Abstract
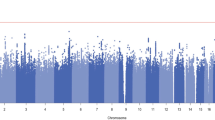
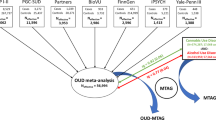
To provide insights into the biology of opioid dependence (OD) and opioid use (i.e., exposure, OE), we completed a genome-wide analysis comparing 4503 OD cases, 4173 opioid-exposed controls, and 32,500 opioid-unexposed controls, including participants of European and African descent (EUR and AFR, respectively). Among the variants identified, rs9291211 was associated with OE (exposed vs. unexposed controls; EUR z = −5.39, p = 7.2 × 10–8). This variant regulates the transcriptomic profiles of SLC30A9 and BEND4 in multiple brain tissues and was previously associated with depression, alcohol consumption, and neuroticism. A phenome-wide scan of rs9291211 in the UK Biobank (N > 360,000) found association of this variant with propensity to use dietary supplements (p = 1.68 × 10–8). With respect to the same OE phenotype in the gene-based analysis, we identified SDCCAG8 (EUR + AFR z = 4.69, p = 10–6), which was previously associated with educational attainment, risk-taking behaviors, and schizophrenia. In addition, rs201123820 showed a genome-wide significant difference between OD cases and unexposed controls (AFR z = 5.55, p = 2.9 × 10–8) and a significant association with musculoskeletal disorders in the UK Biobank (p = 4.88 × 10–7). A polygenic risk score (PRS) based on a GWAS of risk-tolerance (n = 466,571) was positively associated with OD (OD vs. unexposed controls, p = 8.1 × 10–5; OD cases vs. exposed controls, p = 0.054) and OE (exposed vs. unexposed controls, p = 3.6 × 10–5). A PRS based on a GWAS of neuroticism (n = 390,278) was positively associated with OD (OD vs. unexposed controls, p = 3.2 × 10–5; OD vs. exposed controls, p = 0.002) but not with OE (p = 0.67). Our analyses highlight the difference between dependence and exposure and the importance of considering the definition of controls in studies of addiction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cicero TJ, Ellis MS. Nonmedical prescription-opioid use and heroin use. N Engl J Med. 2016;374:1295–6.
Volkow ND, Frieden TR, Hyde PS, Cha SS. Medication-assisted therapies-tackling the opioid-overdose epidemic. N Engl J Med. 2014;370:2063–6.
Centers for Disease Control Prevention. CDC grand rounds: prescription drug overdoses - a U.S. epidemic. MMWR Morb Mortal Wkly Rep. 2012;61:10–13.
Rudd RA, Aleshire N, Zibbell JE, Gladden RM. Increases in drug and opioid overdose deaths-United States, 2000–2014. MMWR Morb Mortal Wkly Rep. 2016;64:1378–82.
Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, et al. 10 years of GWAS discovery: biology, function, and translation. Am J Hum Genet. 2017;101:5–22.
Sullivan PF, Agrawal A, Bulik CM, Andreassen OA, Borglum AD, Breen G, et al. Psychiatric genomics: an update and an agenda. Am J Psychiatry. 2018;175:15–27.
Nelson EC, Agrawal A, Heath AC, Bogdan R, Sherva R, Zhang B, et al. Evidence of CNIH3 involvement in opioid dependence. Mol Psychiatry. 2016;21:608–14.
Gelernter J, Kranzler HR, Sherva R, Koesterer R, Almasy L, Zhao H, et al. Genome-wide association study of opioid dependence: multiple associations mapped to calcium and potassium pathways. Biol Psychiatry. 2014;76:66–74.
Cheng Z, Zhou H, Sherva R, Farrer LA, Kranzler HR, Gelernter J. Genome-wide association study identifies a regulatory variant of RGMA associated with opioid dependence in European Americans. Biol Psychiatry. 2018;84:762–70.
