Abstract
Growth rate is a widely studied parameter for various cell-based biological studies. Growth rates of cell populations can be monitored in chemostats and micro-chemostats, where nutrients are continuously replenished. Here, we present an integrated microfluidic platform that enables long-term culturing of non-adherent cells as well as parallel and mutually independent continuous monitoring of (i) growth rates of cells by means of impedance measurements and of (ii) specific other cellular events by means of high-resolution optical or fluorescence microscopy. Yeast colonies were grown in a monolayer under culturing pads, which enabled high-resolution microscopy, as all cells were in the same focal plane. Upon cell growth and division, cells leaving the culturing area passed over a pair of electrodes and were counted through impedance measurements. The impedance data could then be used to directly determine the growth rates of the cells in the culturing area. The integration of multiple culturing chambers with sensing electrodes enabled multiplexed long-term monitoring of growth rates of different yeast strains in parallel. As a demonstration, we modulated the growth rates of engineered yeast strains using calcium. The results indicated that impedance measurements provide a label-free readout method to continuously monitor the changes in the growth rates of the cells without compromising high-resolution optical imaging of single cells.
Similar content being viewed by others
Introduction
Cells regulate their growth rate in response to external signals, and as cells grow, their metabolism, macromolecular synthesis, and the processes included in cell division must be coordinated1,2,3,4. This coordination of different processes, the way in which cells monitor their nutritional environment, how they integrate this information into the cell cycle, how they regulate their cell cycle, as well as whether and how these regulatory processes change during a cellular life cycle still include many open issues5,6,7. The investigation of these open issues requires a well-developed and broadly understood model system, such as budding and fission yeast8,9, and an experimental setup that can be used to perform such investigations. The chemostat provides a powerful method to systematically study the coupling between growth rates and cellular processes: it allows for experimentally controlling the growth rate of a cell population by adjusting the nutrient supply into a defined culture vessel volume, thereby providing a stable and defined environment for cells10. In a chemostat, the growth kinetics, i.e., the relation between cell growth rate and substrate consumption, is controlled by manipulating the medium addition to the culture vessel.
Micro-chemostats rely on microfluidics technology for culturing cells in a constant and defined environment under continuous perfusion. The cells in these devices grow in chambers or channels of defined size, and their growth rates are usually determined by using microscopy11,12,13,14,15. In contrast to conventional chemostats, the growth rates in these microfluidic platforms are defined by the composition of the supplied media. An advantage of microfluidic devices is that they do allow for monitoring of individual cells over an extended period of time. However, associated growth rate measurements are often limited by the field of view or the overall size of the culture chamber or pad and require dedicated software for cell segmentation and tracking. Detailed cell tracking requires high-temporal-resolution optical measurements, which limits the number of positions that can be imaged by the microscope in a single experiment due to the required stage movements. The limited number of imaging positions considerably reduces the throughput and detracts from the possibility to parallelize experiments under similar or identical conditions. Additionally, the use of fluorescence microscopy for measuring cell growth rates limits the number of fluorophores that are available for tracking other specific events and processes in the cells. Moreover, phototoxic effects may be induced upon frequent imaging16 so that additional control experiments become necessary to assess such phototoxicity effects, which are tedious to perform. Phototoxicity effects can be obviated by the use of label free techniques, such as measuring the optical density of the cell solution in microfluidic platforms17,18. Unfortunately, suitable devices are not amenable to high-resolution optical imaging and to obtaining information at single-cell resolution.
Electrical impedance spectroscopy (EIS) is a label free, non-invasive method for cell or particle counting and analysis19,20,21,22. Impedance cytometers, microfluidic devices with impedance measurement features offer the capability to characterize and analyze cell populations without the need for fluorescent labels23,24,25,26. A common implementation of microfluidic impedance platforms consists of simple microfluidic channels with single or multiple facing electrodes to perform the impedance measurements. Most of these flow-through platforms are stand-alone devices that can be used downstream of cell culture reactors or with cell suspensions, and are not easy to parallelize. Growth rate measurements in cell cultures using electrical cell-substrate impedance sensing (ECIS) were demonstrated for adherent cells27,28. Impedance-based measurements of viable biomass in microtiter plates were also performed for non-adherent cells29,30. However, to the best of our knowledge, there is currently no integrated platform, which features continuous and parallel execution of (i) cell culturing under controlled perfusion conditions, (ii) monitoring of the cell growth rate, and (iii) high-resolution imaging of non-adherent or suspended cells in the same single device.
Therefore, our goal was to develop a platform that allows for long-term culturing as well as continuous parallel and mutually independent monitoring of (i) growth rates by means of impedance measurements and of (ii) specific other cellular events by means of high-resolution optical or fluorescence microscopy. To this end, we used the concept of a previously developed microfluidic device to culture yeast cells in a defined volume underneath pads in a defined environment under continuous perfusion31. We then developed a new device, which, besides some other features, included an impedance readout to count the number of cells leaving culturing areas under the pads. This new microfluidic platform enables continuous impedance-based monitoring of the growth rates of populations of non-adherent cells, such as yeast cells, while it enables simultaneous long-term, high-resolution imaging of cellular events. Combined microscopy and impedance measurements can be conducted in an array-based format, as the device features multiple culturing chambers and sets of sensing electrodes. Multiple yeast cell colonies in different defined culturing environments can be analyzed in parallel. The setup is straightforward and simple to use on an automated microscope including all electrical connections for impedance measurements. Parallel growth rate measurements of different engineered yeast strains over 2 days have been conducted with the device. The growth rates of these strains were modulated by switching between standard medium and calcium-rich medium. The changes in the obtained growth rates were determined by impedance measurements and were similar to those observed by using standard culturing methods. In contrast to microscopy, impedance assessment enables high-throughput monitoring of growth rates without imposing additional strain on cells as a consequence of light exposure. The developed platform enables the user to employ the full repertoire of microscopy methods for tracking a wide range of cellular events, while the growth rate of the cells is continuously and independently monitored through an impedance readout.
