by altering the subcellular localization of beta-catenin. Mol Psychiatry. 2015;20:388–97.
Yang Z, Zhou D, Li H, Cai X, Liu W, Wang L, et al. The genome-wide risk alleles for psychiatric disorders at 3p21.1 show convergent effects on mRNA expression, cognitive function and mushroom dendritic spine. Mol Psychiatry. 2020;25:48–66.
Deans PJM, Raval P, Sellers KJ, Gatford NJF, Halai S, Duarte RRR, et al. Psychosis risk candidate ZNF804A localizes to synapses and regulates neurite formation and dendritic spine structure. Biol Psychiatry. 2017;82:49–61.
Lee J, Lee S, Ryu YJ, Lee D, Kim S, Seo JY, et al. Vaccinia-related kinase 2 plays a critical role in microglia-mediated synapse elimination during neurodevelopment. Glia. 2019;67:1667–79.
Yoon S, Parnell E, Kasherman M, Forrest MP, Myczek K, Premarathne S, et al. Usp9X controls Ankyrin-repeat domain protein homeostasis during dendritic spine development. Neuron. 2020;105:506–21.
Smith KR, Penzes P. Ankyrins: Roles in synaptic biology and pathology. Mol Cell Neurosci. 2018;91:131–9.
Smith KR, Kopeikina KJ, Fawcett-Patel JM, Leaderbrand K, Gao R, Schurmann B, et al. Psychiatric risk factor ANK3/ankyrin-G nanodomains regulate the structure and function of glutamatergic synapses. Neuron. 2014;84:399–415.
Piguel NH, Yoon S, DeSimone FI, Sanders SS, Gao R, Horan KE et al. The 190 kDa Ankyrin-G isoform is required for the dendritic stability of neurons and its palmitoylation is altered by lithium. bioRxiv. 2019; https://doi.org/10.1101/620708.
Nelson AD, Caballero-Floran RN, Rodriguez Diaz JC, Hull JM, Yuan Y, Li J, et al. Ankyrin-G regulates forebrain connectivity and network synchronization via interaction with GABARAP. Mol Psychiatry. 2020;25:2800–17.
Yoon S, Parnell E, Penzes P. TGF-beta-induced phosphorylation of Usp9X stabilizes ankyrin-G and regulates dendritic spine development and maintenance. Cell Rep. 2020;31:107685.
Yoon S, Myczek K, Penzes P. cAMP signaling-mediated phosphorylation of diacylglycerol lipase alpha regulates interaction with Ankyrin-G and dendritic spine morphology. Biol Psychiatry. 2021;90:263–74.
Nanou E, Catterall WA. Calcium channels, synaptic plasticity, and neuropsychiatric disease. Neuron. 2018;98:466–81.
Mack AA, Gao YL, Ratajczak MZ, Kakar S, El-Mallakh RS. Review of animal models of bipolar disorder that alter ion regulation. Neurosci Biobehav Rev. 2019;107:208–14.
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, et al. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2018;46:D1074–D82.
Harrison PJ, Hall N, Mould A, Al-Juffali N, Tunbridge EM. Cellular calcium in bipolar disorder: systematic review and meta-analysis. Mol Psychiatry. 2019;26:4106–16.
Warsh JJ, Andreopoulos S, Li PP. Role of intracellular calcium signaling in the pathophysiology and pharmacotherapy of bipolar disorder: current status. Clin Neurosci Res. 2004;4:201–13.
Schlecker C, Boehmerle W, Jeromin A, DeGray B, Varshney A, Sharma Y, et al. Neuronal calcium sensor-1 enhancement of InsP3 receptor activity is inhibited by therapeutic levels of lithium. J Clin Invest. 2006;116:1668–74.
Franks RD, Dubovsky SL, Lifshitz M, Coen P, Subryan V, Walker SH. Long-term lithium-carbonate therapy causes hyperparathyroidism. Arch Gen Psychiatry. 1982;39:1074–7.
Cipriani A, Saunders K, Attenburrow MJ, Stefaniak J, Panchal P, Stockton S, et al. A systematic review of calcium channel antagonists in bipolar disorder and some considerations for their future development. Mol Psychiatry. 2016;21:1324–32.
Harrison PJ, Tunbridge EM, Dolphin AC, Hall J. Voltage-gated calcium channel blockers for psychiatric disorders: genomic reappraisal. Br J Psychiatry. 2020;216:250–3.