Kalsi G, Euesden J, Coleman JR, Ducci F, Aliev F, Newhouse SJ, et al. Genome-wide association of heroin dependence in Han Chinese. PLoS ONE. 2016;11:e0167388.
Kelly TM, Daley DC, Douaihy AB. Treatment of substance abusing patients with comorbid psychiatric disorders. Addict Behav. 2012;37:11–24.
Clark RE, Baxter JD, Aweh G, O’Connell E, Fisher WH, Barton BA. Risk factors for relapse and higher costs among medicaid members with opioid dependence or abuse: opioid agonists, comorbidities, and treatment history. J Subst Abuse Treat. 2015;57:75–80.
Sanchez-Roige S, Palmer AA, Fontanillas P, Elson SL, Me Research Team tSUDWGotPGC, Adams MJ, et al. Genome-wide association study meta-analysis of the Alcohol Use Disorders Identification Test (AUDIT) in two population-based cohorts. Am J Psychiatry. 2019;176:107–18.
Kranzler HR, Zhou H, Kember RL, Vickers Smith R, Justice AC, Damrauer S, et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat Commun. 2019;10:1499.
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 4th edn, Text Revision (APA Publishing, Washington DC, USA, 2000).
Lam M, Awasthi S, Watson HJ, Goldstein J, Panagiotaropoulou G, Trubetskoy V, et al. RICOPILI: Rapid Imputation for COnsortias PIpeLIne. Bioinformatics 2020;36:930–3.
Walters RK, Polimanti R, Johnson EC, McClintick JN, Adams MJ, Adkins AE, et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat Neurosci. 2018;21:1656–69.
Lai D, Wetherill L, Bertelsen S, Carey CE, Kamarajan C, Kapoor M, et al. Genome-wide association studies of alcohol dependence, DSM-IV criterion count and individual criteria. Genes Brain Behav. 2019;18:e12579.
Consortium. GP, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
O’Connell J, Gurdasani D, Delaneau O, Pirastu N, Ulivi S, Cocca M, et al. A general approach for haplotype phasing across the full spectrum of relatedness. PLoS Genet. 2014;10:e1004234.
Howie BN, Donnelly P, Marchini J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009;5:e1000529.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12:e1001779.
Watanabe K, Stringer S, Frei O, Umicevic Mirkov M, de Leeuw C, Polderman TJC, et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat Genet. 2019;51:1339–48.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J. Schizophrenia Working Group of the Psychiatric Genomics Consortium. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet. 2015;47:291–5.
International HapMap Consortium, Altshuler DM, Gibbs RA, Peltonen L, Altshuler DM, Gibbs RA, et al. Integrating common and rare genetic variation in diverse human populations. Nature. 2010;467:52–58.
de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219.
Watanabe K, Taskesen E, van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1826.
Liberzon A, Subramanian A, Pinchback R, Thorvaldsdottir H, Tamayo P, Mesirov JP. Molecular signatures database (MSigDB) 3.0. Bioinformatics. 2011;27:1739–40.
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature. 2017;550:204–13.
Ramasamy A, Trabzuni D, Guelfi S, Varghese V, Smith C, Walker R, et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat Neurosci. 2014;17:1418–28.
Sul JH, Han B, Ye C, Choi T, Eskin E. Effectively identifying eQTLs from multiple tissues by combining mixed model and meta-analytic approaches. PLoS Genet. 2013;9:e1003491.
Euesden J, Lewis CM, O’Reilly PF. PRSice: Polygenic Risk Score software. Bioinformatics. 2015;31:1466–8.
Karlsson Linner R, Biroli P, Kong E, Meddens SFW, Wedow R, Fontana MA, et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat Genet. 2019;51:245–57.
Nagel M, Jansen PR, Stringer S, Watanabe K, de Leeuw CA, Bryois J, et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat Genet. 2018;50:920–7.
Liu M, Jiang Y, Wedow R, Li Y, Brazel DM, Chen F, et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet. 2019;51:237–44.