Materials and methods
Device fabrication
The device consists of two parts: a poly(dimethylsiloxane) (PDMS) layer containing the microfluidic structures, and a glass slide with a patterned metal layer. The PDMS layer was cast from a mold, fabricated in a three-layer microfabrication process on a 4-inch silicon wafer (Figure S 1a). The first layer for the clamping pads was obtained through selective ion-beam etching of a silicon substrate using a photoresist mask. This process allows for precise adjustment of the height of the first layer, which defines the clamping gap below the pads to be 4 ± 0.3 µm. The next two layers for the cell culture chambers and for the microfluidic channel structures were fabricated by two SU-8 photolithography steps. SU-8 25 and SU-8 100 (MicroChem Co., USA) were sequentially spin-coated to obtain layers of 20 and 200 µm height; the layers were then exposed to UV light through transparency masks for cross-linking. The mold was silanized for 2 h with trichloro(1H,1H,2H,2H-perfluoro-octyl)silane (Sigma-Aldrich, Switzerland) in a vacuum desiccator. A mixture of silicone and curing agent (10:1 w/w, Sylgard® 184, Dow Corning, Germany) was poured on the SU-8 mold and cured for 2 h at 80 °C. The PDMS was then peeled off the mold and cut into single chips. Holes were punched for the device inlets and outlets.
The electrodes were fabricated on 4-inch 500-µm- or 200-µm-thick glass wafers using a lift-off process (Figure S 1b). The wafer was spin-coated with lift-off resist (LOR3B, Microchem Corp., Newton, USA), followed by a positive photoresist (S1813, Rohm-Haas, Schwalbach, Germany) and patterned using photolithography. A sputtering process was applied to deposit a platinum film of 200 nm thickness on top of a 20-nm-thick W/Ti adhesion layer. Metal lift-off was carried out using mr-Rem 400 remover (Micro Resist Technology GmbH, Berlin, Germany). A 500-nm passivation layer of silicon nitride was then deposited on the fabricated metal layer using plasma-enhanced chemical vapor deposition. The passivation layer was removed in the areas of electrodes and contacts pads (using a S1813 positive resist mask) by means of reactive-ion etching. After fabrication, the glass wafer was diced into individual glass slides (70 mm × 32 mm).
Cell loading and device assembly
The cell loading procedure is schematically shown in Figure S 2. The assembly and precise alignment of the structures in the PDMS layer and the electrodes on the glass slide was carried out during the cell loading procedure with a custom-made alignment tool (Figure S 2a). The PDMS layer and glass slide were placed on the top and bottom holder of the alignment tool. The glass slide was brought into close proximity to the PDMS layer without touching it. The PDMS structures were aligned with the electrodes on the glass (X, Y, and theta) by using micrometer screws and a hand-held microscope (Dino-Lite digital microscope, Netherlands). Once aligned, the top holder with the PDMS layer was removed and placed on the table with the culture chambers facing upward; 0.5 µl of cell suspension at a concentration of 3 × 107 cells/ml was pipetted into each chamber. As PDMS is hydrophobic in nature, the liquid did not spread. The top holder was again placed on the alignment tool. The four cone-shaped pins ensured that the top holder re-centered at the same position as aligned before removal. The glass slide was then moved upward and brought in contact with the PDMS layer. The assembled device was afterward removed from the alignment tool.
Setup
Impedance measurements
The experimental setup is shown in Figure S 3a. After cell loading, the device was placed in a custom-made PCB (115 mm × 75 mm), which was used to switch between the sensing electrodes of different analysis units in an automated way (Figure S 3b). The PCB was connected to the impedance spectroscope (HF2, Zurich Instruments AG, Switzerland) and to a custom-made, automated, multiplexed EIS (AMEIS)32 controller board, which controlled the digital signal to switch between analysis units and provided the interface to a PC. Custom-made software was used to select the analysis units and to program the switching protocol and recording duration. The sensing electrodes of selected analysis units were connected to the impedance spectroscope for defined recording durations. Each analysis unit included two electrodes—a stimulating and a recording electrode. An AC signal with an amplitude of 2 V and frequencies of 1.12 and 1.5 MHz was applied to the stimulating electrode. At the recording electrode, the current was transformed into a voltage using a trans-impedance amplifier (HF2TA, Zurich Instruments AG, Switzerland), which was then measured using the impedance spectroscope. The data was stored on the PC for later analysis. The phase signal of the output voltage was recorded and analyzed. The obtained data were bandpass-filtered using MATLAB (The MathWorks Inc., USA) using a frequency range of 0.1–30 Hz. The peaks were extracted from the filtered data by applying a threshold to minimize the number of false positives and false negatives. The threshold values varied, depending on the medium conditions and frequency used for recording. The threshold values ranged between 2 × 10−3–6 × 10−3 degrees for 1.5 MHz and 0.5 × 10−3–0.8 × 10−3 degrees for 1.12 MHz.
Microscopy
The PCB hosting the device was placed in a custom-made holder, which fits onto the stage of an automated, inverted microscope (Figure S 3b). The images were obtained using inverted microscopes (Olympus IX 81 and Nikon Ti Eclipse microscope) placed in an environmental control box, which maintained a stable temperature of 30 °C. Fluorescence images were captured on the Nikon microscope using a Nikon Plan Fluor 40× objective (NA 0.75, WD 0.66). The microscope was controlled using Youscope28, and offline image analysis was performed using ImageJ33 and CellX34. Syringe pumps (neMESYS, Cetoni GmbH, Germany) were connected to the microfluidic chip for media supply (flow rate 10 µl/min, 15 ml of medium is required for a 24-h experiment) and were controlled using Youscope.
