Abstract
Arousal affects cognition, emotion, and behavior and has been implicated in the etiology of psychiatric disorders. Although environmental conditions substantially contribute to the level of arousal, stable interindividual characteristics are well-established and a genetic basis has been suggested. Here we investigated the molecular genetics of brain arousal in the resting state by conducting a genome-wide association study (GWAS). We selected N = 1877 participants from the population-based LIFE-Adult cohort. Participants underwent a 20-min eyes-closed resting state EEG, which was analyzed using the computerized VIGALL 2.1 (Vigilance Algorithm Leipzig). At the SNP-level, GWAS analyses revealed no genome-wide significant locus (p < 5E-8), although seven loci were suggestive (p < 1E-6). The strongest hit was an expression quantitative trait locus (eQTL) of TMEM159 (lead-SNP: rs79472635, p = 5.49E-8). Importantly, at the gene-level, GWAS analyses revealed significant evidence for TMEM159 (p = 0.013, Bonferroni-corrected). By mapping our SNPs to the GWAS results from the Psychiatric Genomics Consortium, we found that all corresponding markers of TMEM159 showed nominally significant associations with Major Depressive Disorder (MDD; 0.006 ≤ p ≤ 0.011). More specifically, variants associated with high arousal levels have previously been linked to an increased risk for MDD. In line with this, the MetaXcan database suggests increased expression levels of TMEM159 in MDD, as well as Autism Spectrum Disorder, and Alzheimer’s Disease. Furthermore, our pathway analyses provided evidence for a role of sodium/calcium exchangers in resting state arousal. In conclusion, the present GWAS identifies TMEM159 as a novel candidate gene which may modulate the risk for psychiatric disorders through arousal mechanisms. Our results also encourage the elaboration of the previously reported interrelations between ion-channel modulators, sleep-wake behavior, and psychiatric disorders.
Similar content being viewed by others
Introduction
Arousal denotes a dimension of functional brain states, which are associated with behavioral phenomena including alert and relaxed wakefulness, drowsiness, as well as shallow and deep sleep. Arousal is linked to cognition, emotion, and behavior [1], and requires regulation and adaptation across varying environmental situations. For instance, arousal is increased in the presence of threat and reduced in situations of viable rest and recovery. Although environmental conditions substantially contribute to the level of arousal, stable interindividual characteristics are well-established and a genetic basis has been suggested [1, 2]. Recently, growing efforts have been made to investigate the molecular genetics of sleep-wake behavior, partly due to encouraging heritability estimates derived from family and twin studies [3, 4]. However, while genome-wide association studies (GWAS) have predominantly focused on parameters indicating the presence and absence of sleep (e.g., sleep duration, timing, and fragmentation) [5, 6], there remains a paucity of evidence concerning the genomics of arousal regulation during the transition from wakefulness to sleep onset.
Arousal has repeatedly been implicated in the etiology of interindividual differences in human behavior. According to the Arousal Regulation Model of Affective Disorders [7, 8], hypostable and hyperstable levels of arousal contribute to manic and depressive-like behavior, respectively. Several empirical studies have supported this model [9,10,11,12]. Notably, similar concepts have previously been proposed on the relationship between arousal and the personality traits extraversion [13] and sensation seeking [14]. In addition, the Research Domain Criteria Project (RDoC) of the US National Institute of Mental Health (NIMH) incorporated arousal as a principal construct into a cross-diagnostic framework designed to describe and classify mental disorders [15]. Despite this well-established theoretical background, objective and reliable assessments of arousal are needed.
One valuable methodology for assessing arousal is the analysis of the human electroencephalogram (EEG). Changes in the power of the EEG frequency bands are intimately related to alterations in arousal [16,17,18,19]. Recently, the EEG- and EOG-based Vigilance Algorithm Leipzig (VIGALL) was introduced [20, 21]. By incorporating information on the spectral composition and cortical distribution of EEG activity, VIGALL enables the computerized assessment of EEG-vigilance as an indicator of brain arousal in the resting state. The algorithm has been validated in functional imaging studies [22,23,24], against parameters of the autonomous nervous system [25, 26], and against subjective ratings [26]. Beyond conventional DSM and ICD disorder categories, VIGALL may constitute one tool to identify etiologically more homogenous psychiatric patient groups.
Recently, strong evidence has been found for shared genetic etiologies among schizophrenia, bipolar disorder, major depression, attention deficit hyperactivity disorder, and autism spectrum disorder [27]. Yet, little is known about the underlying pathophysiologies. Given its theoretical and empirical foundation, the regulation of arousal could be one mechanism linking genes and behavior. Indeed, genetic factors have previously been argued to impact on arousal regulation [1, 28]. In addition, the power of the arousal-indicating EEG frequency bands was observed with heritabilities of ~80% [29]. To date, there have been four GWAS on resting state EEG activity [29,30,31,32]. However, none has assessed arousal and its regulation. Consequently, prior studies used recording periods of <5 min, while longer recordings are preferred when investigating the dynamics of arousal in the resting state [26]. Notably, due to the high-interindividual variability in EEG activity, the assessment of arousal requires adaptive procedures that account for individual peak frequencies and total power. Such procedures have been accounted for in VIGALL.
On this basis, we set out to identify genetic variants linked to brain arousal in the resting state by conducting a genome-wide association study. In particular, we utilized the computerized algorithm VIGALL 2.1 to analyze resting state EEG recordings of 20 min duration without experimenter intervention. To evaluate the functional relevance of the obtained associations, we took into account publicly available gene expression data, performed gene-based analyses, and investigated the gene enrichment of biological pathways. Moreover, we conducted a GWAS catalog screening and queried our top hits in the Psychiatric Genomics Consortium data files. In addition, we derived candidate markers from the four GWAS on other resting EEG activity traits [29,30,31,32].
