Abstract
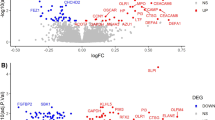
Epigenetic modifications are plausible molecular sources of phenotypic heterogeneity across schizophrenia patients. The current study investigated biological heterogeneity in schizophrenia using peripheral epigenetic profiles to delineate illness subtypes independent of their phenomenological manifestations. We applied epigenome-wide profiling with a DNA methylation array from blood samples of 63 schizophrenia patients and 59 healthy controls. Non-negative matrix factorization (NMF) and k-means clustering were performed to identify DNA methylation-related patient subtypes. The validity of the partition was tested by assessing the profile of the T cell receptor (TCR) repertoires. The uniqueness of the identified subtypes in relation to brain structural and clinical measures were evaluated. Two distinct patterns of DNA methylation profiles were identified in patients. One subtype (60.3% of patients) showed relatively limited changes in methylation levels and cell composition compared to controls, while a second subtype (39.7% of patients) exhibited widespread methylation level alterations among genes enriched in immune cell activity, as well as a higher proportion of neutrophils and lower proportion of lymphocytes. Differentiation of the two patient subtypes was validated by TCR repertoires, which paralleled the partition based on DNA methylation profiles. The subtype with widespread methylation modifications had higher symptom severity, performed worse on cognitive measures, and displayed greater reductions in fractional anisotropy of white matter tracts and evidence of gray matter thickening compared to the other subtype. Identification of a distinct subtype of schizophrenia with unique molecular, cerebral, and clinical features provide a novel parcellation of the schizophrenia syndrome with potential to guide development of individualized therapeutics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
Analysis code are available upon request to the corresponding authors.
References
Clementz BA, Sweeney JA, Hamm JP, Ivleva EI, Ethridge LE, Pearlson GD, et al. Identification of distinct psychosis biotypes using brain-based biomarkers. Am J Psychiatry. 2016;173:373–84.
Almendro V, Marusyk A, Polyak K. Cellular heterogeneity and molecular evolution in cancer. Annu Rev Pathol. 2013;8:277–302.
Schmitt A, Martins-de-Souza D, Akbarian S, Cassoli JS, Ehrenreich H, Fischer A, et al. Consensus paper of the WFSBP Task Force on Biological Markers: criteria for biomarkers and endophenotypes of schizophrenia, part III: molecular mechanisms. World J Biol Psychiatry: Off J World Federation Societies Biol Psychiatry. 2017;18:330–56.
Sun H, Lui S, Yao L, Deng W, Xiao Y, Zhang W, et al. Two patterns of white matter abnormalities in medication-naive patients with first-episode schizophrenia revealed by diffusion tensor imaging and cluster analysis. JAMA Psychiatry. 2015;72:678–86.
Owen MJ, Sawa A, Mortensen PB. Schizophrenia. Lancet. 2016;388:86–97.
Smigielski L, Jagannath V, Rössler W, Walitza S, Grünblatt E. Epigenetic mechanisms in schizophrenia and other psychotic disorders: a systematic review of empirical human findings. Mol Psychiatry. 2020;25:1718–48.
Liu J, Chen J, Ehrlich S, Walton E, White T, Perrone-Bizzozero N, et al. Methylation patterns in whole blood correlate with symptoms in schizophrenia patients. Schizophr Bull. 2014;40:769–76.
Melas PA, Rogdaki M, Ösby U, Schalling M, Lavebratt C, Ekström TJ. Epigenetic aberrations in leukocytes of patients with schizophrenia: association of global DNA methylation with antipsychotic drug treatment and disease onset. FASEB J: Off Publ Federation Am Societies Exp Biol. 2012;26:2712–8.
Grayson DR, Jia X, Chen Y, Sharma RP, Mitchell CP, Guidotti A, et al. Reelin promoter hypermethylation in schizophrenia. Proc Natl Acad Sci USA. 2005;102:9341–6.
Tao R, Davis KN, Li C, Shin JH, Gao Y, Jaffe AE, et al. GAD1 alternative transcripts and DNA methylation in human prefrontal cortex and hippocampus in brain development, schizophrenia. Mol Psychiatry. 2018;23:1496–505.
Gao S, Cheng J, Li G, Sun T, Xu Y, Wang Y, et al. Catechol-O-methyltransferase gene promoter methylation as a peripheral biomarker in male schizophrenia. Eur Psychiatry: J Assoc Eur Psychiatrists. 2017;44:39–46.