Primes G, Fieder M. Real-life helping behaviours in North America: A genome-wide association approach. PLoS ONE. 2018;13:e0190950.
Kreek MJ, Nielsen DA, Butelman ER, LaForge KS. Genetic influences on impulsivity, risk taking, stress responsivity and vulnerability to drug abuse and addiction. Nat Neurosci. 2005;8:1450–7.
Sirugo G, Williams SM, Tishkoff SA. The missing diversity in human genetic studies. Cell. 2019;177:26–31.
Pasman JA, Verweij KJH, Gerring Z, Stringer S, Sanchez-Roige S, Treur JL, et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat Neurosci. 2018;21:1161–70.
Howard DM, Adams MJ, Clarke TK, Hafferty JD, Gibson J, Shirali M, et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat Neurosci. 2019;22:343–52.
Gelernter J, Sun N, Polimanti R, Pietrzak RH, Levey D, Bryois J, et al. Genome-wide association study of post-traumatic stress disorder reexperiencing symptoms in >165,000 US veterans. Nat Neurosci 2019;22:1394–1401.
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. 2019;51:431–44.
Kichaev G, Bhatia G, Loh PR, Gazal S, Burch K, Freund MK, et al. Leveraging polygenic functional enrichment to improve GWAS power. Am J Hum Genet. 2019;104:65–75.
Perez Y, Shorer Z, Liani-Leibson K, Chabosseau P, Kadir R, Volodarsky M, et al. SLC30A9 mutation affecting intracellular zinc homeostasis causes a novel cerebro-renal syndrome. Brain. 2017;140:928–39.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Cole JB, Florez JC, Hirschhorn JN. Comprehensive genomic analysis of dietary habits in UK Biobank identifies hundreds of genetic loci and establishes causal relationships between educational attainment and healthy eating. bioRxiv. 2019:662239.
MacArthur J, Bowler E, Cerezo M, Gil L, Hall P, Hastings E, et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 2017;45:D896–D901.
Lee JJ, Wedow R, Okbay A, Kong E, Maghzian O, Zacher M, et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet. 2018;50:1112–21.
Hoffmann TJ, Ehret GB, Nandakumar P, Ranatunga D, Schaefer C, Kwok PY, et al. Genome-wide association analyses using electronic health records identify new loci influencing blood pressure variation. Nat Genet. 2017;49:54–64.
Yasukochi Y, Sakuma J, Takeuchi I, Kato K, Oguri M, Fujimaki T, et al. Identification of CDC42BPG as a novel susceptibility locus for hyperuricemia in a Japanese population. Mol Genet Genomics. 2018;293:371–9.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7.
Matsuda A, Suzuki Y, Honda G, Muramatsu S, Matsuzaki O, Nagano Y, et al. Large-scale identification and characterization of human genes that activate NF-kappaB and MAPK signaling pathways. Oncogene. 2003;22:3307–18.
Busse JW, Craigie S, Sadeghirad B, Couban R, Hong P, Oparin Y, et al. Management of acute musculoskeletal pain (excluding low back pain): protocol for a systematic review and network meta-analysis of randomised trials. BMJ Open. 2019;9:e024441.
Jiang W, Yu W. Power estimation and sample size determination for replication studies of genome-wide association studies. BMC Genomics. 2016;17:3.
Zou J, Zhou J, Faller S, Brown R, Eskin E. Accurate modeling of replication rates in genome-wide association studies by accounting for winner’s curse and study-specific heterogeneity. bioRxiv 2019: 856898.
Sullivan PF, Geschwind DH. Defining the genetic, genomic, cellular, and diagnostic architectures of psychiatric disorders. Cell. 2019;177:162–83.