Experiments to assess the medium exchange characteristics (Figure S 4) were carried out on the Olympus microscope, and images were taken every second with a CMOS camera (Hamamatsu ORCA Flash 4.0 camera). Syringes were filled with de-ionized water and a solution of Amaranth (4 mg/ml, Sigma-Aldrich, Switzerland) in de-ionized water. Switching between the different syringes was controlled using Youscope. The calculation of the average intensity of the images and further analysis was performed using ImageJ.
Cell culture
Saccharomyces cerevisiae strains—vph1∆ (BY4741, Euroscarf, FRY2033) and wild-type strain (BY4700, FRY1398) were used for the experiments to measure growth rate dynamics in different medium conditions. These strains were cultured overnight in shaking flasks with Yeast extract Peptone Dextrose (YPD) containing 1% yeast extract (BD Biosciences, Germany), 2% peptone (BD Biosciences), and 2% glucose. The overnight culture was diluted and grown for a few hours in YPD to have the cells in the exponential phase before loading them into the device. Cells were grown in YPD with 100 mM CaCl2 to observe changes in the growth rate. Yeast strains carrying Vph1-Citrine::KanMX (FRY 2011), Whi5-Citrine::KanMX (FRY 1935), and Cdc12-Citrine::KanMX (FRY 2145) fluorescent fusion protein constructs were used to perform fluorescence microscopy. These strains were tagged with Citrine fluorophore but at different locations in the cell and cultured in synthetic defined (SD) media.
Growth rate measurements in a plate reader (Infinite M200 PRO, Tecan Group Ltd., Switzerland) were carried out for vph1∆ and wild-type strains in YPD and YPD with 100 mM CaCl2. The absorbance of the cells was recorded at 600 nm every 6 min over a period of 72 h at 30 °C. The growth rate of the cells was measured as an increase in the absorbance over time, and was extracted using MATLAB.
Results and discussion
Device design and function
The device consists of two parts, the PDMS layer containing the microfluidic structures and the glass slide patterned with co-planar platinum electrodes (Fig. 1a). The central part of the assembled device is schematically shown in Fig. 1b. Up to four different medium solutions can be infused through the four inlets and can be mixed in the subsequent meandered channel structure35,36,37(a feature, which has not been used in this paper) to study the effects of medium variation on the cell growth rate. The medium channels around the chamber are 1 mm wide and 200 µm high. The medium is then guided through the three microfluidic cell culture chambers with identical flow rates, as indicated by the white arrows, and is collected at the outlet. Two flow resistors (30 µm wide) are connected in parallel to the culturing chambers. They ensure a controlled, parallel, and unidirectional flow through the chambers and ensure rapid nutrient availability as well as continuous cell removal. The three chambers (4.5 mm long, 5 mm wide, and 20 µm high) allow for simultaneous culturing of three different strains under identical medium conditions.
Photographs of the PDMS layer and the glass slide with the electrodes. For device assembly, the PDMS layer was aligned with and sealed onto a glass slide. The glass appears yellowish due to the silicon nitride insulation layer. b Schematic illustration of the central part of the device consisting of three chambers, each having five analysis units. The 15 analysis units can be used with automated microscopy and feature integrated impedance readout. The channel for the medium is depicted in dark blue with white arrows indicating the flow direction. The vacuum channel to seal the PDMS structure against the glass and to remove bubbles in the liquid is shown in red. c Enlarged sketch of one analysis unit. Yeast cells are clamped under four PDMS pads and proliferate under constant media perfusion. The blue arrow indicates the flow direction of the medium, and black arrows illustrate the cell trajectories. Cross-section AAʹ illustrates that the clamping of yeast cells is between the square PDMS pads and the glass substrate. Cross-section BBʹ shows cells passing through the electric field lines of the co-planar electrodes for impedance measurements
Each chamber has five identical analysis units (Fig. 1c). An analysis unit includes four clamping pads (150 µm in diameter), two hexagonal support and guiding pillars, and one pair of sensing electrodes, placed between the guiding pillars downstream of the clamping pads. The vertical distance between the clamping pads and the glass is 4 ± 0.3 µm, which is sufficient for growing yeast cells in a monolayer under flow conditions. Once the space under the pads is completely filled, cells start to outgrow the pads. These cells are entrained in the flow around the pads and guided over the electrodes for impedance measurement. As the number of cells under the pad remains constant, the number of washed-out cells per time can be used to measure the growth rate of the cells in the chip by using impedance. The sensing electrodes are 300 µm in length, 20 µm in width, 20 µm apart, and located between two hexagonal pillar structures that focus the medium flow. The hexagonal structures also provide support to the chamber ceiling and prevent collapsing. Once the cells have passed the electrodes, they leave the chamber and are removed through the outlet. This prevents clogging of the device and allows for long-term culture.
The device accommodates a total of 15 electrode pairs, five replicates in three chambers, which are all routed to the connection pads along the side of the glass slide. The glass slide is plugged into a PCB that allows for automated sequential impedance recording through the different electrode pairs. The device with the PCB is placed in a custom-made holder mounted on a microscope stage.
Cell culturing and cell flow in the microfluidic device
A flow diagram of a standard experiment is shown in Fig. 2a. After loading, the cells are allowed to grow under the pads and impedance measurements of the outgrowing cells passing the electrodes are possible once the pads are filled. The setup allows for simultaneous optical imaging of the cells under the pads throughout the duration of the experiment.