Abdolmaleky HM, Yaqubi S, Papageorgis P, Lambert AW, Ozturk S, Sivaraman V, et al. Epigenetic dysregulation of HTR2A in the brain of patients with schizophrenia and bipolar disorder. Schizophrenia Res. 2011;129:183–90.
Cheah SY, McLeay R, Wockner LF, Lawford BR, Young RM, Morris CP, et al. Expression and methylation of BDNF in the human brain in schizophrenia. World J Biol Psychiatry: Off J World Federation Societies Biol Psychiatry. 2017;18:392–400.
Rubin LH, Connelly JJ, Reilly JL, Carter CS, Drogos LL, Pournajafi-Nazarloo H, et al. Sex and diagnosis specific associations between DNA methylation of the oxytocin receptor gene with emotion processing and temporal-limbic and prefrontal brain volumes in psychotic disorders. Biol Psychiatry Cogn Neurosci Neuroimaging. 2016;1:141–51.
Montano C, Taub MA, Jaffe A, Briem E, Feinberg JI, Trygvadottir R, et al. Association of DNA methylation differences with schizophrenia in an epigenome-wide association study. JAMA Psychiatry. 2016;73:506–14.
Aberg KA, McClay JL, Nerella S, Clark S, Kumar G, Chen W, et al. Methylome-wide association study of schizophrenia: identifying blood biomarker signatures of environmental insults. JAMA Psychiatry. 2014;71:255–64.
Chan RF, Shabalin AA, Montano C, Hannon E, Hultman CM, Fallin MD, et al. Independent methylome-wide association studies of schizophrenia detect consistent case-control differences. Schizophr Bull. 2020;46:319–27.
Hannon E, Dempster E, Viana J, Burrage J, Smith AR, Macdonald R, et al. An integrated genetic-epigenetic analysis of schizophrenia: evidence for co-localization of genetic associations and differential DNA methylation. Genome Biol. 2016;17:176.
Jaffe AE, Gao Y, Deep-Soboslay A, Tao R, Hyde TM, Weinberger DR, et al. Mapping DNA methylation across development, genotype and schizophrenia in the human frontal cortex. Nat Neurosci. 2016;19:40–7.
Wockner LF, Noble EP, Lawford BR, Young RM, Morris CP, Whitehall VL, et al. Genome-wide DNA methylation analysis of human brain tissue from schizophrenia patients. Transl Psychiatry. 2014;4:e339.
Pai S, Li P, Killinger B, Marshall L, Jia P, Liao J, et al. Differential methylation of enhancer at IGF2 is associated with abnormal dopamine synthesis in major psychosis. Nat Commun. 2019;10:2046.
Chen C, Zhang C, Cheng L, Reilly JL, Bishop JR, Sweeney JA, et al. Correlation between DNA methylation and gene expression in the brains of patients with bipolar disorder and schizophrenia. Bipolar Disord. 2014;16:790–9.
Braun PR, Han S, Hing B, Nagahama Y, Gaul LN, Heinzman JT, et al. Genome-wide DNA methylation comparison between live human brain and peripheral tissues within individuals. Transl Psychiatry. 2019;9:47.
First M, Spitzer R, Gibbon M, Williams J. Structured Clinical Interview for DSM-IV Axis I Disorders, Patient Edition (SCID-I/P). New York: Biometrics Research, New York State Psychiatry Institute; 1995.
Singh SP, Cooper JE, Fisher HL, Tarrant CJ, Lloyd T, Banjo J, et al. Determining the chronology and components of psychosis onset: The Nottingham Onset Schedule (NOS). Schizophr Res. 2005;80:117–30.
Gardner DM, Murphy AL, O’Donnell H, Centorrino F, Baldessarini RJ. International consensus study of antipsychotic dosing. Am J psychiatry. 2010;167:686–93.
Kay SR, Opler LA, Lindenmayer JP. The Positive and Negative Syndrome Scale (PANSS): rationale and standardisation. Br J Psychiatry Suppl 1989; 59–67.
Keefe RS, Goldberg TE, Harvey PD, Gold JM, Poe MP, Coughenour L. The Brief Assessment of Cognition in Schizophrenia: reliability, sensitivity, and comparison with a standard neurocognitive battery. Schizophr Res. 2004;68:283–97.
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, et al. Minfi: a flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinforma (Oxf, Engl). 2014;30:1363–9.
