Abstract
Biomolecular analysis at the single-cell level is increasingly important in the study of cellular heterogeneity and its consequences, particularly in organismic development and complex diseases such as cancer. Single-cell molecular analyses have led to the identification of new cell types1 and the discovery of novel targets for diagnosis and therapy2. While these analyses are performed predominantly on dissociated single cells, emerging techniques seek understanding of cellular state, cellular function and cell–cell interactions within the native tissue environment by combining optical microscopy and single-cell molecular analyses. These techniques include in situ multiplexed imaging of fluorescently labeled proteins and nucleotides, as well as low-throughput ex vivo methods in which specific cells are isolated for downstream molecular analyses. However, these methods are limited in either the number and type of molecular species they can identify or the number of cells that can be analyzed. High-throughput methods are needed for comprehensive profiling of many cells (>1000) to detect rare cell types, discriminate relevant biomarkers from intrinsic population noise, and reduce the time and cost of measurement. Many established, high-throughput single-cell analyses are not directly applicable because they require tissue dissociation, leading to a loss of spatial information3. No current methods exist that can seamlessly connect spatial mapping to single-cell techniques. In this Perspective, we review current methods for spatially resolved single-cell analysis and discuss the prospect of novel multiplexed imaging probes, called laser particles, which allow individual cells to be tagged in tissue and analyzed subsequently using high-throughput, comprehensive single-cell techniques.
Similar content being viewed by others
Current methods for spatially resolved single-cell analysis
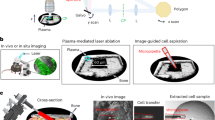
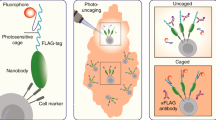
Optical fluorescence microscopy has been a powerful tool to understand cellular function and phenotypic heterogeneity (Fig. 1a). Image-derived phenotypes comprise spatial information such as morphology and location relative to other cells, as well as dynamic behaviors, such as migration and invasion, and result from the complex interplay of different cell types within tissues. Fluorescence imaging at high spatial resolution can also provide molecular analysis in situ within each individual cell (Fig. 1b), thereby enabling correlation of image-derived phenotypes and molecular profiles. The most pervasive approach is the antibody-based fluorescence detection of specific proteins after tissue fixation and permeabilization. Repeated antibody elution and staining steps4 or the use of DNA-barcoded antibodies5 can extend multiplexed detection from 4~40 proteins per sample. Another approach involves imaging RNA in cells with single-molecule fluorescence in situ hybridization (smFISH) using targeted oligonucleotide probes. For multiplexed detection, sequential rounds of smFISH imaging can detect ~100 distinct RNA species in single cells6. Enzymatic amplification of RNA species to generate cDNA nanoballs (amplicons) in fluorescence in situ sequencing (FISSEQ) enables orders of magnitude higher signal intensity compared to that of smFISH7. An approach called STARmap combines RNA amplification, tissue optical clearing and fluorescence-based readout to detect up to ~1000 genes in brain tissue at single-cell resolution8.
a Imaging in tissues provides information on where cells are located with respect to each other and their environment, as well as observation of dynamic behaviors in their native environment. b Biomolecules can be imaged in situ with high-resolution fluorescence imaging but with limited multiplexing. Proteins can be imaged with immunofluorescence. Fluorescence in situ hybridization enables imaging of targeted nucleic acids, including DNA and RNA. Fluorescence in situ sequencing enables imaging of untargeted transcriptomic mRNA expression. c Laser microdissection and photoactivation/photoconversion enable the selection of specific cells of interest for downstream molecular analyses. Spatial barcoding of tissue sections enables high-throughput spatial transcriptomics, but not at single-cell resolution
Although in situ molecular profiling techniques retain spatial information, their scope is limited compared to that of widespread single-cell technologies that analyze dissociated suspensions of cells devoid of spatial information. Current multiplexed in situ techniques primarily measure RNA, do not cover the whole transcriptome (>10,000 expressed genes), and require laborious imaging steps. In contrast, single-cell technologies based on next-generation sequencing (NGS) techniques comprise a suite of “omic” methods that can more comprehensively profile the genome, epigenome, transcriptome and proteome3. These methods include more established techniques, such as high-throughput single-cell DNA and RNA sequencing (scDNA-seq and scRNA-seq), to capture the whole genome and transcriptome, respectively, as well as single-cell mass cytometry for multiplexed analysis of proteins9. Emerging epigenetic techniques that probe how gene expression is regulated include ChIP-seq (chromatin immunoprecipitation and sequencing) and ATAC-seq (assay for transposase-accessible chromatin using sequencing). However, as mentioned, all of these technologies require tissue dissociation into cellular suspensions, which leads to the loss of all spatial information.