Methods and materials
Sample
Participants were Central Europeans drawn from the LIFE-Adult study, a population-based cohort study of 10,000 randomly selected inhabitants of the city of Leipzig, Germany [33]. The sample was composed of subjects aged 40–79 years with resting state EEG data obtained from 3462 subjects. Of those, 2727 subjects underwent genotyping and surpassed all corresponding quality checks (described below). We did not include subjects reporting current intakes of EEG-affecting drugs or who had a history of stroke, multiple sclerosis, Parkinson’s disease, epilepsy, skull fracture, cerebral tumor, or meningitis (leaving 2171 subjects). On the basis of data from structured clinical interviews for DSM-IV axis I disorders, we selected subjects without a history of substance dependence or psychotic disorders, and who were free of current affective and anxiety disorders (leaving 2087 subjects). Moreover, EEGs with substantial artifacts ( ≥ 15% of all EEG segments) and those displaying pathological activity, alpha variant rhythms, or low-voltage alpha were not included. The final sample comprised N = 1877 eligible subjects (942 male; mean age: 67.8 years). Participants gave written informed consent and received an expense allowance. All procedures were conducted according to the Declaration of Helsinki and were approved by the Ethics Committee of the University of Leipzig (263-2009-14122009).
Physiological data collection and processing
Physiological data were collected and processed, as previously described [26]. All EEGs were recorded according to the same standardized operating procedure. EEGs were recorded at ~0830, 1100, and 1330 hours. Subjects lay comfortably on a lounge chair within a sound-attenuated light-dimmed booth. To achieve similar initial levels of arousal, all subjects completed a brief arithmetic task that required them to count backwards by sixes starting at one hundred. Subsequently, subjects were instructed to close their eyes, relax and not to struggle against any potential drowsiness during the 20-min resting condition. Electroencephalic activity was recorded by 25 electrodes against the common average reference with AFz ground, was sampled at 1000 Hz, and amplified using a QuickAmp amplifier (Brain Products GmbH, Gilching, Germany). Impedances were kept below 10 kΩ. Vertical and horizontal eye movements were recorded by two bipolar electrodes. EEG offline processing was performed using Brain Vision Analyzer 2.0 (Brain Products GmbH, Gilching, Germany). EEGs were filtered (70 Hz low-pass and 0.5 Hz high-pass with 48 dB/Oct slope, 50 Hz notch) and rectified from muscle, sweating, cardiac, and eye movement artifacts using an extended infomax Independent Component Analysis (ICA) with classic principal component analysis (PCA) sphering. Sleep spindles and K-complexes were manually marked by experienced raters, as previously described [26]. For detailed preprocessing steps, please see also the publicly available VIGALL 2.1 manual [20].
Assessment of brain arousal
The assessment of brain arousal was performed as previously described [26]. Brain arousal was measured through resting state EEG-vigilance as determined using the Brain Vision Analyzer add-on VIGALL 2.1 (https://research.uni-leipzig.de/vigall) [20]. On the basis of the spectral composition and cortical distribution of electroencephalic activity, VIGALL assigns one of seven EEG-vigilance stages to each one-second EEG epoch. EEG-vigilance stages correspond to active wakefulness (stage 0), relaxed wakefulness (stages A1, A2, A3), drowsiness (stages B1, B2/3), and sleep onset (stage C). Assigned EEG-vigilance stages were transformed into values ranging from 7 (active wakefulness) to 1 (sleep onset). We calculated three outcome variables: mean vigilance, stability score, and slope index [11, 34]. While the former provides an estimate for the average level of EEG-vigilance during rest, the stability score and slope index particularly focus on the dynamics of EEG-vigilance. Values of mean vigilance range between 1 and 7, stability scores range between 1 and 14, and values of the slope index range between −7.2 and 1.2. Lower values indicate lower levels and steeper declines, respectively, of EEG-vigilance. All three outcome variables were validated, showed test-retest reliability, and have been established as an adequate way to summarize resting state EEG results [26, 34,35,36].
Genotyping, preprocessing, and imputation
Genotyping was performed at the Institute of Laboratory Medicine, University of Leipzig, Germany. Genomic DNA was extracted from peripheral blood leukocytes using an automated protocol on the Autopure LS instrument (Qiagen, Hilden, Germany). Genotyping was conducted using an Axiom Genome-Wide CEU 1 Array Plate (Affymetrix, Inc., Santa Clara, California, USA) with 587,352 single nucleotide polymorphisms (SNPs). Genotyping was performed according to the instructions of the manufacturer. Sample quality control included: dish-QC <0.82, call rate <0.97, reported vs. genotype-wise computed sex mismatch, and cryptic relatedness. On the basis of a subset of about 200,000 high-quality SNPs (call rate >0.998), we additionally conducted principle component analyses using smartpca 8000 of EIGENSOFT 3.0. Outliers (6 SD) were removed. In total, 4985 LIFE-Adult individuals were available for analysis. Irregularities in X–Y intensity plots were used to further remove seven samples for gonosomal analyses. Autosomal SNP quality control included: call rate <0.97, Hardy–Weinberg p-value <1E-6, plate association p-value <1E-7, and application of cluster-plot quality metrics as recommended by Affymetrix (i.e., Fisher’s Linear Discriminant <3.6, heterozygous cluster strength offset ≤0.1 or invalid homozygote ratio offset). For X-chromosomal SNPs, filter criteria proposed by König et al. [37]. were applied: p-value of exact test for Hardy–Weinberg equilibrium for women <1E−4, call rate <0.98, and minor allele frequency (MAF) <0.01 after setting all heterozygous calls in male samples to missing. A total of 546,429 SNPs passed all quality criteria. We imputed these data on the 1000 Genomes reference phase 1, release V3 of CEU. Annotation referred to HG19, dbSNP-build 135. We successfully mapped 538,181 measured SNPs to the reference. Data were pre-phased using SHAPEIT (version v2.r778) with standard settings for European populations. Imputation was conducted with IMPUTE2 (version 2.3.0) and resulted in 39,300,191 imputed SNPs. Post-imputation quality control required that a SNP has a MAF ≥ 1% and an Info-score ≥ 0.8. This resulted in 8,042,262 SNPs eligible for GWAS analyses.