Acknowledgements
The Psychiatric Genomics Consortium Substance Use Disorders Working Group receives support from the National Institute on Drug Abuse and the National Institute of Mental Health via U01 MH109532 and U01 MH109528. We gratefully acknowledge prior support from the National Institute on Alcohol Abuse and Alcoholism. Statistical analyses for the PGC were carried out on the Genetic Cluster Computer (http://www.geneticcluster.org) hosted by SURFsara and financially supported by the Netherlands Scientific Organization (NWO 480-05-003) along with a supplement from the Dutch Brain Foundation and the VU University Amsterdam. A.A. acknowledges DA032573; A.C.H. acknowledges support from NIH grants AA07535, AA07729, AA13320, AA13321, and AA11998; A.E.A. acknowledges support from AA011408 and AA017828; A.M.G. acknowledges support from U10 AA08401; B.P.R. was supported by AA011408, AA017828, and AA022537; B.T.W. acknowledges support from AA011408, AA017828, and AA022537; C.J.H. acknowledges DA032555, DA035804, DA011015, and DA042755; D.B.H. acknowledges support from R01DA036583; E.J.C. acknowledges support from DA023026, DA011301, and DA024413; E.O.J. acknowledges support from R01 DA044014; H.M. acknowledges support from DA025109, DA024413, and DA016977; J.G. acknowledges support from DA12690 and DA047527; J.K.H. acknowledges support from DA011015; K.S.K. acknowledges support from AA011408, AA017828, and AA022537; L.D. is supported by an Australian National Health and Medical Research Council (NHMRC) Principal Research Fellowship; L.J.B. acknowledges support from R01DA036583; L.M.H. acknowledges support from AA011408 and AA017828; L.M.H. acknowledges support from AA011408 and AA017828; M.C.S. acknowledges support from DA035804; P.A.F.M. acknowledges funding support from NIH grants: DA012854 and R25DA027995; R.A.G. acknowledges support from AA017444; R.E.P. is supported by NIH K01 grant MH113848; R.P. acknowledges support from DA12690 and DA047527; S.A.B. acknowledges support from AA011408, AA017828, AA022537, and AA022717; S.M.H. acknowledges support from R21AA024888 and K08DA032680; T.B.B. acknowledges support from MH100549; T.L.W. acknowledges support from R01 DA021905 and R01 DA035804; W.E.C. acknowledges support from R01HD093651, R01DA036523, and P30DA023026, R01MH117559. Alcohol Dependence in African Americans (ADAA) study was funded by NIH grant R01 AA017444. Funding support for the Comorbidity and Trauma Study (CATS) (dbGAP accession number: phs000277.v1.p1) was provided by the National Institute on Drug Abuse (R01 DA17305); GWAS genotyping services at the CIDR at The Johns Hopkins University were supported by the National Institutes of Health (contract N01-HG-65403). The data collection and analysis of the Center on Antisocial Drug Dependence (CADD) was supported by the following grants: DA011015, DA012845, DA021913, DA021905, DA032555, and DA035804. The Collaborative Study on the Genetics of Alcoholism (COGA) is supported by NIH Grant U10AA008401 from the National Institute on Alcohol Abuse and Alcoholism (NIAAA) and the National Institute on Drug Abuse (NIDA). Funding support for this GWAS genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research, was provided by the National Institute on Alcohol Abuse and Alcoholism, the NIH GEI (U01HG004438), and the NIH contract “High throughput genotyping for studying the genetic contributions to human disease” (HHSN268200782096C). COGA Principal Investigators: B. Porjesz, V. Hesselbrock, H. Edenberg, L. Bierut, includes eleven different