Clark MB, Wrzesinski T, Garcia-Bea AB, Hall NAL, Kleinman JE, Hyde T, et al. Long-read sequencing reveals the complex splicing profile of the psychiatric risk gene CACNA1C in human brain. Mol Psychiatry. 2020;25:37–47.
Pachoud B, Adamantidis A, Ravassard P, Luppi PH, Grisar T, Lakaye B, et al. Major impairments of glutamatergic transmission and long-term synaptic plasticity in the hippocampus of mice lacking the melanin-concentrating hormone receptor-1. J Neurophysiol. 2010;104:1417–25.
Ye H, Cui XY, Ding H, Cui SY, Hu X, Liu YT, et al. Melanin-concentrating hormone (MCH) and MCH-R1 in the locus coeruleus may be involved in the regulation of depressive-like behavior. Int J Neuropsychopharmacol. 2018;21:1128–37.
Garcia-Fuster MJ, Parks GS, Clinton SM, Watson SJ, Akil H, Civelli O. The melanin-concentrating hormone (MCH) system in an animal model of depression-like behavior. Eur Neuropsychopharmacol. 2012;22:607–13.
Al-Massadi O, Dieguez C, Schneeberger M, Lopez M, Schwaninger M, Prevot V et al. Multifaceted actions of melanin-concentrating hormone on mammalian energy homeostasis. Nat Rev Endocrinol. 2021;17:745–55.
Smith DG, Qi H, Svenningsson P, Wade M, Davis RJ, Gehlert DR, et al. Behavioral and biochemical responses to d-amphetamine in MCH1 receptor knockout mice. Synapse. 2008;62:128–36.
Roy M, David N, Cueva M, Giorgetti M. A study of the involvement of melanin-concentrating hormone receptor 1 (MCHR1) in murine models of depression. Biol Psychiatry. 2007;61:174–80.
Chee MJ, Hebert AJ, Briancon N, Flaherty SE 3rd, Pissios P, Maratos-Flier E. Conditional deletion of melanin-concentrating hormone receptor 1 from GABAergic neurons increases locomotor activity. Mol Metab. 2019;29:114–23.
Marsh DJ, Weingarth DT, Novi DE, Chen HY, Trumbauer ME, Chen AS, et al. Melanin-concentrating hormone 1 receptor-deficient mice are lean, hyperactive, and hyperphagic and have altered metabolism. Proc Natl Acad Sci. 2002;99:3240–5.
Millan MJ, Gobert A, Panayi F, Rivet JM, Dekeyne A, Brocco M, et al. The melanin-concentrating hormone1 receptor antagonists, SNAP-7941 and GW3430, enhance social recognition and dialysate levels of acetylcholine in the frontal cortex of rats. Int J Neuropsychopharmacol. 2008;11:1105–22.
Shimazaki T, Yoshimizu T, Chaki S. Melanin-concentrating hormone MCH1 receptor antagonists: a potential new approach to the treatment of depression and anxiety disorders. CNS Drugs. 2006;20:801–11.
Chaki S, Yamaguchi J, Yamada H, Thomsen W, Tran TA, Semple G, et al. ATC0175: an orally active melanin-concentrating hormone receptor 1 antagonist for the potential treatment of depression and anxiety. CNS Drug Rev. 2005;11:341–52.
Gehlert DR, Rasmussen K, Shaw J, Li X, Ardayfio P, Craft L, et al. Preclinical evaluation of melanin-concentrating hormone receptor 1 antagonism for the treatment of obesity and depression. J Pharm Exp Ther. 2009;329:429–38.
Borowsky B, Durkin MM, Ogozalek K, Marzabadi MR, DeLeon J, Lagu B, et al. Antidepressant, anxiolytic and anorectic effects of a melanin-concentrating hormone-1 receptor antagonist. Nat Med. 2002;8:825–30.
Kishi T, Ikuta T, Matsuda Y, Sakuma K, Okuya M, Nomura I et al. Pharmacological treatment for bipolar mania: a systematic review and network meta-analysis of double-blind randomized controlled trials. Mol Psychiatry. 2021; https://doi.org/10.1038/s41380-021-01334-4.
Pacchiarotti I, Anmella G, Colomer L, Vieta E. How to treat mania. Acta Psychiatr Scand. 2020;142:173–92.
Cipriani A, Barbui C, Salanti G, Rendell J, Brown R, Stockton S, et al. Comparative efficacy and acceptability of antimanic drugs in acute mania: a multiple-treatments meta-analysis. Lancet. 2011;378:1306–15.