Several techniques restrict molecular analysis to specific cells that can be defined and selected during imaging (Fig. 1c), thereby retaining spatial information, but with a tremendous cost to throughput. Methods based on photoactivation or photoconversion10 involve tagging of selected cells while imaging and subsequent isolation of the selected cells or cellular material for downstream analyses. Laser capture microdissection (LCM) is a more mature technology that involves the isolation of cells from tissue by laser ablation under the microscope. LCM has been employed in an automated manner to couple with single-cell genomic11, transcriptomic12 and proteomic profiling13 to obtain spatially resolved molecular profiles. Instead of isolating individual cells for analysis, significantly higher throughput can be achieved with spatial barcoding. One method uses an array of DNA-barcoded oligonucleotides on glass slides that capture mRNA from permeabilized tissue sections14. Subsequent RNA-seq is mapped to the spatial coordinates of the array and thus the tissue. While this approach improves throughput compared to LCM, it is limited to RNA profiling and does not maintain depth information about cells within the tissue slice. At present, there is no high-throughput method that permits comprehensive molecular analysis with the full complement of single-cell omics technologies while retaining spatial information.
Current methods for multiplexed cell tagging
Cell tagging enables the tracking of cellular identities over time and space and potentially across analytical platforms to correlate spatial and molecular analysis. However, conventional fluorescent probes allow only a few different cell types to be labeled with different colors due to spectral overlap. Recently, the development of multiplexed imaging techniques has permitted the labeling and tracking of a far greater number of cellular populations, a new paradigm for biological imaging, which has gained traction given the increasing appreciation of intercellular diversity. Tracking individual cells through imaging gives access to their properties (e.g., spatial location, morphology) over time, as well as the spatial relationships between different cells. The combinatorial expression of multiple fluorescent proteins in each cell can achieve up to ~100 resolvable colors using ratiometric coding15. This technique has been used to discriminate neurons in brain tissue15 and for long-term cell tracking in zebrafish skin16. Quantum dots have also been employed for image-based multiplexing in vitro17. This ratiometric or image-based coding, however, is subject to error for thick tissues due to variable attenuation and scattering. Other approaches proposed to overcome this limitation include multiplexed lifetime imaging with lanthanide nanoparticles18 and Raman multiplexed probes with 24 colors19. However, none of these techniques are easily scalable to a much larger number (>1000) of cells.
Laser particles for single-cell analysis
A new class of nano/microsized imaging probes has emerged that harness narrowband laser emission to enable massively multiplexed cell tagging. These laser particles (LPs) are particularly promising for single-cell analysis for several reasons. First, they provide a means for narrowband laser emission from inside cells, This enables cell tagging by spectral barcoding, which is far more reliable than barcoding based on intensity levels or imaging features that are of limited use in tissue due to wavelength-dependent absorption and scattering. Second, these spectral barcodes can be repeatedly read in a nondestructive manner. This allows cell tracking over time and space in biological systems, as well as the retention of individual cellular identity across different analytical platforms, which enables the association of spatial information and other image-derived phenotypes with molecular profiles.
Several potential designs have been reported, including fluorescent polystyrene particles20,21, upconverting nanoparticle microlasers22, perovskite nanowires23, and plasmonic nanoparticles24. Our recent study employed silica-coated III–V semiconductor microdisk LPs with diameters of ~2 μm and total thicknesses of ~400 nm25. Our microdisk LPs have several key properties that make them more suitable for cellular barcoding than the previous designs, including relatively small sizes (~0.1% of the cell volume), stability in aqueous environments, biocompatibility, and tunable wavelengths over a wide range. Silica-coated microdisk LPs were readily internalized into a variety of cell types through macropinocytosis and had no appreciable effect on cell viability, cell cycle time, or motility when internalized inside cells25. Using five different semiconductor alloy compositions and stochastically varying the microdisk diameter through nanofabrication enabled single-mode emission over a wide range in the tissue-penetrating NIR-II window, from 1170 to 1580 nm, with subnanometer linewidths (Fig. 2a). Given 1-nm bins within which LP emission is stable, this range provides more than 400 unique colors, with room for further expansion using different materials (e.g., InGaAsP and InAlGaAs for NIR-I and NIR-II, InGaN for visible). The use of the NIR range also preserves compatibility with conventional fluorescent probes in the visible wavelengths that can label different cellular features (e.g., nuclei, membrane). The availability of hundreds of spectrally distinct colors for multiplexed imaging enables longitudinal cell tracking even in dense, scattering tissues. In our study25, we used microdisk LPs to track thousands of individual cancer cells over several days in a 3D tumor spheroid invasion model. Our data enabled the classification of individual cells according to their motility and the identification of cells moving together in small packs within the spheroid by analyzing spatial correlations in velocities.