GWAS analyses
Genome-wide association analyses were performed using PLINK 1.9. Separate analyses were carried out for the dependent variables mean vigilance, stability score, and slope index. We calculated additive models of gene-doses adjusting for sex, age, and the first three principal components. Top hits were defined by p-values < 1E-6. Genome-wide significance was set at p < 5E-8. To identify independent loci, lead-SNPs were pruned applying a linkage disequilibrium (LD) cut-off of R2 = 0.5. Further, for analyzing the congruence with eQTL hits, we screened 23 eQTL data sources with 67 listed eQTL studies [38, 39], providing information on whether a gene-locus regulates gene expression levels. Minimum LD between top hits and eQTL hits was set at R2 = 0.5. Furthermore, seeking to identify pleiotropic effects, top hits were compared with previous GWAS hits according to the recent NHGRI-EBI GWAS catalog (https://www.ebi.ac.uk/gwas/api/search/downloads/full, downloaded May 5, 2017). Minimum LD between top hits and GWAS hits was set at R2 = 0.5. Beyond this, we queried the results of all EEG-vigilance top hits in the publicly available PGC GWAS data files on major depressive disorder (MDD) [40], bipolar disorder (BD) [41], and attention deficit hyperactivity disorder (ADHD) [42]. We also investigated the enrichment of Reactome pathways among genes that are located within a 50 kb top hit distance or for which our top hit regions are replicated eQTLs (as suggested by the eQTL screening). Selected genes had to be listed in the Reactome database. In addition, we conducted a candidate marker analysis by selecting reported SNPs from the four GWAS on other resting EEG activity traits [29,30,31,32].
Gene-based analyses
For gene-based analyses, we utilized the approaches ‘PrediXcan’ (expression-based) [43] and ‘VEGAS2’ (locus-based) [44]. Analyses followed the guidelines, as described by the authors. For the former approach, we applied the DGN-model as it was trained by the authors on the largest sample (922 whole-blood samples from the Depression Genes and Networks, DGN). Only SNPs with MAF ≥0.01 and Info-score ≥0.8 were entered into analyses. For the latter approach, VEGAS2, we used standard settings including a margin of 50 kb for each gene and gene-locus assignment, as provided by the authors. Identified associating genes were validated by querying the MetaXcan online resource that contains association results derived from the GWAS summary statistics of over 100 complex traits [45].
Statistical power
Power analyses were conducted using r package pwr (version 1.2-1), with effect sizes quantified as ηp² (i.e., variance of the phenotype explained uniquely by the genetic variant in a multiple linear regression model adjusting for sex, age, and the first three principle components). Given N = 1,877 and α = 5E-8, power calculations revealed that SNP associations with a true effect size of ηp2 = 0.011, ηp2 = 0.016, and ηp2 = 0.021 were identified with a chance of 20%, 50%, and 80% (1−β), respectively. Supplementary Figure S1 shows the probabilities (1−β) of associations to surpass the threshold of genome-wide significance (α = 5E-8), given true effect sizes of up to ηp2 = 0.05.
Results
The descriptive statistics for VIGALL 2.1 variables of EEG-vigilance obtained from the present sample are shown in Table 1. Histograms and normal quantile plots for each EEG-vigilance variable are provided in Supplementary Figure S2.
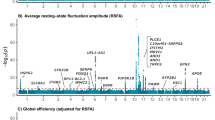
The distribution of p-values derived from GWAS analyses did not reveal evidence of general inflation of test-statistics (λ = 1.0042 for mean vigilance, λ = 0.9981 for the stability score, and λ = 0.9963 for the slope index), suggesting that association results are not biased by population stratification (for qq-plots see Supplementary Fig. S3). GWAS analyses revealed that no locus surpassed the threshold of genome-wide significance (p < 5E-8). However, SNPs in seven chromosomal regions were associated with at least one of the three EEG-vigilance variables at p < 1E-6 (considered as suggestive evidence). Figure 1 shows the Manhattan plot of results merged for the three outcome variables (for Manhattan plots of separate variables see Supplementary Fig. S4).
Manhattan plot showing the merged SNP associations for the three EEG-vigilance variables (best p-value for each SNP across mean vigilance, stability score, and slope index). The solid horizontal line indicates the genome-wide level of significance (5E-8). The dashed horizontal line indicates the level of suggestive evidence (1E-6)
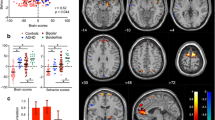
The strongest association was found for a region on chromosome 16 that tags DNAH3 and TMEM159 (lead-SNP: rs79472635, slope index: p = 5.49E-8), coding for dynein axonemal heavy chain 3 and transmembrane protein 159, respectively. Carriers of the minor allele exhibited relatively shallow declines of EEG-vigilance (see Fig. 2). With the slope index serving as the trait of interest, we obtained three further independent hits with p < 1E-6. One of them is located in SLC8A2 (rs830142, p = 3.13E-7), encoding solute carrier family 8 member A2. Another maps with SLC8A1 antisense RNA 1 (rs6544273, p = 7.95E-7), which is 120 kb upstream of the corresponding protein-coding SLC8A1 gene (encoding solute carrier family 8 member A1). In addition, we obtained rs13163913 (p = 6.38E-7) located 26 kb upstream of EEF1A1P20 (eukaryotic translation elongation factor 1 alpha 1 pseudogene 20). With the mean vigilance serving as the trait of interest, analyses revealed one hit in THSD7A (rs13247707, p = 1.05E-7), encoding thrombospondin type I domain containing 7A, and one further hit in HDAC9 (rs17432268, p = 9.37E-7), coding for histone deacetylase 9. For the stability score, we obtained rs11585842101 near RYBP (p = 1.87E-7), encoding RING1 and YY1 binding protein. Table 2 summarizes the lead-SNPs of our top hit loci. For regional plots, see Fig. 2 (DNAH3/TMEM159) and Supplementary Figure S5 (all other top hit loci). Detailed GWAS results are presented in Supplementary Table S1.