Lee KM, Hawi ZH, Parkington HC, Parish CL, Kumar PV, Polo JM, et al. The application of human pluripotent stem cells to model the neuronal and glial components of neurodevelopmental disorders. Mol Psychiatry. 2020;25:368–78.
Hoffman GE, Schrode N, Flaherty E, Brennand KJ. New considerations for hiPSC-based models of neuropsychiatric disorders. Mol Psychiatry. 2019;24:49–66.
Brennand KJ, Simone A, Tran N, Gage FH. Modeling psychiatric disorders at the cellular and network levels. Mol Psychiatry. 2012;17:1239–53.
Falk A, Heine VM, Harwood AJ, Sullivan PF, Peitz M, Brustle O, et al. Modeling psychiatric disorders: from genomic findings to cellular phenotypes. Mol Psychiatry. 2016;21:1167–79.
Mishra HK, Ying NM, Luis A, Wei H, Nguyen M, Nakhla T, et al. Circadian rhythms in bipolar disorder patient-derived neurons predict lithium response: preliminary studies. Mol Psychiatry. 2021;26:3383–94.
Stern S, Santos R, Marchetto MC, Mendes APD, Rouleau GA, Biesmans S, et al. Neurons derived from patients with bipolar disorder divide into intrinsically different sub-populations of neurons, predicting the patients’ responsiveness to lithium. Mol Psychiatry. 2018;23:1453–65.
Santos R, Linker SB, Stern S, Mendes APD, Shokhirev MN, Erikson G, et al. Deficient LEF1 expression is associated with lithium resistance and hyperexcitability in neurons derived from bipolar disorder patients. Mol Psychiatry. 2021;26:2440–56.
Ripke S, Walters JT, O’Donovan MC, Schizophrenia Working Group of the Psychiatric Genomics Consortium. Mapping genomic loci prioritises genes and implicates synaptic biology in schizophrenia. medRxiv. 2020; https://doi.org/10.1101/2020.09.12.20192922.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Li W, Cai X, Li HJ, Song M, Zhang CY, Yang Y, et al. Independent replications and integrative analyses confirm TRANK1 as a susceptibility gene for bipolar disorder. Neuropsychopharmacology. 2021;46:1103–12.
Tesli M, Skatun KC, Ousdal OT, Brown AA, Thoresen C, Agartz I, et al. CACNA1C risk variant and amygdala activity in bipolar disorder, schizophrenia and healthy controls. PLoS One. 2013;8:e56970.
Liu F, Gong XH, Yao XD, Cui LL, Yin ZY, Li C, et al. Variation in the CACNB2 gene is associated with functional connectivity of the hippocampus in bipolar disorder. BMC Psychiatry. 2019;19:62.
Chen JS, Tan JW, Greenshaw AJ, Sawalha J, Liu Y, Zhang XF, et al. CACNB2 rs11013860 polymorphism correlates of prefrontal cortex thickness in bipolar patients with first-episode mania. J Affect Disord. 2020;268:82–7.
Rietschel M, Mattheisen M, Frank J, Treutlein J, Degenhardt F, Breuer R, et al. Genome-wide association-, replication-, and neuroimaging study implicates HOMER1 in the etiology of major depression. Biol Psychiatry. 2010;68:578–85.
Schultz CC, Muhleisen TW, Nenadic I, Koch K, Wagner G, Schachtzabel C, et al. Common variation in NCAN, a risk factor for bipolar disorder and schizophrenia, influences local cortical folding in schizophrenia. Psychol Med. 2014;44:811–20.
Raum H, Dietsche B, Nagels A, Witt SH, Rietschel M, Kircher T, et al. A genome-wide supported psychiatric risk variant in NCAN influences brain function and cognitive performance in healthy subjects. Hum Brain Mapp. 2015;36:378–90.
Dannlowski U, Kugel H, Grotegerd D, Redlich R, Suchy J, Opel N, et al. NCAN cross-disorder risk variant is associated with limbic gray matter deficits in healthy subjects and major depression. Neuropsychopharmacol. 2015;40:2510–6.
Assmann A, Richter A, Schutze H, Soch J, Barman A, Behnisch G, et al. Neurocan genome-wide psychiatric risk variant affects explicit memory performance and hippocampal function in healthy humans. Eur J Neurosci. 2021;53:3942–59.
