Abstract
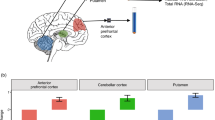
Genome-wide association studies have identified single nucleotide polymorphisms (SNPs) associated with schizophrenia risk. Integration of RNA-sequencing data from postmortem human brains with these risk SNPs identified transcripts associated with increased schizophrenia susceptibility, including a class of exon 9-spliced isoforms of Sorting nexin-19 (SNX19d9) and an isoform of Arsenic methyltransferase (AS3MT) splicing out exons 2 and 3 (AS3MTd2d3). However, the biological function of these transcript variants is unclear. Defining the cell types where these risk transcripts are dominantly expressed is an important step to understand function, in prioritizing specific cell types and/or neural pathways in subsequent studies. To identify the cell type-specific localization of SNX19 and AS3MT in the human dorsolateral prefrontal cortex (DLPFC), we used single-molecule in situ hybridization techniques combined with automated quantification and machine learning approaches to analyze 10 postmortem brains of neurotypical individuals. These analyses revealed that both pan-SNX19 and pan-AS3MT were more highly expressed in neurons than non-neurons in layers II/III and VI of DLPFC. Furthermore, pan-SNX19 was preferentially expressed in glutamatergic neurons, while pan-AS3MT was preferentially expressed in GABAergic neurons. Finally, we utilized duplex BaseScope technology, to delineate the localization of SNX19d9 and AS3MTd2d3 splice variants, revealing consistent trends in spatial gene expression among pan-transcripts and schizophrenia risk-related transcript variants. These findings demonstrate that schizophrenia risk transcripts have distinct localization patterns in the healthy human brains, and suggest that SNX19 transcripts might disrupt the normal function of glutamatergic neurons, while AS3MT may lead to disturbances in the GABAergic system in the pathophysiology of schizophrenia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Pardiñas AF, Holmans P, Pocklington AJ, Escott-Price V, Ripke S, Carrera N, et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat Genet. 2018;50:381–9.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H, et al. Systematic localization of common disease-associated variation in regulatory DNA. Science. 2012;337:1190–5.
Genovese G, Fromer M, Stahl EA, Ruderfer DM, Chambert K, Landén M, et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat Neurosci. 2016;19:1433–41.
Takata A, Matsumoto N, Kato T. Genome-wide identification of splicing QTLs in the human brain and their enrichment among schizophrenia-associated loci. Nat Commun. 2017;8:14519.
Reble E, Dineen A, Barr CL. The contribution of alternative splicing to genetic risk for psychiatric disorders. Genes Brain Behav. 2018;17:e12430.
Walker RL, Ramaswami G, Hartl C, Mancuso N, Gandal MJ, de la Torre-Ubieta L, et al. Genetic control of expression and splicing in developing human brain informs disease mechanisms. Cell. 2019;179:750.e22.
Li J, Wang L, Jiang T, Wang J, Li X, Liu X, et al. eSNPO: An eQTL-based SNP Ontology and SNP functional enrichment analysis platform. Sci Rep. 2016;6:30595.
Rockman MV, Kruglyak L. Genetics of global gene expression. Nat Rev Genet. 2006;7:862–72.
Luo X-J, Mattheisen M, Li M, Huang L, Rietschel M, Børglum AD, et al. Systematic integration of brain eQTL and GWAS identifies ZNF323 as a novel schizophrenia risk gene and suggests recent positive selection based on compensatory advantage on pulmonary function. Schizophr Bull. 2015;41:1294–308.
Li M, Jaffe AE, Straub RE, Tao R, Shin JH, Wang Y, et al. A human-specific AS3MT isoform and BORCS7 are molecular risk factors in the 10q24.32 schizophrenia-associated locus. Nat Med. 2016;22:649–56.
Ma L, Semick SA, Chen Q, Li C, Tao R, Price AJ, et al. Schizophrenia risk variants influence multiple classes of transcripts of sorting nexin 19 (SNX19). Mol Psychiatry. 2020;25:831–43.
Sweet RA, Fish KN, Lewis DA. Mapping synaptic pathology within cerebral cortical circuits in subjects with schizophrenia. Front Hum Neurosci. 2010;4:44.
Velmeshev D, Schirmer L, Jung D, Haeussler M, Perez Y, Mayer S, et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science. 2019;364:685–9.
Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127.
Maynard KR, Collado-Torres L, Weber LM, Uytingco C, Barry BK, Williams SR, et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat Neurosci. 2021. https://doi.org/10.1038/s41593-020-00787-0. Epub ahead of print.
Erben L, He M-X, Laeremans A, Park E, Buonanno A. A novel ultrasensitive in situ hybridization approach to detect short sequences and splice variants with cellular resolution. Mol Neurobiol. 2018;55:6169–81.
Maynard KR, Tippani M, Takahashi Y, Phan BN, Hyde TM, Jaffe AE, et al. dotdotdot: an automated approach to quantify multiplex single molecule fluorescent in situ hybridization (smFISH) images in complex tissues. Nucleic Acids Res. 2020;48:e66.
Lipska BK, Deep-Soboslay A, Weickert CS, Hyde TM, Martin CE, Herman MM, et al. Critical factors in gene expression in postmortem human brain: Focus on studies in schizophrenia. Biol Psychiatry. 2006;60:650–8.
Wang F, Flanagan J, Su N, Wang LC, Bui S, Nielson A, et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn. 2012;14:22–9.