Fortin JP, Triche TJ Jr., Hansen KD. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinforma (Oxf, Engl). 2017;33:558–60.
Teschendorff AE, Marabita F, Lechner M, Bartlett T, Tegner J, Gomez-Cabrero D, et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinforma (Oxf, Engl). 2013;29:189–96.
Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Gayther SA, Apostolidou S, et al. An epigenetic signature in peripheral blood predicts active ovarian cancer. PloS ONE. 2009;4:e8274.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinforma (Oxf, Engl). 2012;28:882–3.
Brunet JP, Tamayo P, Golub TR, Mesirov JP. Metagenes and molecular pattern discovery using matrix factorization. Proc Natl Acad Sci USA. 2004;101:4164–9.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9.
Piñero J, Queralt-Rosinach N, Bravo À, Deu-Pons J, Bauer-Mehren A, Baron M, et al. DisGeNET: a discovery platform for the dynamical exploration of human diseases and their genes. Database: J Biol Databases Curation. 2015;2015:bav028.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics: a J Integr Biol. 2012;16:284–7.
Peters TJ, Buckley MJ, Statham AL, Pidsley R, Samaras K, VL R, et al. De novo identification of differentially methylated regions in the human genome. Epigenetics Chromatin. 2015;8:6.
Rahmani E, Yedidim R, Shenhav L, Schweiger R, Weissbrod O, Zaitlen N, et al. GLINT: a user-friendly toolset for the analysis of high-throughput DNA-methylation array data. Bioinforma (Oxf, Engl). 2017;33:1870–2.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinforma. 2012;13:86.
Koestler DC, Jones MJ, Usset J, Christensen BC, Butler RA, Kobor MS, et al. Improving cell mixture deconvolution by identifying optimal DNA methylation libraries (IDOL). BMC Bioinforma. 2016;17:120.
Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlén SE, Greco D, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PloS ONE. 2012;7:e41361.
Shugay M, Bagaev DV, Turchaninova MA, Bolotin DA, Britanova OV, Putintseva EV, et al. VDJtools: unifying post-analysis of T cell receptor repertoires. PLoS Computational Biol. 2015;11:e1004503.
Cannon TD, Kaprio J, Lonnqvist J, Huttunen M, Koskenvuo M. The genetic epidemiology of schizophrenia in a Finnish twin cohort. A population-based modeling study. Arch Gen Psychiatry. 1998;55:67–74.
van Os J, Rutten BP, Poulton R. Gene-environment interactions in schizophrenia: review of epidemiological findings and future directions. Schizophr Bull. 2008;34:1066–82.
Richetto J, Meyer U. Epigenetic modifications in Schizophrenia and related disorders: molecular scars of environmental exposures and source of phenotypic variability. Biol Psychiatry. 2020;89:215–26.
Gürel Ç, Kuşçu GC, Yavaşoğlu A, Biray, Avcı Ç. The clues in solving the mystery of major psychosis: the epigenetic basis of schizophrenia and bipolar disorder. Neurosci Biobehav Rev. 2020;113:51–61.
Alharbi RA, Pettengell R, Pandha HS, Morgan R. The role of HOX genes in normal hematopoiesis and acute leukemia. Leukemia. 2013;27:1000–8.
Bhatlekar S, Fields JZ, Boman BM. Role of HOX genes in stem cell differentiation and cancer. Stem Cells Int. 2018;2018:3569493.
Mandeville I, Aubin J, LeBlanc M, Lalancette-Hebert M, Janelle MF, Tremblay GM, et al. Impact of the loss of Hoxa5 function on lung alveogenesis. Am J Pathol. 2006;169:1312–27.
Person RE, Li FQ, Duan Z, Benson KF, Wechsler J, Papadaki HA, et al. Mutations in proto-oncogene GFI1 cause human neutropenia and target ELA2. Nat Genet. 2003;34:308–12.
Xia J, Bolyard AA, Rodger E, Stein S, Aprikyan AA, Dale DC, et al. Prevalence of mutations in ELANE, GFI1, HAX1, SBDS, WAS and G6PC3 in patients with severe congenital neutropenia. Br J Haematol. 2009;147:535–42.
Velu CS, Baktula AM, Grimes HL. Gfi1 regulates miR-21 and miR-196b to control myelopoiesis. Blood. 2009;113:4720–8.