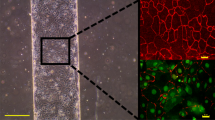
Flow diagram of an experiment: cells were loaded into the device and allowed to grow under the pads (b). Time t = 0 defines the beginning of the experiment. From time t = tEIS simultaneous impedance measurements of the outgrowing cells (c) and imaging of the filled pads by means of microscopy is possible. b Bright-field images of the cells growing between the pad and the 500-µm-thick glass slide that form a 2D monolayer. c Bright-field image of four filled clamping pads of an analysis unit and flow paths of cells leaving the pads as extracted from a video. The obtained cell traces indicate that all cells passed between the hexagonal guiding structures. The dotted line indicates the boundaries of the region, within which the cells were guided through the hexagonal structures and passed over the electrodes
Cells were loaded by pipetting the selected cell suspension (0.5 µl) into each chamber of the PDMS layer, which were then sealed with a glass slide hosting the electrodes by applying vacuum through the vacuum line. During sealing, the electrodes have to be precisely aligned with respect to the hexagonal structures in the PDMS layer. The alignment and sealing has to be quick, so that evaporation of the small volume of the cell suspension is minimized. To this end, an alignment machine that enables a simple and quick loading procedure was developed in-house (Figure S 2). The loaded volume is low enough to fill only the cell culture chamber; neither overflows of the cell suspension into neighboring chambers nor cross contamination were observed.
After sealing, the PDMS layer and glass slide were held together only through vacuum, applied to the vacuum channel that surrounds the medium microchannel inside the PDMS layer (red feature in Fig. 1b). This feature is different from what has been previously presented by Frey et al.31 and enables a comfortable use of the device also in laboratories lacking plasma bonding equipment. The device can be perfused for the duration of the experiment at flow rates up to 500 µl/min without leakage. The non-permanent bonding between PDMS and glass enables the facile re-use of both parts of the device. As there is a substantial risk of compound carry-over due to the absorbing nature of PDMS, only the glass slide was reused. The second function of the vacuum channel was the removal of any bubbles formed in the device after loading or during the experiment, thus ensuring robust long-term cell culturing under constant medium perfusion.
After loading, only cells below the pads were clamped. All unclamped cells outside the pads were washed away by the medium flow. The cells clamped between the PDMS and glass slide grew only in a two-dimensional (2D) plane forming a monolayer of cells, as shown in time-lapse images of one of the pads in Fig. 2b. Cell counting can be carried out by using bright-field imaging with 500-µm-thick glass slides. Clamping was found to cause no stress for the cells, as was shown previously using a Msn2 reporter31.
A key issue in designing the microfluidic device was to ensure that all the cells growing out of the clamping pads were guided over the sensing electrodes. Figure 2c shows a picture of the analysis units with the four pads, placed in a diamond-like configuration, and the hexagonal pillar structures further downstream. Loading of the chip leads to stochastic distribution of cells under the pads. The arrangement of the four pads in each analysis unit and their geometric area (150 µm side length) provides a sizeable number of clamped cells per analysis unit. The diamond-like shape of the pad minimizes the area of stagnant flow and prevents cell accumulation. The cell occupancy of the pads and the flow paths of the cells leaving the pads in Fig. 2c were extracted from a video. The hexagonal structures were designed to act as a funnel for collecting all cells growing out of the pads (dotted line in Fig. 2c). A finite-element model of the analysis unit evidences that the flow lines around the culturing pads pass through the hexagonal structures (Figure S 4c). It also shows the fluid velocity around the analysis unit (Figure S 4c). At the same time, the channel between the two hexagonal structures was designed sufficiently large so that all cells pass over the pair of electrodes due to the laminar-flow conditions in the chamber. A narrower channel would have entailed the risk of either clogging or of losing cells that pass outside the channel owing to the increased fluidic resistance. The five analysis units are arranged at sufficient distance within the culture chamber to completely obviate cross-talk under the given laminar-flow regime.
Impedance measurements of cells
The electrodes downstream of the clamping pads were used to perform impedance-based counting of the cells coming from the pads. An AC voltage signal was applied to the stimulating electrode, and the current flowing through the system was converted into a voltage by using a trans-impedance amplifier. When a cell moved over the electrodes, it caused a transient variation of the intra-electrode impedance, which, in turn, produced a transient change in the recorded current and, hence, the output voltage38,39. A characteristic peak was observed upon passage of a cell over the electrodes in the phase component of the output voltage at 1.12 and 1.5 MHz. Impedance measurements of the cells correlated well with optical monitoring of the cells flowing over the electrodes (Fig. 3a). The impedance peaks were then used to count the cells and quantify the growth rate.
Impedance phase signals measured at 1.5 MHz, recorded while cells pass over the electrodes.Figures a1–a5 show snapshots of events, which yielded two consecutive peaks. The first peak occurred, when the cell marked with a green circle passed over the electrode, followed by the one marked with a red circle. The blue arrow indicates the direction of the medium flow. b Number of peaks counted during 4-min windows through impedance measurements, plotted vs. time for two different analysis units having two and three filled pads. Each circle represents the number of peaks counted during the respective 4-min measurement window. tEIS denotes the time interval from the beginning of the experiment until impedance values were recorded, which was 13 h in this case. c Peak ratenorm plotted for both analysis units obtained by normalizing the peak count (obtained from impedance data) for the average cell number and the recording window duration (in hours). The green and red lines by and large coincide and indicate the mean growth rate for a given cell type extracted from all the measurement windows for a given analysis unit
We examined the effect of the electrode area, confined by the hexagonal structures, on the impedance signal. The detection volume is defined by the width of the channel segment above the electrodes, the electrode spacing, and the height of the chamber, which is within the reach of the electric field lines. This volume defines the magnitude of the impedance change upon passage of cells. Minimization of this detection volume increased the measurable impedance signal, so that the passage of cells could be detected. With larger detection volumes, for example upon misalignment of PDMS layer and glass slide, the detection of cells was no more possible (Figure S 5).
To enable the multiple use of the glass slide hosting the electrodes, the stability of the electrodes was analyzed. A frequency sweep on all electrodes of the device evidenced low variation in phase and magnitude signal between the beginning and the end of an experiment, which indicated the robustness of the electrodes (Figure S 6a). The low variation also reflected the uniformity of the fabricated electrode pairs across the chip. Further, the signal-to-noise ratio obtained from a re-used electrode pair at identical medium conditions and measurement frequency was measured for two separate experiments (Figure S 6b). Signal peaks could be detected in both experiments, and there was no significant difference in the observed signal-to-noise ratios of the two independent experiments. These findings illustrate that a glass slide with electrodes can be used for multiple experiments.