Association results for top hit rs79472635 located on Chromosome 16 in DNAH3 (near TMEM159). a Regional association plot with the slope index serving as dependent variable. b Simple moving averages of EEG-vigilance across the 20-min resting condition stratified by genotype groups of rs79472635. Stages of EEG-vigilance were determined using VIGALL 2.1
eQTL mapping
The screening of available eQTL data sources revealed that six out of seven lead-SNPs were either themselves reported to, or had a proxy SNP (R² ≥ .5) shown to be associated with, gene expression levels [38, 39]. Our strongest hit, rs79472635 located in DNAH3, has directly been associated with expression levels of TMEM159 (p = 1E-29). Direct evidence has also been reported for rs6544273 in SLC8A1-AS1 to contribute to variation in expression levels of SLC8A1 (p = 1.95E-4), for a link between rs830142 in SLC8A2 and expression levels of GLTSCR1 (p = 5.71E-7), for rs115842101 near RYBP and expression levels of both PPP4R2 (p = 3.82E-4) and RYBP (p = .002), and for rs13163913 near EEF1A1P20 and expression levels of FAM174A (p = 1.46E-51). All eQTL hits—including those referring to proxy SNPs—are provided in Supplementary Table S2.
GWAS catalog screening
In comparing our seven top hit regions with entries in the GWAS catalog, we identified no SNP that is in LD (R2 ≥ 0.5) with variants previously reported genome-wide significant for another trait. Supplementary Table S3 provides additional results of the GWAS catalog screening for SNPs with EEG-vigilance associations at p < 1E-5 and by applying a more liberal minimum LD (R2 = 0.2).
GWAS of the Psychiatric Genomics Consortium (PGC)
We queried the results of all EEG-vigilance top hits in the publicly available PGC GWAS data files on MDD [40], BD [41], and ADHD [42]. The mapping revealed that all corresponding top hits of our strongest locus (DNAH3/TMEM159) showed nominally significant associations with MDD (0.006 ≤ p ≤ 0.011). In this regard, carriers of alleles that we observed with less steep declines of EEG-vigilance (slope index) were reported to be at higher risk for major depression. Detailed PGC GWAS results of our top hits are provided in Supplementary Table S4.
Gene-based analyses
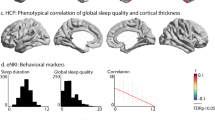
At the gene-level, the PrediXcan analysis revealed genome-wide significance for TMEM159 (slope index: p = 1.18E-6, Bonferroni-corrected p = 0.013). Higher expression levels of TMEM159 correlated with less steep declines of EEG-vigilance (see Fig. 3). Results of VEGAS2 were consistent with this finding, where this locus was labeled with central gene ZP2 and reached genome-wide significance (slope index: p = 2E-6, Bonferroni-corrected p = .048). As the latter analysis applied a margin of 50 kb around each gene, the first three hits—ZP2, DNAH3, and TMEM159—essentially describe the same locus on chromosome 16p12.3. We regard TMEM159 as the most relevant gene because the vast majority of eQTLs at the genetic site with the strongest SNP associations are related to the expression of TMEM159 (suppl. Fig. S6). Detailed gene-based results are provided in Supplementary Tables S5 and S6.
Gene-based association results of the expression-based approach ‘PrediXcan’. a Quantile-quantile-plots showing the expected vs. observed p-values for EEG-vigilance variable slope index. b Boxplot showing the observed scores of EEG-vigilance variable slope index as a function of imputed expression levels of TMEM159. Higher expression levels of TMEM159 correlated with less steep declines of resting state EEG-vigilance
We queried TMEM159 in the MetaXcan online resource [45]. Using the same statistical model (elastic net based on DGN whole-blood samples), expression levels of TMEM159 have been found with nominal significance in independent cohorts of patients with Major Depressive Disorder (p = 0.027, PGC), Autism Spectrum Disorder (p = 0.029, PGC), and Alzheimer’s Disease (p = 0.049, International Genomics of Alzheimer’s Project IGAP consortium) [46]. In this regard, higher expression levels of TMEM159 correlated with an increased risk for respective disorders. Supplementary Table S7 shows detailed MetaXcan association results for TMEM159.
Pathway analyses
Gene enrichment analyses revealed several pathways reaching the FDR-corrected level of significance. Based on the top hits in SLC8A1-AS1 and SLC8A2, we obtained evidence for pathways sodium/calcium exchangers (q = 8E-4), reduction of cytosolic Ca++ levels (q = 8E-4), and SLC-mediated transmembrane transport (q = 0.028). The above-mentioned pathways also surpassed the FDR-corrected level of significance when repeating analyses for genomic loci with EEG-vigilance associations at p < 1E-5. Detailed pathway results are shown in Supplementary Tables S8 and S9.