Zahorec R. Ratio of neutrophil to lymphocyte counts–rapid and simple parameter of systemic inflammation and stress in critically ill. Bratisl Lek Listy. 2001;102:5–14.
Steiner J, Frodl T, Schiltz K, Dobrowolny H, Jacobs R, Fernandes BS, et al. Innate immune cells and C-reactive protein in acute first-episode psychosis and schizophrenia: relationship to psychopathology and treatment. Schizophr Bull. 2020;46:363–73.
Bustan Y, Drapisz A, Ben Dor DH, Avrahami M, Schwartz-Lifshitz M, Weizman A, et al. Elevated neutrophil to lymphocyte ratio in non-affective psychotic adolescent inpatients: evidence for early association between inflammation and psychosis. Psychiatry Res. 2018;262:149–53.
Brinn A, Stone J. Neutrophil-lymphocyte ratio across psychiatric diagnoses: a cross-sectional study using electronic health records. BMJ Open. 2020;10:e036859.
Trepanier MO, Hopperton KE, Mizrahi R, Mechawar N, Bazinet RP. Postmortem evidence of cerebral inflammation in schizophrenia: a systematic review. Mol Psychiatry. 2016;21:1009–26.
van Kesteren CF, Gremmels H, de Witte LD, Hol EM, Van Gool AR, Falkai PG, et al. Immune involvement in the pathogenesis of schizophrenia: a meta-analysis on postmortem brain studies. Transl Psychiatry. 2017;7:e1075.
Brown AS, Derkits EJ. Prenatal infection and schizophrenia: a review of epidemiologic and translational studies. Am J Psychiatry. 2010;167:261–80.
Khandaker GM, Zimbron J, Dalman C, Lewis G, Jones PB. Childhood infection and adult schizophrenia: a meta-analysis of population-based studies. Schizophrenia Res. 2012;139:161–8.
Fillman SG, Weickert TW, Lenroot RK, Catts SV, Bruggemann JM, Catts VS, et al. Elevated peripheral cytokines characterize a subgroup of people with schizophrenia displaying poor verbal fluency and reduced Broca’s area volume. Mol Psychiatry. 2016;21:1090–8.
Lizano P, Lutz O, Xu Y, Rubin LH, Paskowitz L, Lee AM, et al. Multivariate relationships between peripheral inflammatory marker subtypes and cognitive and brain structural measures in psychosis. Mol Psychiatry 2020. https://doi.org/10.1038/s41380-020-00914-0. e-pub ahead of print 15 October 2020.
Gonzalez H, Pacheco R. T-cell-mediated regulation of neuroinflammation involved in neurodegenerative diseases. J Neuroinflammation. 2014;11:201.
Engelhardt B. Molecular mechanisms involved in T cell migration across the blood-brain barrier. J Neural Transm (Vienna). 2006;113:477–85.
Prinz M, Priller J. The role of peripheral immune cells in the CNS in steady state and disease. Nat Neurosci. 2017;20:136–44.
Larochelle C, Alvarez JI, Prat A. How do immune cells overcome the blood-brain barrier in multiple sclerosis? FEBS Lett. 2011;585:3770–80.
Montagne A, Zhao Z, Zlokovic BV. Alzheimer’s disease: a matter of blood-brain barrier dysfunction? J Exp Med. 2017;214:3151–69.
Zenaro E, Pietronigro E, Della Bianca V, Piacentino G, Marongiu L, Budui S, et al. Neutrophils promote Alzheimer’s disease-like pathology and cognitive decline via LFA-1 integrin. Nat Med. 2015;21:880–6.
Núñez C, Stephan-Otto C, Usall J, Bioque M, Lobo A, González-Pinto A, et al. Neutrophil count is associated with reduced gray matter and enlarged ventricles in first-episode psychosis. Schizophr Bull. 2019;45:846–58.
Hanson DR, Gottesman II. Theories of schizophrenia: a genetic-inflammatory-vascular synthesis. BMC Med Genet. 2005;6:7.
Pong S, Karmacharya R, Sofman M, Bishop JR, Lizano P. The role of brain microvascular endothelial cell and blood-brain barrier dysfunction in Schizophrenia. Complex Psychiatry. 2020;6:30–46.
Li Q, Zhou J, Cao X, Liu Q, Li Q, Li W, et al. Clonal characteristics of T-cell receptor repertoires in violent and non-violent patients with Schizophrenia. Front Psychiatry. 2018;9:403.