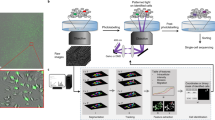
a Left: SEM of a silica-coated, III–V semiconductor microdisk laser particle (LP). Middle: Narrowband emission measured from the LP upon optical pumping with a 1060 nm laser emitting 3 ns pulses at 2 MHz and 20 pJ/pulse. Right: Use of different materials and fabrication of different microdisk diameters by nanolithography yields 400 LPs with single-mode emission from 1170 to 1580 nm. Reproduced from Ref 25. b Spectral barcoding by using combinations of different LPs per cell. c A human cell (HEK293) carrying three LPs. Magenta (LP), green (membrane), blue (nucleus). d Left: Fabrication of multiplet LPs. Right: An example of emission from a triplet LP, one of ~11 million barcodes possible assuming 400 spectral channels. e Proposed workflow for comprehensive single-cell analysis using LPs. Tissue-containing laser particles are imaged to capture tissue architecture and cellular behaviors. LP emission is measured before and after dissociation to track cell identity. Any downstream molecular analysis of single cells (e.g., transcriptomics and proteomics) can be correlated to their location and behavior in their native tissue
Despite progress in the development of multiplexed imaging probes, the number of resolvable colors, or available barcodes, is still far fewer than desired for most applications (~10,000 cells per 1 mm3 of tissue volume). To tag cells uniquely, the number of barcodes should exceed the number of cells. Spectral barcoding with LPs is a practical and scalable approach to break through this limitation. LPs with different emissions can be combined to readily achieve millions of unique barcodes (Fig. 2b). One approach is the uptake of multiple LPs in each cell (Fig. 2c), as long as LPs can be colocalized to specific cells by segmentation of cell boundaries. However, this approach would not be suitable for dividing cells since barcode fidelity would be lost upon LP segregation. A more tractable approach is using multiple LPs stacked together in a single particle to generate an emission composed of multiple laser lines25 (Fig. 2d). To associate spatial information with single-cell analyses, the emission spectra of LP tagged cells can be read during imaging to catalog the position and identity of cells within tissue and following tissue dissociation at an appropriate step before subsequent analyses (Fig. 2e). For plate-based sequencing, LP emission can be measured during or after the sorting of single cells into individual microwells. LP cell tagging is also compatible with high-throughput methods, including microdroplet-based26 and split-pool barcoding27, which introduce cell-specific DNA barcodes for highly parallel analysis as long as an association between the optical LP barcode and the DNA barcode is made.
Further development of the LP technology could enable the phenotypic and molecular characterization of every cell in a tissue, limited only by light penetration during imaging. One critical area of research is cancer biology. A major question is how, when, and which cancer cells within a heterogeneous tumor become metastatic. Live imaging and multiplexed cell tracking with LPs in dense tumor tissue can delineate key steps in the metastatic cascade, including migration, intravasation, and extravasation, as well as interactions with the tumor microenvironment. Linking these cellular phenotypes with molecular profiles could address the largely unexplored roles of cellular cooperation and non-cell-autonomous behaviors at the single-cell level. Analysis of interacting ligands and receptors on individual cells may lead to the construction of a comprehensive, spatially resolved cell–cell interaction network to reveal functional mechanisms in metastatic tumor progression. Beyond research applications, similar experiments conducted on patient-derived tumor models could identify biomarkers for diagnosis or targets for therapy.
Another application of single-cell analyses, particularly scRNA-seq, is unbiased cell classification, which has been successful in identifying new cell types in a number of different tissues. One reason for their success is a data-driven approach that is unencumbered by previous biases towards cellular identity. Techniques that integrate multiple omics layers of data (e.g., transcriptomic and epigenomic) provide a more complete picture of a cell’s molecular framework that is thus more powerful than using any single layer alone3. Cellular phenotypes captured by imaging can be considered additional layers of information that reflect a cell’s native environment. Since cells do not naturally exist in isolation, this information is essential for understanding cellular state, function, and cell–cell interactions in their native tissue environment. A recent initiative by an international consortium to characterize every cell in the human body, the Human Cell Atlas, has highlighted the need for spatial information to generate comprehensive reference maps as a basis for diagnosing, monitoring, and treating disease28. Spatially resolved single-cell analysis will likely transform applications such as tissue engineering and precision medicine. Comprehensive datasets combining imaging and molecular profiles could also be used to train machine-learning algorithms for image-based cell classification and in silico diagnostics.