Cells flowing over a pair of co-planar electrodes show a characteristic signature, which depends on both cell size and cell position and speed40. The convolution of these effects currently limits the ability of the system in distinguishing between single or budded cells, and small cell clusters. Other studies have shown that optimized geometries and setups, featuring multiple facing and focusing electrodes, provide the capability to distinguish between different cell clusters26 or to analyze intra-cellular features41. However, integration of complex sensing structures, such as sandwich structures41 or electrode posts perpendicular to substrate plane42, into the current setup would increase device complexity and compromise ease of cell loading, culturing, and imaging. The current setup meets the requirements for measuring cell growth rates and changes in those, as the passage of clustered cells over the electrodes happened very rarely, as has been confirmed with microscopy observations. This statement is supported by the fact that the growth rates determined in our system coincide with those measured by other methods. In order to compensate for not being able to measure budding cells, a correction factor (mentioned later in the paper) that is commonly used in yeast biology and includes budded cells was applied to the measured growth rates.
Measuring growth of cell colonies
The loading process of the cells into the chamber did not produce a homogeneous cell distribution over all pad regions. Upon using low concentrations of cells in the loaded suspension in order to only clamp a few or a single cell under each pad, there is a high probability that some pads remain empty. After cell loading and starting medium perfusion, clamped cells grew and formed a 2D colony under the respective pads (Fig. 2b). Figure S 7a displays the number of cells under the pad over time, while the pad was getting filled. Once the pad was full, cells started to outgrow and were continuously removed from the colony through the medium flow around the pads. From this time on, the colony size under the pad remained stable and a counting of the outgrowing cells with respect to time could then be used as a direct measure for the growth rate of the cells under the pad.
Figure 3b presents peak counts (cells passing over the electrodes) within 4-min time windows for two different analysis units once the space under the pads was fully occupied. The analysis units featured three and two pads that had been filled with the same yeast strain. As expected, peak counts were lower for the analysis unit with two filled pads compared to that with three filled pads. To compare the measurements of the different analysis units, peak counts by the electrodes were normalized with regard to the number of cells under the respective pads. This was possible because the device allowed for simultaneous imaging of the pads during recording of the impedance signals.
Once the area under the pad was fully occupied, the variability of the cell number under the pad was low. Relative inter-pad variability in the number of cells under filled pads (in the same experiment and between experiments) was found to be approximately 8% for a given cell type (Figure S 7b). Averaged cell counts under the pads at the beginning and at the end of the impedance recording sessions were used for normalization. Further, the impedance peak count was also divided by the elapsed time of the recording window (4 min), which yielded the normalized peak rate (peak ratenorm). The peak ratenorm, calculated for both analysis units, was proportional to the growth rate of the cells under the respective pads. As shown in Fig. 3c, the calculated mean value of the peak ratenorm was found to be the same for the two analysis units (0.288 ± 0.009/h and 0.288 ± 0.015/h) in which the same strain had been cultured.
Parallel growth rate measurements of multiple strains
The setup allows for automated switching between the sensing electrodes of different analysis units and for sequential impedance recording in a round-robin configuration during the experiments (Figure S 3). We used this feature to analyze the variations in the growth rates of different yeast strains cultured on the same device by using the impedance readout. Two strains (vph1∆ and wild type) were loaded and cultured in different chambers in parallel. Both strains were first grown in YPD until the pads were filled. The medium was then switched to YPD with calcium and switched back to solely YPD after approximately 10 h. The strains are known to have different growth rates when calcium is added to the medium43. The switching time between the two media was ~40 s at a flow rate of 10 µl/min (Figure S 4) for all analysis units, such that the recordings in all analysis units under new media conditions could be performed after less than a minute. Switching time was therefore not a limiting factor in the experiments.
Figure 4a, b shows the normalized peak rates for the strains under different medium conditions. The average cell numbers under the pads at the beginning and at the end of the experiments for each medium condition were used for normalizing the peak rates. As expected, the mean peak rates for vph1∆ and the wild-type strain, grown in YPD, were similar (0.32 ± 0.02/h vs. 0.30 ± 0.01/h). When the media was switched to YPD containing calcium, the vph1∆ strain grew slower. The mean peak rate calculated for this condition was 0.196 ± 0.004/h for vph1∆ vs. 0.25 ± 0.01/h for the wild type. The cells fully recovered to their faster growth rates upon switching back to YPD without calcium (t = 20–26 h). We observed a higher variability in the peak rate for vph1∆ in YPD at the beginning of the experiment (t = 0–6 h) as compared to the end (t = 20–26 h). This finding can be attributed to the fact that the pads were initially not completely occupied, which resulted in a higher variation in the cell number under the pads and the related peak count.
Normalized peak rates of three replicates (represented by different symbols) of the Vph1∆ strain in YPD with calcium (100 mM) and in YPD without calcium. The line indicates the mean peak rate for each of the replicates. tEIS denotes the time from the beginning of the experiment until the respective impedance values were recorded, which was 13 h in this case for both strains. b Normalized peak rates of the wild-type strain (replicates represented by different symbols) in YPD with calcium (100 mM) and in YPD without calcium. c Growth rates as obtained from the plate reader (filled circles) and the impedance device (empty circles) under different medium conditions (YPD with and without calcium) for Vph1∆ (green) and wild-type cells (red). Each circle indicates one replicate. Squares are means, error bars represent standard deviations
The absolute growth rate of a monitored cell colony was obtained by taking into account that a fraction of cells passing over the sensing electrodes was in a budded state but yielded a single peak (Fig. 4c). As mentioned before, the current device design does not allow for differentiating between single and budded yeast cells. Therefore, the absolute growth rate was obtained after accounting for budded cells in the obtained peak rates. A budding index (percentage of budded cells) of ~0.6 (60% of all are budded) was measured for the cells used in these experiments through image analysis of the cells flowing over the electrodes. This value is comparable to the budding index value of 0.68 obtained from FACS analysis of the same cells grown under identical medium conditions in a shaking flask.