Britanova OV, Putintseva EV, Shugay M, Merzlyak EM, Turchaninova MA, Staroverov DB, et al. Age-related decrease in TCR repertoire diversity measured with deep and normalized sequence profiling. J Immunol (Baltim, Md: 1950). 2014;192:2689–98.
Kohler S, Thiel A. Life after the thymus: CD31+ and CD31− human naive CD4+ T-cell subsets. Blood. 2009;113:769–74.
Alves Sousa AP, Johnson KR, Ohayon J, Zhu J, Muraro PA, Jacobson S. Comprehensive analysis of TCR-beta repertoire in patients with neurological immune-mediated disorders. Sci Rep. 2019;9:344.
Amor S, Puentes F, Baker D, van der Valk P. Inflammation in neurodegenerative diseases. Immunology. 2010;129:154–69.
Debnath M. Adaptive immunity in Schizophrenia: functional implications of T cells in the etiology, course and treatment. J Neuroimmune Pharm. 2015;10:610–9.
Shah C, Zhang W, Xiao Y, Yao L, Zhao Y, Gao X, et al. Common pattern of gray-matter abnormalities in drug-naive and medicated first-episode schizophrenia: a multimodal meta-analysis. Psychol Med. 2017;47:401–13.
Pasternak O, Westin CF, Bouix S, Seidman LJ, Goldstein JM, Woo TU, et al. Excessive extracellular volume reveals a neurodegenerative pattern in schizophrenia onset. J Neurosci: Off J Soc Neurosci. 2012;32:17365–72.
Abel KM, Drake R, Goldstein JM. Sex differences in schizophrenia. Int Rev Psychiatry (Abingdon, Engl). 2010;22:417–28.
Mendrek A, Mancini-Marïe A. Sex/gender differences in the brain and cognition in schizophrenia. Neurosci Biobehav Rev. 2016;67:57–78.
Kirkpatrick B, Buchanan RW, Ross DE, Carpenter WT Jr. A separate disease within the syndrome of schizophrenia. Arch Gen Psychiatry. 2001;58:165–71.
Zhou J, Li M, Wang X, He Y, Xia Y, Sweeney J, et al. Drug response-related DNA methylation changes in Schizophrenia, bipolar disorder, and major depressive disorder. Front Neurosci. 2021;15:674273.
Funding
This study was supported by National Natural Science Foundation of China (grants 82120108014, 82071908, 81761128023, 81901702 and 81901705); China Postdoctoral Science Foundation (No.2020M673243); Sichuan Science and Technology Program (No. 2021JDTD0002 and 2020YFS0117); 1.3.5 project for disciplines of excellence, Post-Doctoral Research Project, West China Hospital, Sichuan University (Grant: 2ZYJC18020, ZYYC08001, and 018HXBH058) and the University of Cincinnati Schizophrenia Research Fund. LS also acknowledges the support from Humboldt Foundation Friedrich Wihelm Bessel Research Award and Chang Jiang Scholars.
Author information
Authors and Affiliations
Contributions
LS, GQ, XD and LC contributed to study design. LC, XY, HN, LS and WX contributed to acquisition of data. LC, PX and WX contributed to analysis and interpretation of the data. LC and PX contributed to the drafting of the paper. SJ, BJ, LS and XD made critical revision of the paper for important intellectual content. LS, GQ, XD and LC had full access to all data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Luo, C., Pi, X., Hu, N. et al. Subtypes of schizophrenia identified by multi-omic measures associated with dysregulated immune function. Mol Psychiatry 26, 6926–6936 (2021). https://doi.org/10.1038/s41380-021-01308-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-021-01308-6
This article is cited by
-
Immunophenotypes in psychosis: is it a premature inflamm-aging disorder?
Molecular Psychiatry (2024)
-
DNA methylation and the opposing NMDAR dysfunction in schizophrenia and major depression disorders: a converging model for the therapeutic effects of psychedelic compounds in the treatment of psychiatric illness
Molecular Psychiatry (2023)
-
Biological subtyping of psychiatric syndromes as a pathway for advances in drug discovery and personalized medicine
Nature Mental Health (2023)
-
A Potential Immune-Related miRNAs Regulatory Network and Corresponding Diagnostic Efficacy in Schizophrenia
Neurochemical Research (2023)
-
Neuroimaging biomarkers define neurophysiological subtypes with distinct trajectories in schizophrenia
Nature Mental Health (2023)