Weber GF, Menko AS. Color image acquisition using a monochrome camera and standard fluorescence filter cubes. BioTech. 2005;38:52.54,56.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Guillozet-Bongaarts AL, Hyde TM, Dalley RA, Hawrylycz MJ, Henry A, Hof PR, et al. Altered gene expression in the dorsolateral prefrontal cortex of individuals with schizophrenia. Mol Psychiatry. 2014;19:478–85.
Hoftman GD, Datta D, Lewis DA. Layer 3 excitatory and inhibitory circuitry in the prefrontal cortex: developmental trajectories and alterations in schizophrenia. Biol Psychiatry. 2017;81:862–73.
Zeng H, Shen EH, Hohmann JG, Oh SW, Bernard A, Royall JJ, et al. Large-scale cellular-resolution gene profiling in human neocortex reveals species-specific molecular signatures. Cell. 2012;149:483–96.
Zhang Y, Sloan SA, Clarke LE, Caneda C, Plaza CA, Blumenthal PD, et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron. 2016;89:37–53.
Alganem K, Shukla R, Eby H, Abel M, Zhang X, McIntyre WB, et al. Kaleidoscope: A New Bioinformatics Pipeline Web Application for In Silico Hypothesis Exploration of Omics Signatures. BioRxiv. 2020.05.01.070805; https://doi.org/10.1101/2020.05.01.070805.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47:D607–13.
Enwright Iii JF, Huo Z, Arion D, Corradi JP, Tseng G, Lewis DA. Transcriptome alterations of prefrontal cortical parvalbumin neurons in schizophrenia. Mol Psychiatry. 2018;23:1606–13.
Skene NG, Bryois J, Bakken TE, Breen G, Crowley JJ, Gaspar HA, et al. Genetic identification of brain cell types underlying schizophrenia. Nat Genet. 2018;50(May):825–33.
Hagemann-Jensen M, Ziegenhain C, Chen P, Ramsköld D, Hendriks GJ, Larsson AJM, et al. Single-cell RNA counting at allele and isoform resolution using Smart-seq3. Nat Biotechnol. 2020;38:708–14.
Mathys H, Davila-Velderrain J, Peng Z, Gao F, Mohammadi S, Young JZ, et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature. 2019;570:332–7.
Jolly S, Lang V, Koelzer VH, Sala Frigerio C, Magno L, Salinas PC, et al. Single-cell quantification of mRNA expression in the human brain. Sci Rep. 2019;9:12353.
Burgess DJ. Spatial transcriptomics coming of age. Nat Rev Genet. 2019;20:317.
Lein E, Borm LE, Linnarsson S. The promise of spatial transcriptomics for neuroscience in the era of molecular cell typing. Science. 2017;358:64–9.
Moncada R, Chiodin M, Devlin JC, Baron M, Hajdu CH, Simeone D, et al. Building a tumor atlas: integrating single-cell RNA-Seq data with spatial transcriptomics in pancreatic ductal adenocarcinoma. BioRxiv. 2018. https://doi.org/10.1101/254375.
Moncada R, Barkley D, Wagner F, Chiodin M, Devlin JC, Baron M, et al. Integrating microarray-based spatial transcriptomics and single-cell RNA-seq reveals tissue architecture in pancreatic ductal adenocarcinomas. Nat Biotechnol. 2020;38:333–42.
Maynard KR, Jaffe AE, Martinowich K. Spatial transcriptomics: putting genome-wide expression on the map. Neuropsychopharmacology. 2020;45:232–3.
Eng C-HL, Lawson M, Zhu Q, Dries R, Koulena N, Takei Y, et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature. 2019;568:235–9.
D’Ambrosio E, Dahoun T, Pardiñas AF, Veronese M, Bloomfield MAP, Jauhar S, et al. The effect of a genetic variant at the schizophrenia associated AS3MT/BORCS7 locus on striatal dopamine function: A PET imaging study. Psychiatry Res Neuroimaging. 2019;291:34–41.
Korovaitseva GI, Gabaeva MV, Yunilainen OA, Golimbet VE. Effect of VNTR polymorphism of the AS3MT gene and obstetrical complications on the severity of schizophrenia. Bull Exp Biol Med. 2019;168:84–6.
Jaffe AE, Straub RE, Shin JH, Tao R, Gao Y, Collado-Torres L, et al. Developmental and genetic regulation of the human cortex transcriptome illuminate schizophrenia pathogenesis. Nat Neurosci. 2018;21:1117–25.
Tao R, Davis KN, Li C, Shin JH, Gao Y, Jaffe AE, et al. GAD1 alternative transcripts and DNA methylation in human prefrontal cortex and hippocampus in brain development, schizophrenia. Mol Psychiatry. 2018;23:1496–505.
Ursini G, Punzi G, Chen Q, Marenco S, Robinson JF, Porcelli A, et al. Convergence of placenta biology and genetic risk for schizophrenia. Nat Med. 2018;24:792–801.
Acknowledgements
We are thankful for the contributions of the Clinical Brain Disorders Branch of the National Institute of Mental Health in assisting the Lieber Institute for Brain Development in the acquisition and curation of brain tissue donations. We are also thankful to Ran Tao and Richard Straub for their help with case selection. Funding for the study was provided by the Lieber Institute for Brain Development.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Takahashi, Y., Maynard, K.R., Tippani, M. et al. Single molecule in situ hybridization reveals distinct localizations of schizophrenia risk-related transcripts SNX19 and AS3MT in human brain. Mol Psychiatry 26, 3536–3547 (2021). https://doi.org/10.1038/s41380-021-01046-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-021-01046-9