The growth rate values measured with the impedance device have also been compared to the ones obtained by using an automated plate reader for both strains under the different medium conditions. Figure 4c displays the growth rates obtained from the plate reader and obtained from the impedance device for both media: YPD with and without calcium. After adjusting the values of the impedance measurement with the budding index, the determined growth rates were comparable yet slightly lower than those obtained from the plate reader. The relative change in growth rate upon switching the medium from YPD with calcium to YPD for the vph1∆ strain in the plate reader and in the impedance device amounted to 58% and 63%. For the wild-type strain, the relative changes were 29% and 20%, respectively.
In summary, the developed platform enabled monitoring relative changes in growth rates of different strains in parallel upon changing the medium conditions. The change was more pronounced for vhp1∆ as compared to the wild type. Moreover, we showed that the replicates of the same strain yielded similar mean peak rates under the respective medium conditions, which demonstrates that the results of the impedance measurements for a given strain of cells were reproducible and consistent under identical conditions in the different analysis units. Additionally, we verified that the growth rate changes monitored in our platform were similar to those obtained with classical analysis methods.
Optimization of the multiplexed impedance readout
The presented device features a serial readout of 15 analysis units in a sequential way. A computer-controlled switch-board was used to route the selected electrode pair to the impedance analyzer for a defined time window. During this time, the cells passing the electrodes were counted according to the obtained impedance peaks. To maximize the information that can be obtained from multiplexing several analysis units, the recording time for each analysis unit needs to be optimized. This optimization can be achieved by defining a minimal time that is required per analysis unit to obtain an accurate estimate of the cell growth rate.
We continuously recorded from a cell colony of a single analysis unit over 8 h to analyze the factors that may define a suitable recording time. Figure 5a shows the peak counts for time windows of 60 and 240 s. More variability in the peak count per cycle was observed for a window of 60 s as compared to 240 s. In both cases, however, the mean peak count over all measurement cycles was identical within measurement errors. Figure 5b depicts the peak count in dependence of the time window. The variability in the peak number per cycle decreased with increasing time window, as the number of peaks was averaged over a longer duration, and was found to be relatively constant for measurement windows of 300 s or more.
Peak counts per second plotted for window lengths of 60 s (blue) and 240 s (red). The black line indicates the mean of the peak count for both window lengths. tEIS denotes the time from the beginning of the experiment until the respective impedance values were recorded, which was 24 h in this case. b Peak counts per second for window lengths varying from 1 to 600 s. c Mean values with error bars indicating SEM values for 60 s window length for different numbers of cycles. Below is the same standard error of the mean (SEM) for 60 s window length for different numbers of cycles. The x-axis at the bottom indicates the total measurement time in seconds (cycles × window length). d Mean values with error bars indicating SEM values and SEM values for 240 s window length
Larger time windows, however, reduce the sampling frequency of the analysis units during an experiment and, therefore, the temporal resolution of the obtained data (fewer red dots as compared to the blue dots in Fig. 5a). The window length and number of cycles ultimately define the obtainable measurement precision. Figure 5c, d shows the mean and standard error of the mean (SEM) for a window length of 60 and 240 s in dependence of the number of measurement cycles. As an example, a SEM of 0.004 was obtained, when an analysis unit was sampled ~85 times with a 60 s window but only ~22 times with a 240 s window. It can be inferred that for obtaining a certain precision, fewer cycles are required for recording with longer time windows from a given analysis unit. The total recording time was similar (5100 and 5280 s) for the two examples detailed above.
The number of cells passing over the electrodes is a stochastic process that can be modeled by a Poisson distribution. One feature of the Poisson model is that the precision of the estimate only depends on the total recording time, irrespective of the length of each recording interval. Consequently, it does not matter if several longer-interval or many short-interval recordings are conducted, as long as the overall recording duration is the same. It has to be noted that the window size per analysis unit should be chosen with respect to the growth rate of the cells—an analysis unit featuring slowly growing cells should be assigned a longer recording window as compared to a unit with rapidly growing cells to achieve, in both cases, good precision of the relative growth rate measurements.
Cell imaging in the microfluidic device
The advantage of the “micro-chemostats” is that they are amenable to high-resolution imaging. We tested the imaging capabilities in the devices by using a Citrine tag endogenously fused to different yeast proteins. For high-resolution fluorescence imaging, the electrodes were patterned on a 200-µm-thick glass. The fluorescent fusion proteins localized to the vacuolar membrane (Vph1-Citrine), at the bud neck (Cdc12-Citrine) and shuffled between the cytoplasm and the nucleus in a cell-cycle-dependent manner (Whi5-Citrine). All three cell types were inoculated separately, mixed in equal ratios, and cultured in the device.
The different cell types and their growth over time were clearly distinguishable according to their different levels of fluorophore signals (Fig. 6a). The intensity of the Citrine tag was highest for Vph1 tagged cells (~2940 copies/cell44), followed by Cdc12 (~680 copies/cell) and Whi5 (~120 copies/cell) tagged cells. Figure 6b shows the images of single cells featuring the different tags. The appearance of the Cdc12 marker (in yellow) at the bud neck and the localization of Whi5 in the nucleus shortly before cell division were clearly visible in the device (see also video S 1). The use of a thin glass slide makes it possible to image intracellular components and enables high-resolution microscopy comparable to that with commonly used glass slides.