EEG-candidate marker analysis
From the four GWAS on other resting EEG activity traits [29,30,31,32], we derived 415 candidate SNPs in 272 loci previously reported with associations at p < 1E-4. Results from the candidate SNP analysis are shown in supplementary Table S10 (markers not included are listed in Table S11). We obtained 55 SNPs in 27 loci reaching nominal significance (p < 0.05) for at least one of our three EEG-vigilance variables. None of these associations remained significant after trait-wise correction for multiple testing (all q > 0.228). This was also the case after carrying out thinning (including only one SNP per locus that showed the strongest literature p-value) and repeating analyses in subgroups of EEG candidates (grouped according to their literature phenotypes alpha frequency, alpha power, beta power, delta power, theta power, total power; all q > 0.198). In addition, qq-plots did not reveal considerable evidence for an inflation of test statistics (suppl. Figure S7–9).
Discussion
The present GWAS is the first to identify genetic variants linked to brain arousal in the resting state. Based on a sample of N = 1877 subjects, we investigated the associations between 8,042,262 SNPs and EEG-vigilance during a 20-min eyes-closed resting condition. At the SNP-level, GWAS analyses revealed no locus with genome-wide significance (p < 5E-8), although seven loci were suggestive (p < 1E-6). Importantly, at the gene-level, GWAS analyses revealed significant evidence for TMEM159 (p = 0.013, Bonferroni-corrected). By mapping our SNPs to the GWAS results from the Psychiatric Genomics Consortium (PGC), we found that all corresponding markers of TMEM159 showed nominally significant associations with Major Depressive Disorder [40]. In line with this, the MetaXcan database suggests increased expression levels of TMEM159 in Major Depression, as well as Autism Spectrum Disorder, and Alzheimer’s Disease. Further, our pathway analyses indicate the potential involvement of sodium/calcium channel activity in arousal regulation.
Our strongest hit, rs79472635 (p = 5.49E-8), is an intronic variant located on chromosome 16 in DNAH3, encoding dynein axonemal heavy chain 3. Although DNAH3 is the nearest gene, we regard TMEM159 (encoding transmembrane protein 159) as the most relevant gene in this locus, because the vast majority of eQTLs at the genetic site are related to the expression of TMEM159. TMEM159 is also known as promethin and is expressed in various human organs including the brain, thyroid, and adrenal glands [47]. Importantly, variants of TMEM159 that we observed with persistently high arousal levels have previously been shown to increase the risk for major depression at nominal significance levels (PGC GWAS) [40]. This finding is in agreement with the postulations of the Arousal Regulation Model of Affective Disorders [7], according to which hyperstable levels of arousal contribute to the pathogenesis of depressive episodes. Indeed, depressive patients often experience insomnia, show delayed sleep onset, and benefit from sleep deprivation [48,49,50]. In addition, several studies have provided direct evidence for enhanced levels of brain arousal in depression [9,10,11,12]. Moreover, we observed that increased imputed expression levels of TMEM159 correlated with persistently high arousal levels, and the MetaXcan database suggests, by analogy, increased TMEM159 expression levels in independent cohorts of patients with Major Depression, Autism Spectrum Disorder, and Alzheimer’s Disease [45]. Therefore, TMEM159 appears to be an encouraging novel candidate gene that may modulate the risk for psychiatric disorders through arousal mechanisms.
Interestingly, TMEM159 was shown in lab experiments to interact with the transmembrane protein ‘flower’ (CACFD1) [51] that is involved in Ca2+ influx and resting Ca2+ levels [52]. Modulators of calcium metabolism may exert a variety of biological actions including effects on neurotransmission [53]. In fact, we obtained other hits that point towards a role of calcium metabolism in resting state arousal.
Two of the seven top hit loci tag genes that code for members of the solute carrier 8 (SLC8) family. One of them is a region on chromosome 19 within an intronic sequence of SLC8A2 (lead-SNP: rs830142, p = 3.13E-7). The other is located on chromosome 2 in an intronic sequence of SLC8A1 antisense RNA 1 (lead-SNP: rs6544273, p = 7.95E-7) and is associated with expression levels of the corresponding protein-coding SLC8A1 gene. Members of the SLC8 family are sodium-calcium exchanger proteins, which regulate cytoplasmic calcium concentrations by extruding calcium through the plasma membrane during relaxation of excitable cells. Their splice variants have been implicated in various phenomena including hippocampal long term potentiation, blood pressure regulation and hypertension, as well as cerebral ischemia [54].
Based on the independent hits in SLC8A1-AS1 and SLC8A2, our analyses revealed evidence for an involvement of sodium/calcium exchanger related pathways. In keeping with this, findings from animal experiments underscore the role of calcium channels in sleep-wake activity [55], vigilance state control [56], generation of sleep-spindles and delta oscillations [57, 58], and arousal events during sleep [59]. In a prior candidate gene study, initial evidence has been reported for an association of EEG-vigilance and a SNP within CACNA1C, encoding a subunit of a voltage-gated calcium channel (VGCC) [60]. Further, the overexpression of distinct VGCC subunits has been shown to promote neocortical hyperexcitability in mice [61]. In turn, alterations in arousal-phenotypes are well documented among psychiatric disorders and have been suggested to causally contribute to mental illness [7, 62]. Therefore, arousal could serve as a partial mediator between genetic calcium channel modulators and psychiatric phenotypes. In fact, an enrichment of genomic signals associated with calcium channel activity was demonstrated in Bipolar Disorder [41] and in a large-scale cross-disorder study of Major Depression, Bipolar Disorder, Attention Deficit Hyperactivity Disorder, Autism, and Schizophrenia [63]. Moreover, a role of ion-channelopathies in psychiatric disorders is in agreement with the pharmacodynamics of mood-stabilizing drugs such as valproate, lamotrigine, and carbamazepine. Thus, genetic modulators of ion-channel activity are encouraging candidates in both domains: arousal and psychiatric disorders. Future studies should further elaborate their possible causal relations.