centers: University of Connecticut (V. Hesselbrock); Indiana University (H.J. Edenberg, J. Nurnberger Jr., T. Foroud); University of Iowa (S. Kuperman, J. Kramer); SUNY Downstate (B. Porjesz); Washington University in St. Louis (L. Bierut, J. Rice, K. Bucholz, A. Agrawal); University of California at San Diego (M. Schuckit); Rutgers University (J. Tischfield, A. Brooks); Department of Biomedical and Health Informatics, The Children’s Hospital of Philadelphia; Department of Genetics, Perelman School of Medicine, University of Pennsylvania, Philadelphia PA (L. Almasy), Virginia Commonwealth University (D. Dick), Icahn School of Medicine at Mount Sinai (A. Goate), and Howard University (R. Taylor). Other COGA collaborators include: L. Bauer (University of Connecticut); J. McClintick, L. Wetherill, X. Xuei, Y. Liu, D. Lai, S. O’Connor, M. Plawecki, S. Lourens (Indiana University); G. Chan (University of Iowa; University of Connecticut); J. Meyers, D. Chorlian, C. Kamarajan, A. Pandey, J. Zhang (SUNY Downstate); J.-C. Wang, M. Kapoor, S. Bertelsen (Icahn School of Medicine at Mount Sinai); A. Anokhin, V. McCutcheon, S. Saccone (Washington University); J. Salvatore, F. Aliev, B. Cho (Virginia Commonwealth University); and Mark Kos (University of Texas Rio Grande Valley). A. Parsian are the NIAAA Staff Collaborators. M. Reilly was an NIAAA staff collaborator. We continue to be inspired by our memories of Henri Begleiter and Theodore Reich, founding PI and Co-PI of COGA, and also owe a debt of gratitude to other past organizers of COGA, including Ting-Kai Li, currently a consultant with COGA, P. Michael Conneally, Raymond Crowe, and Wendy Reich, for their critical contributions. We thank Kim Doheny and Elizabeth Pugh from CIDR and Justin Paschall from the NCBI dbGaP staff for valuable assistance with genotyping and quality control in developing the dataset available at dbGaP (phs000125.v1.p1; also: phs000763.v1.p1; phs000976.v1.p1). Support for the Study of Addiction: Genetics and Environment (SAGE) was provided through the NIH Genes, Environment and Health Initiative [GEI; U01 HG004422; dbGaP study accession phs000092.v1.p1]. SAGE is one of the genome-wide association studies funded as part of the Gene Environment Association Studies (GENEVA) under GEI. Assistance with phenotype harmonization and genotype cleaning, as well as with general study coordination, was provided by the GENEVA Coordinating Center [U01 HG004446]. Assistance with data cleaning was provided by the National Center for Biotechnology Information. Support for collection of datasets and samples was provided by the Collaborative Study on the Genetics of Alcoholism [COGA; U10 AA008401], the Collaborative Genetic Study of Nicotine Dependence [COGEND; P01 CA089392; see also phs000404.v1.p1], and the Family Study of Cocaine Dependence [FSCD; R01 DA013423, R01 DA019963]. Funding support for genotyping, which was performed at the Johns Hopkins University Center for Inherited Disease Research (CIDR), was provided by the NIH GEI [U01HG004438], the National Institute on Alcohol Abuse and Alcoholism, the National Institute on Drug Abuse, and the NIH contract “High throughput genotyping for studying the genetic contributions to human disease” [HHSN268200782096C]. The Gene–Environment Development Initiative: Great Smoky Mountains Study (phs000852.v1.p1) was supported by the National Institute on Drug Abuse (U01DA024413, R01DA11301), the National Institute of Mental Health (R01MH063970, R01MH063671, R01MH048085, K01MH093731, and K23MH080230), NARSAD, and