Time-lapse images showing the growth of different cells under one of the pads (40× NA 0.75). The three strains are indicated through different colors in the bright-field images. The strains were tagged with the Citrine fluorophore at the bud neck (Cdc12; yellow), in the vacuole (Vph1; blue), and in the nucleus (Whi5; green). The pad hosts all three strains that grow, divide, and feature different fluorescence intensities of Citrine (video S 1). b Enlarged images of cells with the different tags. Arrows indicate the cellular distribution and transitional localization of Whi5 in the nucleus of a cell
Conclusion
We described a platform enabling long-term culturing and high-resolution imaging of yeast cells, while the growth rates of populations of cells could be assessed in parallel and label free by means of impedance measurements. The platform allows for simultaneous culturing of up to three strains under different microfluidic-flow and medium conditions, and features five sets of analysis units per strain for parallelized in situ impedance analysis of the growth rate. The availability of multiple analysis units enables the user to conduct experiments in the same platform in parallel in several replications and to include all necessary control experiments, while having the capability to perform optical imaging. Importantly, the use of non-permanent vacuum-based bonding between the microfluidic structure and the glass slide hosting the electrodes allows for simple sample loading into the device and multiple usage of the glass slides.
The yeast cells grow in a monolayer under the clamping pads, they divide and develop—potentially from a single cell—to a cell colony of constant size, which is defined by the clamping pad dimensions. We validated all functions of the platform by continuously monitoring the changes in the growth rates of differently engineered yeast strains, loaded in separate culturing compartments of the device, upon changing media conditions by using the impedance readout. The observed growth rates and their changes coincide with those recorded by using standard methods. We also confirmed the quality of the optical imaging by using a set of endogenously tagged proteins that are frequently used in yeast research. This experiment shows that continuous bright-field and fluorescence imaging of the cells at sub-cellular resolution were possible with our platform.
Growth rate measurements were based on impedance and complementary to measurements performed by microscopy. Furthermore, the automated recording and switching between several impedance analysis units was completely independent of the position, temporal resolution, or field of view of the microscope used in parallel. Such a platform could serve as a useful experimental setup for nutrition and cell metabolism studies that require a continuous monitoring of growth rates in a steady and controlled environment along with tracking of fast cellular processes by means of high-resolution time-lapse imaging.
References
Brauer, MatthewJ. et al. Cordination of growth rate, cell cycle,stress response and metabolic activity in yeast. Mol. Biol. Cell 19, 308–317 (2008).
Burnetti, AnthonyJ. et al. Cell cycle start is coupled to entry into the yeast metabolic cycle across diverse strains and growth rates. Mol. Biol. Cell XXXIII, 81–87 (2015).
Ziv, N., Siegal, M. L. & Gresham, D. Genetic and nongenetic determinants of cell growth variation assessed by high-throughput microscopy. Mol. Biol. Evol. 30, 2568–2578 (2013).
Slavov, N. & Botstein, D. Coupling among growth rate response, metabolic cycle, and cell division cycle in yeast. Mol. Biol. Cell 22, 1997–2009 (2011).
Amodeo, A. A. & Skotheim, J. M. Cell-size control. Cold Spring Harb. Perspect. Biol. 8, a019083 (2016).
Wang, J. D. & Levin, P. A. Metabolism, cell growth and the bacterial cell cycle. Nat. Rev. Microbiol. 7, 822–827 (2009).
Cai, L. & Tu, B. P. Driving the cell cycle through metabolism. Annu. Rev. Cell Dev. Biol. 28, 59–87 (2012).
Botstein, D. & Fink, G. R. Yeast: an experimental organism for 21st century biology. Genetics 189, 695–704 (2011).
Boone, C. Yeast systems biology: our best shot at modeling a cell. Genetics 198, 435–437 (2014).
Hoskisson, P. A. & Hobbs, G. Continuous culture - making a comeback? Microbiology 151, 3153–3159 (2005).
Groisman, A. et al. A microfluidic chemostat for experiments with bacterial and yeast cells. Nat. Methods 2, 685–689 (2005).
Balagadde, F. K., You, L., Hansen, C. L., Arnold, F. H. & Quake, S. R. Long-term monitoring of bacteria undergoing programmed population control in a microchemostat. Science 309, 137–140 (2005).
Denervaud, N. et al. A chemostat array enables the spatio-temporal analysis of the yeast proteome. Proc. Natl Acad. Sci. USA 110, 15842–15847 (2013).
Long, Z. et al. Microfluidic chemostat for measuring single cell dynamics in bacteria. Lab Chip 13, 947–954 (2013).
Nobs, J.-B. & Maerkl, S. J. Long-term single cell analysis of S. pombe on a microfluidic microchemostat array. PLoS ONE 9, e93466 (2014).
Rines, D. R., Thomann, D., Dorn, J. F., Goodwin, P. & Sorger, P. K. Live cell imaging of yeast. Cold Spring Harb. Protoc. 2011, 1026–1041 (2011).
Lee, K. S., Boccazzi, P., Sinskey, A. J. & Ram, R. J. Microfluidic chemostat and turbidostat with flow rate, oxygen, and temperature control for dynamic continuous culture. Lab Chip 11, 1730 (2011).
Zhang, Z. et al. Microchemostat—microbial continuous culture in a polymer-based, instrumented microbioreactor. Lab Chip 6, 906–913 (2006).
Hassan, U. & Bashir, R. Electrical cell counting process characterization in a microfluidic impedance cytometer. Biomed. Microdevices 16, 697–704 (2014).
van Berkel, C. et al. Integrated systems for rapid point of care (PoC) blood cell analysis. Lab Chip 11, 1249 (2011).
Holmes, D. et al. Leukocyte analysis and differentiation using high speed microfluidic single cell impedance cytometry. Lab Chip 9, 2881–2889 (2009).
Sun, T. & Morgan, H. Single-cell microfluidic impedance cytometry: a review. Microfluid Nanofluidics 8, 423–443 (2010).