Li M, Wang Y, Zheng XB, Ikeda M, Iwata N, Luo XJ, et al. Meta-analysis and brain imaging data support the involvement of VRK2 (rs2312147) in schizophrenia susceptibility. Schizophr Res. 2012;142:200–5.
Sohn H, Kim B, Kim KH, Kim MK, Choi TK, Lee SH. Effects of VRK2 (rs2312147) on white matter connectivity in patients with schizophrenia. Plos One. 2014;9:e103519.
Rasetti R, Sambataro F, Chen Q, Callicott JH, Mattay VS, Weinberger DR. Altered cortical network dynamics: a potential intermediate phenotype for schizophrenia and association with ZNF804A. Arch Gen Psychiatry. 2011;68:1207–17.
Esslinger C, Walter H, Kirsch P, Erk S, Schnell K, Arnold C, et al. Neural mechanisms of a genome-wide supported psychosis variant. Science. 2009;324:605.
Walter H, Schnell K, Erk S, Arnold C, Kirsch P, Esslinger C, et al. Effects of a genome-wide supported psychosis risk variant on neural activation during a theory-of-mind task. Mol Psychiatry. 2011;16:462–70.
Nakagawasai O, Onogi H, Mitazaki S, Sato A, Watanabe K, Saito H, et al. Behavioral and neurochemical characterization of mice deficient in the N-type Ca2+ channel alpha1B subunit. Behav Brain Res. 2010;208:224–30.
Beuckmann CT, Sinton CM, Miyamoto N, Ino M, Yanagisawa M. N-type calcium channel alpha1B subunit (Cav2.2) knock-out mice display hyperactivity and vigilance state differences. J Neurosci. 2003;23:6793–7.
Kabir ZD, Che A, Fischer DK, Rice RC, Rizzo BK, Byrne M, et al. Rescue of impaired sociability and anxiety-like behavior in adult cacna1c-deficient mice by pharmacologically targeting eIF2alpha. Mol Psychiatry. 2017;22:1096–109.
Dedic N, Pohlmann ML, Richter JS, Mehta D, Czamara D, Metzger MW, et al. Cross-disorder risk gene CACNA1C differentially modulates susceptibility to psychiatric disorders during development and adulthood. Mol Psychiatry. 2018;23:533–43.
Lee AS, Ra S, Rajadhyaksha AM, Britt JK, De Jesus-Cortes H, Gonzales KL, et al. Forebrain elimination of cacna1c mediates anxiety-like behavior in mice. Mol Psychiatry. 2012;17:1054–5.
Dao DT, Mahon PB, Cai X, Kovacsics CE, Blackwell RA, Arad M, et al. Mood disorder susceptibility gene CACNA1C modifies mood-related behaviors in mice and interacts with sex to influence behavior in mice and diagnosis in humans. Biol Psychiatry. 2010;68:801–10.
Renthal W, Maze I, Krishnan V, Covington HE 3rd, Xiao G, Kumar A, et al. Histone deacetylase 5 epigenetically controls behavioral adaptations to chronic emotional stimuli. Neuron. 2007;56:517–29.
Tsankova NM, Berton O, Renthal W, Kumar A, Neve RL, Nestler EJ. Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat Neurosci. 2006;9:519–25.
Middleton SJ, Kneller EM, Chen S, Ogiwara I, Montal M, Yamakawa K, et al. Altered hippocampal replay is associated with memory impairment in mice heterozygous for the Scn2a gene. Nat Neurosci. 2018;21:996–1003.
Yi X, Li M, He G, Du H, Li X, Cao D, et al. Genetic and functional analysis reveals TENM4 contributes to schizophrenia. iScience. 2021;24:103063.
Chai Z, Wang C, Huang R, Wang Y, Zhang X, Wu Q, et al. CaV2.2 gates calcium-independent but voltage-dependent secretion in mammalian sensory neurons. Neuron. 2017;96:1317–26 e4.
Bunda A, LaCarubba B, Bertolino M, Akiki M, Bath K, Lopez-Soto J, et al. Cacna1b alternative splicing impacts excitatory neurotransmission and is linked to behavioral responses to aversive stimuli. Mol Brain. 2019;12:81.
Moosmang S, Haider N, Klugbauer N, Adelsberger H, Langwieser N, Muller J, et al. Role of hippocampal Cav1.2 Ca2+ channels in NMDA receptor-independent synaptic plasticity and spatial memory. J Neurosci. 2005;25:9883–92.