From the four GWAS on other resting EEG activity traits [29,30,31,32], we derived 415 candidate SNPs and tested their potential association with EEG-vigilance. None of the obtained associations withstood multiple testing correction and the distribution of observed p-values was not different from a random distribution. Notably, previous studies focused on EEG power and peak frequencies during a recording period of less than 5 min. In contrast, we assessed the dynamics of EEG-vigilance by incorporating information on the spectral composition and cortical distribution of EEG activity across 20 min of rest. Therefore, VIGALL features adaptive procedures that account for the interindividual variability in total power and peak frequencies. Hence, we consider phenotypical difference as highly plausible reason for the lack of corroboration.
A common finding of GWAS is that, despite substantial heritability estimates from family and twin studies, single genetic variants exhibit only small associations. This general observation underscores the need for studies carried out in large research consortia with high statistical power. Because arousal regulation has been proposed to constitute a state-modulated trait [7], multicenter studies will require adequately standardized operating procedures (SOPs) to keep the proportion of variance caused by situational factors at a minimum. Daytime of assessment might be one of the most salient potential confounders to consider, although this variable did not decisively affect our own association results (see suppl. Figure S10). Notably, our analyses enable to deduce with a certainty of 99% (1−β) that none of the investigated SNPs account for >3% of the variance in EEG-vigilance as assessed using VIGALL (with α = 5E-8 and N = 1877). However, the power was only 20% to detect SNPs that explain 1.1% of the variance in EEG-vigilance. Hence, the sample size is a limitation of the present GWAS.
In the present study, we selected participants without a history of substance dependence or psychotic disorders, and who were free of current affective and anxiety disorders. Noteworthy, the prevalence of these disorders was low in the LIFE-Adult cohort. With our selection we aimed at improving the sample homogeneity. Nevertheless, this may have also led to reduced effect sizes of variants that impact psychiatric disorders through arousal mechanisms. On this account, it should be emphasized that the present effect size estimates refer to non-clinical populations.
In conclusion, the present GWAS is the first to address the dynamics of EEG-derived brain arousal in the resting state. We identified TMEM159 as a novel candidate gene which may modulate the risk for psychiatric disorders through arousal mechanisms. Additionally, two independent associations in SLC8A1-AS1 and SLC8A2 revealed evidence for an involvement of sodium/calcium exchanger related pathways, which is in line with previous findings on the interrelations of ion-channel activity, sleep-wake behavior, and psychiatric disorders. Future studies are needed to replicate our findings and to unravel the physiological mechanisms behind these associations.
References
Pfaff D, Ribeiro A, Matthews J, Kow L-M. Concepts and mechanisms of generalized central nervous system arousal. Ann N Y Acad Sci. 2008;1129:11–25.
De Valck E, Cluydts R. Sleepiness as a state-trait phenomenon, comprising both a sleep drive and a wake drive. Med Hypotheses. 2003;60:509–12.
Luyster FS, Strollo PJ Jr, Zee PC, Walsh JK. Sleep: a health imperative. Sleep. 2012;35:727–34.
Wing YK, Zhang J, Lam SP, Li SX, Tang NL, Lai KY, et al. Familial aggregation and heritability of insomnia in a community-based study. Sleep Med. 2012;13:985–90.
Spada J, Scholz M, Kirsten H, Hensch T, Horn K, Jawinski P et al. Genome-wide association analysis of actigraphic sleep phenotypes in the LIFE Adult Study. J Sleep Res. 2016;25:690–701.
Lane JM, Liang J, Vlasac I, Anderson SG, Bechtold DA, Bowden J, et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat Genet. 2017;49:274–81.
Hegerl U, Hensch T. The vigilance regulation model of affective disorders and ADHD. Neurosci Biobehav Rev. 2014;44:45–57.
Geissler J, Romanos M, Hegerl U, Hensch T. Hyperactivity and sensation seeking as autoregulatory attempts to stabilize brain arousal in ADHD and mania? Atten deficit Hyperact Disord. 2014;6:159–73.
Schmidt FM, Pschiebl A, Sander C, Kirkby KC, Thormann J, Minkwitz J, et al. Impact of serum cytokine levels on EEG-measured arousal regulation in patients with major depressive disorder and healthy controls. Neuropsychobiology. 2016;73:1–9.
Wittekind DA, Spada J, Gross A, Hensch T, Jawinski P, Ulke C, et al. Early report on brain arousal regulation in manic vs depressive episodes in bipolar disorder. Bipolar Disord. 2016;18:502–10.
Ulke C, Sander C, Jawinski P, Mauche N, Huang J, Spada J et al. Sleep disturbances and upregulation of brain arousal during daytime in depressed versus non-depressed elderly subjects. World J Biol Psychiatry Off J World Federation of Soc Biol Psychiatry. 2017;18:633–40.
Schmidt FM, Sander C, Dietz M-E, Nowak C, Schroder T, Mergl R, et al. Brain arousal regulation as response predictor for antidepressant therapy in major depression. Sci Rep. 2017;7:45187.
Eysenck H-J. The biological basis of personality. American lecture series. 689. Springfield, IlI: Thomas; 1967.
Zuckerman M. Sensation seeking: Beyond the optimal level of arousal. L. Erlbaum Associates; Distributed by the Halsted Press Division of Wiley: Hillsdale, N.J., New York; 1979.
Cuthbert BN, Insel TR. Toward the future of psychiatric diagnosis: the seven pillars of RDoC. BMC Med. 2013;11:126.
De Gennaro L, Ferrara M, Curcio G, Cristiani R. Antero-posterior EEG changes during the wakefulness-sleep transition. Clin Neurophysiol Off J Int Fed Clin Neurophysiol. 2001;112:1901–11.