the William T. Grant Foundation. We are grateful to all the GSMS and CCC study participants who contributed to this work. The following grants supported data collection and analysis of CADD: DA011015, DA012845, DA021913, DA021905, DA032555, and DA035804. Gene-Environment-Development Initiative -GEDI – Virginia Commonwealth University (VTSABD; dbGAP in progress) was supported by the National Institute on Drug Abuse (U01DA024413, R01DA025109), the National Institute of Mental Health (R01MH045268, R01MH055557, and R01MH068521), and the Virginia Tobacco Settlement Foundation grant 8520012. We are grateful to all the VTSABD-YAFU-TSA study participants who contributed to this work. Yale-Penn (phs000425.v1.p1; phs000952.v1.p1) was supported by National Institutes of Health Grants RC2 DA028909, R01 DA12690, R01 DA12849, R01 DA18432, R01 AA11330, and R01 AA017535 and the Veterans Affairs Connecticut and Philadelphia Veterans Affairs Mental Illness Research, Educational, and Clinical Centers. Australian Alcohol and Nicotine studies (OZALC; phs000181.v1.p1) were supported by National Institutes of Health Grants AA07535,AA07728, AA13320, AA13321, AA14041, AA11998, AA17688,DA012854, and DA019951; by Grants from the Australian National Health and Medical Research Council (241944, 339462, 389927, 389875, 389891, 389892, 389938, 442915, 442981, 496739, 552485, and 552498); by Grants from the Australian Research Council (A7960034, A79906588, A79801419, DP0770096, DP0212016, and DP0343921); and by the 5th Framework Programme (FP-5) GenomEUtwin Project (QLG2-CT-2002-01254). Genome-wide association study genotyping at Center for Inherited Disease Research was supported by a Grant to the late Richard Todd, M.D., Ph.D., former Principal Investigator of Grant AA13320. Substance Use Disorder Working Group of the Psychiatric Genomics Consortium: Raymond K. Walters, Renato Polimanti, Emma C. Johnson, Jeanette N. McClintick, Mark J. Adams, Amy E. Adkins, Fazil Aliev, Silviu-Alin Bacanu, Anthony Batzler, Sarah Bertelsen, Joanna M. Biernacka, Tim B. Bigdeli, Li-Shiun Chen, Toni-Kim Clarke, Yi-Ling Chou, Franziska Degenhardt, Anna R. Docherty, Alexis C. Edwards, Pierre Fontanillas, Jerome C. Foo, Louis Fox, Josef Frank, Ina Giegling, Scott Gordon, Laura M. Hack, Annette M. Hartmann, Sarah M. Hartz, Stefanie Heilmann-Heimbach, Stefan Herms, Colin Hodgkinson, Per Hoffmann, Jouke Jan Hottenga, Martin A. Kennedy, Mervi Alanne-Kinnunen, Bettina Konte, Jari Lahti, Marius Lahti-Pulkkinen, Dongbing Lai, Lannie Ligthart, Anu Loukola, Brion S. Maher, Hamdi Mbarek, Andrew M. McIntosh, Matthew B. McQueen, Jacquelyn L. Meyers, Yuri Milaneschi, Teemu Palviainen, John F. Pearson, Roseann E. Peterson, Samuli Ripatti, Euijung Ryu, Nancy L. Saccone, Jessica E. Salvatore, Sandra Sanchez-Roige, Melanie Schwandt, Richard Sherva, Fabian Streit, Jana Strohmaier, Nathaniel Thomas, Jen-Chyong Wang, Bradley T. Webb, Robbee Wedow, Leah Wetherill, Amanda G. Wills, 23andMe Research Team, Jason D. Boardman, Danfeng Chen, Doo-Sup Choi, William E. Copeland, Robert C. Culverhouse, Norbert Dahmen, Louisa Degenhardt, Benjamin W. Domingue, Sarah L. Elson, Mark A. Frye, Wolfgang Gäbel, Caroline Hayward, Marcus Ising, Margaret Keyes, Falk Kiefer, John Kramer, Samuel Kuperman, Susanne Lucae, Michael T. Lynskey, Wolfgang Maier, Karl Mann, Satu Männistö, Bertram Müller-Myhsok, Alison D. Murray, John I. Nurnberger, Aarno Palotie, Ulrich Preuss, Katri Räikkönen, Maureen D. Reynolds, Monika Ridinger, Norbert Scherbaum, Marc A. Schuckit, Michael Soyka, Jens