Sala C, Futai K, Yamamoto K, Worley PF, Hayashi Y, Sheng M. Inhibition of dendritic spine morphogenesis and synaptic transmission by activity-inducible protein homer1a. J Neurosci. 2003;23:6327–37.
Kammermeier PJ, Worley PF. Homer 1a uncouples metabotropic glutamate receptor 5 from postsynaptic effectors. P Natl Acad Sci. 2007;104:6055–60.
Smothers CT, Szumlinski KK, Worley PF, Woodward JJ. Altered NMDA receptor function in primary cultures of hippocampal neurons from mice lacking the Homer2 gene. Synapse. 2016;70:33–9.
Vierra NC, Kirmiz M, van der List D, Santana LF, Trimmer JS. Kv2.1 mediates spatial and functional coupling of L-type calcium channels and ryanodine receptors in mammalian neurons. Elife. 2019;8:e49953.
Kobayashi Y, Okada T, Miki D, Sekino Y, Koganezawa N, Shirao T, et al. Properties of primary cilia in melanin-concentrating hormone receptor 1-bearing hippocampal neurons in vivo and in vitro. Neurochem Int. 2021;142:104902.
Eltokhi A, Gonzalez-Lozano MA, Oettl LL, Rozov A, Pitzer C, Roth R et al. Imbalanced post- and extrasynaptic SHANK2A functions during development affect social behavior in SHANK2-mediated neuropsychiatric disorders. Mol Psychiatry. 2021;26:6482–504.
Suzuki N, Numakawa T, Chou J, de Vega S, Mizuniwa C, Sekimoto K, et al. Teneurin-4 promotes cellular protrusion formation and neurite outgrowth through focal adhesion kinase signaling. FASEB J. 2014;28:1386–97.
Suzuki N, Fukushi M, Kosaki K, Doyle AD, de Vega S, Yoshizaki K, et al. Teneurin-4 is a novel regulator of oligodendrocyte differentiation and myelination of small-diameter axons in the CNS. J Neurosci. 2012;32:11586–99.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81971259 to ML, 82071275 to XX, 31872778 and 82171506 to ZH, 81930033 to YF); Yunnan Fundamental Research Project (202101AW070020 to XX); Nature Science Foundation of China Key Project (81630030 and 81920108018 to TL); Special Foundation for Brain Research from Science and Technology Program of Guangdong (2018B030334001 to TL and YF/JC); the Clinical Research Center of Shanghai Mental Health Center Key Project (CRC2018ZD02 to YF); Project for Hangzhou Medical Disciplines of Excellence and Key Project for Hangzhou Medical Disciplines (202004A11 to TL); National Key Research and Development Program of China (2016YFC1307100 to YF); the Western Light Innovative Research Team of Chinses Academy of Sciences. XX was also supported by the Chinese Academy of Sciences Western Light Program. ML was also supported by CAS Pioneer Hundred Talents Program and the 1000 Young Talents Program. ZH was also supported by Xiangya Hospital Start-up Research Grants, Key Research and Development Program from Hunan Province 2021DK2001 and the innovative team program 2019RS1010 from Department of Science & Technology of Hunan Province, the innovation-driven team project 2020CX016 from Central South University, 111 Grant (B13036), and Hunan 100 Talents Program. We would like to apologize to those authors whose work were not cited or elaborately represented in this review due to space constraints.
Author information
Authors and Affiliations
Contributions
ML oversaw the project. ML, YF, TL, and ZH conceived and designed the study, drafted the first version of the manuscript. XX and JC contributed to discussion and preparation of the manuscript. All authors revised the manuscript critically and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, M., Li, T., Xiao, X. et al. Phenotypes, mechanisms and therapeutics: insights from bipolar disorder GWAS findings. Mol Psychiatry 27, 2927–2939 (2022). https://doi.org/10.1038/s41380-022-01523-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-022-01523-9
This article is cited by
-
Identification of a psychiatric risk gene NISCH at 3p21.1 GWAS locus mediating dendritic spine morphogenesis and cognitive function
BMC Medicine (2023)
-
PAI1 inhibits the pathogenesis of primary focal hyperhidrosis by targeting CHRNA1
Orphanet Journal of Rare Diseases (2023)
-
Genetic evidence for the “dopamine hypothesis of bipolar disorder”
Molecular Psychiatry (2023)
-
Bidirectional genetic overlap between bipolar disorder and intelligence
BMC Medicine (2022)