Cheung, K., Gawad, S. & Renaud, P. Impedance spectroscopy flow cytometry: on-chip label-free cell differentiation. Cytometry A 65, 124–132 (2005).
Cheung, K. C. et al. Microfluidic impedance-based flow cytometry. Cytometry A 77A, 648–666 (2010).
McGrath, J. S. et al. Analysis of parasitic protozoa at the single-cell level using microfluidic impedance cytometry. Sci. Rep. 7, 2601 (2017).
Haandbæk, N., Bürgel, S. C., Rudolf, F., Heer, F. & Hierlemann, A. Characterization of single yeast cell phenotypes using microfluidic impedance cytometry and optical imaging. ACS Sensors 1, (1020–1027 (2016).
Wegener, J., Keese, C. R. & Giaever, I. Electric cell–substrate impedance sensing (ECIS) as a noninvasive means to monitor the kinetics of cell spreading to artificial surfaces. Exp. Cell. Res. 259, 158–166 (2000).
Giaever, I. & Keese, C. R. A morphological biosensor for mammalian cells. Nature 366, 591–592 (1993).
Krommenhoek, E. E. et al. Lab-scale fermentation tests of microchip with integrated electrochemical sensors for pH, temperature, dissolved oxygen and viable biomass concentration. Biotechnol. Bioeng. 99, 884–892 (2008).
Luchterhand, B. et al. Newly designed and validated impedance spectroscopy setup in microtiter plates successfully monitors viable biomass online. Biotechnol. J. 10, 1259–1268 (2015).
Frey, O., Rudolf, F., Schmidt, G. W. & Hierlemann, A. Versatile, simple-to-use microfluidic cell-culturing chip for long-term, high-resolution, time-lapse imaging. Anal. Chem. 87, 4144–4151 (2015).
Bürgel, S. C., Diener, L., Frey, O., Kim, J.-Y. & Hierlemann, A. Automated, multiplexed electrical impedance spectroscopy platform for continuous monitoring of microtissue spheroids. Anal. Chem. 88, 10876–10883 (2016).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Mayer, C., Dimopoulos, S., Rudolf, F. & Stelling, J. Using CellX to quantify intracellular events. Curr. Protoc. Mol. Biol. 14, 14.22 (2013).
Schmidt, G. W., Frey, O. & Rudolf, F. The CellClamper: a convenient microfluidic device for time-lapse imaging of yeast. Methods Mol. Biol. 1672, 537–555 (2018).
Liu, R. H. et al. Passive mixing in a three-dimensional serpentine microchannel. J. Microelectromech. Syst. 9, 190–197 (2000).
Yang, J.-T. & Lin, K.-W. Mixing and separation of two-fluid flow in a micro planar serpentine channel. J. Micromech. Microeng. 16, 2439–2448 (2006).
Gawad, S., Schild, L. & Renaud, P. H. Micromachined impedance spectroscopy flow cytometer for cell analysis and particle sizing. Lab Chip 1, 76–82 (2001).
Morgan, H., Sun, T., Holmes, D., Gawad, S. & Green, N. G. Single cell dielectric spectroscopy. J. Phys. D Appl. Phys. 61, 61–70 (2007).
Spencer, D. & Morgan, H. Positional dependence of particles in microfludic impedance cytometry. Lab Chip 11, 1234–1239 (2011).
Haandbæk, N., Bürgel, S. C., Heer, F. & Hierlemann, A. Characterization of subcellular morphology of single yeast cells using high frequency microfluidic impedance cytometer. Lab Chip 14, 369–377 (2014).
Kilchenmann, S. C., Rollo, E., Maoddi, P. & Guiducci, C. Metal-coated SU-8 structures for high-density 3-D microelectrode arrays. J. Microelectromech. Syst. 25, 425–431 (2016).
Manolson, M. et al. The VPH1 gene encodes a 95-kDa integral membrane polypeptide required for in vivo assembly and activity of the yeast vacuolar H(+)-ATPase. J. Biol. Chem. 267, 14294–14303 (1992).
Chong, Y. et al. Yeast proteome dynamics from single cell imaging and automated analysis. Cell 161, 1413–1424 (2015).
Acknowledgements
The work was financially supported by the Swiss SystemsX.ch IPhD program, by the FP7 of the EU through the MTN ISOLATE, Contract Number 289995, and the Ambizione Grant 142440 of the Swiss National Science Foundation for Olivier Frey. We extend our gratitude to Moritz Lang, ETH Zurich, for helping out with the Youscope software. We are grateful to Mario M. Modena, Nassim Rousset, Andreas P. Cuny, all at ETH Zurich for their inputs and assistance. The authors would like to thank Paul Argast in the workshop, Alexander Stettler and Albert Martel in the cleanroom facility, and the staff in the single-cell facility at D-BSSE, ETH Zurich, for their help and support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chawla, K., Bürgel, S.C., Schmidt, G.W. et al. Integrating impedance-based growth-rate monitoring into a microfluidic cell culture platform for live-cell microscopy. Microsyst Nanoeng 4, 8 (2018). https://doi.org/10.1038/s41378-018-0006-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41378-018-0006-5
This article is cited by
-
Analytical modelling and numerical simulations for microfluidic chip enabling accurate sheathless impedance spectroscopy
Research on Biomedical Engineering (2024)
-
piRT-IFC: Physics-informed real-time impedance flow cytometry for the characterization of cellular intrinsic electrical properties
Microsystems & Nanoengineering (2023)
-
Tumour-on-chip microfluidic platform for assessment of drug pharmacokinetics and treatment response
Communications Biology (2021)
-
Investigation of daughter cell dissection coincidence of single budding yeast cells immobilized in microfluidic traps
Analytical and Bioanalytical Chemistry (2021)
-
Design and analysis of microfluidic cell counter using spice simulation
SN Applied Sciences (2019)