In conclusion, advances in cellular imaging and molecular analyses have pushed the envelope of what can be observed and measured, but their impact has been hampered by the inability to employ these technologies in tandem on the same cells. Laser particles are a novel class of multiplexed imaging probes with the potential to bridge these technologies by enabling massively multiplexed imaging, cell tagging, and comprehensive single-cell analysis.
References
Montoro, D. T. et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 560, 319–324 (2018).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Chappell, L., Russell, A. J. C. & Voet, T. Single-cell (multi)omics technologies. Annu. Rev. Genom. Hum. Genet. 19, 15–41 (2018).
Gut, G., Herrmann, M. D. & Pelkmans, L. Multiplexed protein maps link subcellular organization to cellular states. Science 361, eaar7042 (2018).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981.e15 (2018).
Chen, K. H. et al. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Lee, J. H. et al. Highly multiplexed subcellular RNA sequencing in situ. Science 343, 1360–1363 (2014).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Spitzer, M. H. & Nolan, G. P. Mass cytometry: single cells, many features. Cell 165, 780–791 (2016).
Lovatt, D. et al. Transcriptome in vivo analysis (TIVA) of spatially defined single cells in live tissue. Nat. Methods 11, 190–196 (2014).
Casasent, A. K. et al. Multiclonal invasion in breast tumors identified by topographic single cell sequencing. Cell 172, 205–217.e12 (2018).
Moor, A. E. et al. Spatial reconstruction of single enterocytes uncovers broad zonation along the intestinal villus axis. Cell 175, 1156–1167.e15 (2018).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Chen, C. H. et al. Multicolor cell barcoding technology for long-term surveillance of epithelial regeneration in zebrafish. Dev. Cell 36, 668–680 (2016).
Rees, P. et al. Nanoparticle vesicle encoding for imaging and tracking cell populations. Nat. Methods 11, 1177–1181 (2014).
Fan, Y. et al. Lifetime-engineered NIR-II nanoparticles unlock multiplexed in vivo imaging. Nat. Nanotechnol. 13, 941–946 (2018).
Wei, L. et al. Super-multiplex vibrational imaging. Nature 544, 465–470 (2017).
Humar, M. & Yun, S. H. Intracellular microlasers. Nat. Photonics 9, 572–576 (2015).
Schubert, M. et al. Lasing within live cells containing intracellular optical microresonators for barcode-type cell tagging and tracking. Nano Lett. 15, 5647–5652 (2015).
Fernandez-Bravo, A. et al. Continuous-wave upconverting nanoparticle microlasers. Nat. Nanotechnol. 13, 572–577 (2018).
Cho, S. et al. Laser particle stimulated emission microscopy. Phys. Rev. Lett. 117, 193902 (2016).
Galanzha, E. I. et al. Spaser as a biological probe. Nat. Commun. 8, 15528 (2017).
Martino, N. et al. Wavelength-encoded laser particles for massively-multiplexed cell tagging. Nat. Photon. https://doi.org/10.1101/465104. (2019)
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Cao, J. Y. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Regev, A. et al. The human cell atlas. eLife 6, e27041 (2017).
Acknowledgements
S.H.Y. acknowledges grants from US National Institutes of Health (DP1-OD022296, P41-EB015903, R01-CA192878), Human Frontier Science Program (RGP0034/2016), and MGH Research Scholar Program. Light: Science and Applications thanks Prof Dayong Jin for handling this article and for his advice on enhancing its presentation.
Editor Information
Light: Science & Applications thanks Prof Dayong Jin for handling this article and for his advice on enhancing its presentation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
S.J.J.K. receives compensation as an employee of LASE Innovation Inc., a company focused on commercializing technologies based on LPs. S.H.Y. has financial interests in LASE Innovation Inc., and holds patents on technologies related to the devices developed in this work. S.H.Y.’s interests were reviewed and are managed by Massachusetts General Hospital and Partners HealthCare in accordance with their conflict of interest policies. The remaining authors declate that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kwok, S.J.J., Martino, N., Dannenberg, P.H. et al. Multiplexed laser particles for spatially resolved single-cell analysis. Light Sci Appl 8, 74 (2019). https://doi.org/10.1038/s41377-019-0183-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41377-019-0183-5
This article is cited by
-
Micro/Nano Lasers for Biomolecular Sensing and Cellular Analysis
Photonic Sensors (2025)
-
A high-precision multi-dimensional microspectroscopic technique for morphological and properties analysis of cancer cell
Light: Science & Applications (2023)
-
Review of biosensing with whispering-gallery mode lasers
Light: Science & Applications (2021)
-
Laser particles with omnidirectional emission for cell tracking
Light: Science & Applications (2021)
-
Whispering-gallery microlasers for cell tagging and barcoding: the prospects for in vivo biosensing
Light: Science & Applications (2021)