Tsuno N, Shigeta M, Hyoki K, Kinoshita T, Ushijima S, Faber PL, et al. Spatial organization of EEG activity from alertness to sleep stage 2 in old and younger subjects. J Sleep Res. 2002;11:43–51.
Strijkstra AM, Beersma, Domien GM, Drayer B, Halbesma N, Daan S. Subjective sleepiness correlates negatively with global alpha (8–12 Hz) and positively with central frontal theta (4–8 Hz) frequencies in the human resting awake electroencephalogram. Neurosci Lett. 2003;340:17–20.
Corsi-Cabrera M, Guevara MA, Del Río-Portilla Y, Arce C, Villanueva-Hernández Y. EEG bands during wakefulness, slow-wave and paradoxical sleep as a result of principal component analysis in man. Sleep. 2000;23:738–44.
Hegerl U, Sander C, Ulke C, Böttger D, Hensch T, Huang J et al. Vigilance Algorithm Leipzig (VIGALL) Version 2.1 Manual. 2016. https://research.uni-leipzig.de/vigall/ Accessed 12 Oct 2017.
Sander C, Hensch T, Wittekind DA, Böttger D, Hegerl U. Assessment of wakefulness and brain arousal regulation in psychiatric research. Neuropsychobiology. 2015;72:195–205.
Guenther T, Schönknecht P, Becker G, Olbrich S, Sander C, Hesse S, et al. Impact of EEG-vigilance on brain glucose uptake measured with [(18)F]FDG and PET in patients with depressive episode or mild cognitive impairment. Neuroimage. 2011;56:93–101.
Olbrich S, Mulert C, Karch S, Trenner M, Leicht G, Pogarell O, et al. EEG-vigilance and BOLD effect during simultaneous EEG/fMRI measurement. Neuroimage. 2009;45:319–32.
Huang J, Hensch T, Ulke C, Sander C, Spada J, Jawinski P, et al. Evoked potentials and behavioral performance during different states of brain arousal. BMC Neurosci. 2017;18:21.
Olbrich S, Sander C, Matschinger H, Mergl R, Trenner M, Schönknecht P, et al. Brain and body: associations between EEG-vigilance and the autonomous nervous system activity during rest. J Psychophysiol. 2011;25:190–200.
Jawinski P, Kittel J, Sander C, Huang J, Spada J, Ulke C et al. Recorded and reported sleepiness: the association between brain arousal in resting state and subjective daytime sleepiness. Sleep. 2017;40:zsx099.
Lee SH, Ripke S, Neale BM, Faraone SV, Purcell SM, Perlis RH, et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet. 2013;45:984–94.
Pfaff DW. Brain arousal and information theory: neural and genetic mechanisms. Cambridge, Mass: Harvard University Press; 2006.
Malone SM, Burwell SJ, Vaidyanathan U, Miller MB, McGue M, Iacono WG. Heritability and molecular-genetic basis of resting EEG activity: a genome-wide association study. Psychophysiology. 2014;51:1225–45.
Meyers JL, Zhang J, Wang JC, Su J, Kuo SI, Kapoor M et al. An endophenotype approach to the genetics of alcohol dependence: a genome wide association study of fast beta EEG in families of African ancestry. Mol Psychiatry. 2017;22:1767–75.
Meyers JL, Zhang J, Manz N, Rangaswamy M, Kamarajan C, Wetherill L et al. A genome wide association study of fast beta EEG in families of European ancestry. Int J Psychophysiol Off J Int Organ Psychophysiol. 2017;115:74–85.
Hodgkinson CA, Enoch M-A, Srivastava V, Cummins-Oman JS, Ferrier C, Iarikova P et al. Genome-wide association identifies candidate genes that influence the human electroencephalogram. Proc Natl Acad Sci USA. 2010; 107: 8695–8700.
Loeffler M, Engel C, Ahnert P, Alfermann D, Arelin K, Baber R, et al. The LIFE-Adult-Study: objectives and design of a population-based cohort study with 10,000 deeply phenotyped adults in Germany. BMC Public Health. 2015;15:691.
Huang J, Sander C, Jawinski P, Ulke C, Spada J, Hegerl U, et al. Test-retest reliability of brain arousal regulation as assessed with VIGALL 2.0. Neuropsychiatr Electrophysiol. 2015;1:263.
Jawinski P, Mauche N, Ulke C, Huang J, Spada J, Enzenbach C, et al. Tobacco use is associated with reduced amplitude and intensity dependence of the cortical auditory evoked N1-P2 component. Psychopharmacology. 2016;233:2173–83.
Jawinski P, Tegelkamp S, Sander C, Häntzsch M, Huang J, Mauche N et al. Time to wake up: No impact of COMT Val158Met gene variation on circadian preferences, arousal regulation and sleep. Chronobiol Int. 2016;33:893–905.
König IR, Loley C, Erdmann J, Ziegler A. How to include chromosome X in your genome-wide association study. Genet Epidemiol. 2014;38:97–103.
Kirsten H, Al-Hasani H, Holdt L, Gross A, Beutner F, Krohn K, et al. Dissecting the genetics of the human transcriptome identifies novel trait-related trans-eQTLs and corroborates the regulatory relevance of non-protein coding loci†. Hum Mol Genet. 2015;24:4746–63.
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science. 2015;348:648–60.
Ripke S, Wray NR, Lewis CM, Hamilton SP, Weissman MM, Breen G, et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol Psychiatry. 2013;18:497–511.
Sklar P, Ripke S, Scott LJ, Andreassen OA, Cichon S, Craddock N, et al. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet. 2011;43:977–83.