Treutlein, Stephanie Witt, Norbert Wodarz, Peter Zill, Daniel E. Adkins, Joseph M. Boden, Dorret I. Boomsma, Laura J. Bierut, Sandra A. Brown, Kathleen K. Bucholz, Sven Cichon, E. Jane Costello, Harriet de Wit, Nancy Diazgranados, Danielle M. Dick, Johan G. Eriksson, Lindsay A. Farrer, Tatiana M. Foroud, Nathan A. Gillespie, Alison M. Goate, David Goldman, Richard A. Grucza, Dana B. Hancock, Kathleen Mullan Harris, Andrew C. Heath, Victor Hesselbrock, John K. Hewitt, Christian J. Hopfer, John Horwood, William Iacono, Eric O. Johnson, Jaakko A. Kaprio, Victor M. Karpyak, Kenneth S. Kendler, Henry R. Kranzler, Kenneth Krauter, Paul Lichtenstein, Penelope A. Lind, Matt McGue, James MacKillop, Pamela A. F. Madden, Hermine H. Maes, Patrik Magnusson, Nicholas G. Martin, Sarah E. Medland, Grant W. Montgomery, Elliot C. Nelson, Markus M. Nöthen, Abraham A. Palmer, Nancy L. Pedersen, Brenda W. J. H. Penninx, Bernice Porjesz, John P. Rice, Marcella Rietschel, Brien P. Riley, Richard Rose, Dan Rujescu, Pei-Hong Shen, Judy Silberg, Michael C. Stallings, Ralph E. Tarter, Michael M. Vanyukov, Scott Vrieze, Tamara L. Wall, John B. Whitfield, Hongyu Zhao, Benjamin M. Neale, Joel Gelernter, Howard J. Edenberg & Arpana Agrawal
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
H.R.K. is a member of the American Society of Clinical Psychopharmacology’s Alcohol Clinical Trials Initiative, which over the last three years was sponsored by Alkermes, Ethypharm, Indivior, Lilly, Lundbeck, Otsuka, Pfizer, Arbor Pharmaceuticals, and Amygdala Neurosciences, Inc. H.R.K. and J.G. are named as inventors on PCT patent application #15/878,640 entitled: “Genotype-guided dosing of opioid agonists,” filed on 24 January 2018. L.J.B. and A.M.G. are listed as inventors on Issued U.S. Patent 8080,371, “Markers for Addiction” covering the use of certain SNPs in determining the diagnosis, prognosis, and treatment of addiction. The spouse of N.S. is listed as an inventor on Issued U.S. Patent 8,080,371, “Markers for Addiction” covering the use of certain SNPs in determining the diagnosis, prognosis, and treatment of addiction. The other authors do not report any conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Full list of Substance Use Disorder Working Group members appears in the Acknowledgments
Rights and permissions
About this article
Cite this article
Polimanti, R., Walters, R.K., Johnson, E.C. et al. Leveraging genome-wide data to investigate differences between opioid use vs. opioid dependence in 41,176 individuals from the Psychiatric Genomics Consortium. Mol Psychiatry 25, 1673–1687 (2020). https://doi.org/10.1038/s41380-020-0677-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-020-0677-9
This article is cited by
-
Cross-phenotype relationship between opioid use disorder and suicide attempts: new evidence from polygenic association and Mendelian randomization analyses
Molecular Psychiatry (2023)
-
Genome-wide analysis of a model-derived binge eating disorder phenotype identifies risk loci and implicates iron metabolism
Nature Genetics (2023)
-
Testing the causal relationships of physical activity and sedentary behaviour with mental health and substance use disorders: a Mendelian randomisation study
Molecular Psychiatry (2023)
-
Polygenic scores for psychiatric disorders in a diverse postmortem brain tissue cohort
Neuropsychopharmacology (2023)
-
Single-nucleus genomics in outbred rats with divergent cocaine addiction-like behaviors reveals changes in amygdala GABAergic inhibition
Nature Neuroscience (2023)