Demontis D, Walters RK, Martin J, Mattheisen M, Als TD, Agerbo E et al. Discovery of the First Genome-Wide Significant Risk Loci For ADHD. 2017.
Gamazon ER, Wheeler HE, Shah KP, Mozaffari SV, Aquino-Michaels K, Carroll RJ, et al. A gene-based association method for mapping traits using reference transcriptome data. Nat Genet. 2015;47:1091–8.
Mishra A, Macgregor S. VEGAS2: software for more flexible gene-based testing. Twin Res Human Genet Off J Int Soc Twin Stud. 2015;18:86–91.
Barbeira AN, Dickinson SP, Torres JM, Bonazzola R, Zheng J, Torstenson ES et al. Exploring the Phenotypic Consequences of Tissue Specific Gene Expression Variation Inferred from GWAS Summary Statistics. 2016.
Lambert JC, Ibrahim-Verbaas CA, Harold D, Naj AC, Sims R, Bellenguez C, et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat Genet. 2013;45:1452–8.
Fagerberg L, Hallström BM, Oksvold P, Kampf C, Djureinovic D, Odeberg J, et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteom. 2014;13:397–406.
Baglioni C, Battagliese G, Feige B, Spiegelhalder K, Nissen C, Voderholzer U, et al. Insomnia as a predictor of depression: A meta-analytic evaluation of longitudinal epidemiological studies. J Affect Disord. 2011;135:10–19.
Armitage R. Sleep and circadian rhythms in mood disorders. Acta Psychiatr Scand Suppl. 2007;115:104–15.
Giedke H, Schwärzler F. Therapeutic use of sleep deprivation in depression. Sleep Med Rev. 2002;6:361–77.
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, et al. STRINGv10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015;43:D447–52.
Yao C-K, Lin YQ, Ly CV, Ohyama T, Haueter CM, Moiseenkova-Bell VY, et al. A synaptic vesicle-associated Ca2+ channel promotes endocytosis and couples exocytosis to endocytosis. Cell. 2009;138:947–60.
Berridge MJ, Lipp P, Bootman MD. The versatility and universality of calcium signalling. Nat Rev Mol Cell Biol. 2000;1:11–21.
Khananshvili D. The SLC8 gene family of sodium-calcium exchangers (NCX) - structure, function, and regulation in health and disease. Mol Asp Med. 2013;34:220–35.
Kraus RL, Li Y, Gregan Y, Gotter AL, Uebele VN, Fox SV, et al. In vitro characterization of T-type calcium channel antagonist TTA-A2 and in vivo effects on arousal in mice. J Pharmacol Exp Ther. 2010;335:409–17.
Beuckmann CT, Sinton CM, Miyamoto N, Ino M, Yanagisawa M. N-type calcium channel alpha1B subunit (Cav2.2) knock-out mice display hyperactivity and vigilance state differences. J Neurosci Off J Soc Neurosci. 2003;23:6793–7.
Astori S, Wimmer RD, Prosser HM, Corti C, Corsi M, Liaudet N et al. The Ca(V)3.3 calcium channel is the major sleep spindle pacemaker in thalamus. Proc Natl Acad Sci USA. 2011; 108: 13823–8.
Lee J, Shin H-S. T-type calcium channels and thalamocortical rhythms in sleep: a perspective from studies of T-type calcium channel knockout mice. CNSNDDT. 2007;6:63–70.
Anderson MP, Mochizuki T, Xie J, Fischler W, Manger JP, Talley EM et al. Thalamic Cav3.1 T-type Ca2+ channel plays a crucial role in stabilizing sleep. Proc Natl Acad Sci USA. 2005; 102: 1743–8.
Jawinski P, Sander C, Mauche N, Spada J, Huang J, Schmidt A, et al. Brain arousal regulation in carriers of bipolar disorder risk alleles. Neuropsychobiology. 2015;72:65–73.
Faria LC, Gu F, Parada I, Barres B, Luo ZD, Prince DA. Epileptiform activity and behavioral arrests in mice overexpressing the calcium channel subunit alpha2delta-1. Neurobiol Dis. 2017;102:70–80.
Krystal AD. Psychiatric disorders and sleep. Neurol Clin. 2012;30:1389–413.
Smoller JW, Ripke S, Lee PH, Neale B, Nurnberger JI, Santangelo S, et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet. 2013;381:1371–9.
Acknowledgements
This publication is supported by LIFE—Leipzig Research Center for Civilization Diseases, Universität Leipzig. LIFE is funded by means of the European Union, by the European Regional Development Fund (ERDF) and by means of the Free State of Saxony within the framework of the excellence initiative. This publication was written within the framework of the cooperation between the German Depression Foundation and the ‘Deutsche Bahn Stiftung gGmbH’. GWAS analysis was supported by the Federal Ministry of Education and Research (BMBF), project Grant No. FKZ 031L0026 (LHA). We thank Elise Paul for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jawinski, P., Kirsten, H., Sander, C. et al. Human brain arousal in the resting state: a genome-wide association study. Mol Psychiatry 24, 1599–1609 (2019). https://doi.org/10.1038/s41380-018-0052-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-018-0052-2
This article is cited by
-
Generalized and social anxiety disorder interactomes show distinctive overlaps with striosome and matrix interactomes
Scientific Reports (2021)
-
Fatigue and brain arousal in patients with major depressive disorder
European Archives of Psychiatry and Clinical Neuroscience (2021)
-
Effects and feasibility of hyperthermic baths in comparison to exercise as add-on treatment to usual care in depression: a randomised, controlled pilot study
BMC Psychiatry (2020)
-
Vulnerability to bipolar disorder is linked to sleep and sleepiness
Translational Psychiatry (2019)
-
Impact of brain arousal and time-on-task on autonomic nervous system activity in the wake-sleep transition
BMC Neuroscience (2018)