Abstract
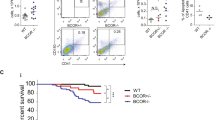
Myelodysplastic syndromes (MDS) are myeloid neoplasms presenting with dysplasia in the bone marrow (BM) and peripheral cytopenia. In most patients anemia develops. We screened for genes that are expressed abnormally in erythroid progenitor cells (EP) and contribute to the pathogenesis of MDS. We found that the Coxsackie-Adenovirus receptor (CAR = CXADR) is markedly downregulated in CD45low/CD105+ EP in MDS patients compared to control EP. Correspondingly, the erythroblast cell lines HEL, K562, and KU812 stained negative for CAR. Lentiviral transduction of the full-length CXADR gene into these cells resulted in an increased expression of early erythroid antigens, including CD36, CD71, and glycophorin A. In addition, CXADR-transduction resulted in an increased migration against a serum protein gradient, whereas truncated CXADR variants did not induce expression of erythroid antigens or migration. Furthermore, conditional knock-out of Cxadr in C57BL/6 mice resulted in anemia and erythroid dysplasia. Finally, decreased CAR expression on EP was found to correlate with high-risk MDS and decreased survival. Together, CAR is a functionally relevant marker that is down-regulated on EP in MDS and is of prognostic significance. Decreased CAR expression may contribute to the maturation defect and altered migration of EP and thus their pathologic accumulation in the BM in MDS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bennett JM, Kouides PA, Forman SJ. The myelodysplastic syndromes: morphology, risk assessment, and clinical management (2002). Int J Hematol. 2002;76:228–38.
Valent P, Horny HP, Bennett JM, Fonatsch C, Germing U, Greenberg P, et al. Definitions and standards in the diagnosis and treatment of the myelodysplastic syndromes: consensus statements and report from a working conference. Leuk Res. 2007;31:727–36.
Nimer SD. Myelodysplastic syndromes. Blood. 2008;111:4841–51.
Corey SJ, Minden MD, Barber DL, Kantarjian H, Wang JC, Schimmer AD. Myelodysplastic syndromes: the complexity of stem-cell diseases. Nat Rev Cancer. 2007;7:118–29.
Haase D, Germing U, Schanz J, Pfeilstocker M, Nosslinger T, Hildebrandt B, et al. New insights into the prognostic impact of the karyotype in MDS and correlation with subtypes: evidence from a core dataset of 2124 patients. Blood. 2007;110:4385–95.
Mills KI, Kohlmann A, Williams PM, Wieczorek L, Liu WM, Li R, et al. Microarray-based classifiers and prognosis models identify subgroups with distinct clinical outcomes and high risk of AML transformation of myelodysplastic syndrome. Blood. 2009;114:1063–72.
Bejar R, Levine R, Ebert BL. Unraveling the molecular pathophysiology of myelodysplastic syndromes. J Clin Oncol Off J Am Soc Clin Oncol. 2011;29:504–15.
Schanz J, Steidl C, Fonatsch C, Pfeilstocker M, Nosslinger T, Tuechler H, et al. Coalesced multicentric analysis of 2351 patients with myelodysplastic syndromes indicates an underestimation of poor-risk cytogenetics of myelodysplastic syndromes in the international prognostic scoring system. J Clin Oncol Off J Am Soc Clin Oncol. 2011;29:1963–70.
Lindsley RC, Ebert BL. Molecular pathophysiology of myelodysplastic syndromes. Ann Rev Pathol. 2013;8:21–47.
Smith AE, Mohamedali AM, Kulasekararaj A, Lim Z, Gaken J, Lea NC, et al. Next-generation sequencing of the TET2 gene in 355 MDS and CMML patients reveals low-abundance mutant clones with early origins, but indicates no definite prognostic value. Blood. 2010;116:3923–32.
Makishima H, Visconte V, Sakaguchi H, Jankowska AM, Abu Kar S, Jerez A, et al. Mutations in the spliceosome machinery, a novel and ubiquitous pathway in leukemogenesis. Blood. 2012;119:3203–10.
Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G, et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia. 2014;28:241–7.
Duetz C, Westers TM, van de Loosdrecht AA. Clinical implication of multi-parameter flow cytometry in myelodysplastic syndromes. Pathobiol: J Immunopathol, Mol Cell Biol. 2019;86:14–23.
Malcovati L, Della Porta MG, Lunghi M, Pascutto C, Vanelli L, Travaglino E, et al. Flow cytometry evaluation of erythroid and myeloid dysplasia in patients with myelodysplastic syndrome. Leukemia. 2005;19:776–83.
Della Porta MG, Malcovati L, Invernizzi R, Travaglino E, Pascutto C, Maffioli M, et al. Flow cytometry evaluation of erythroid dysplasia in patients with myelodysplastic syndrome. Leukemia. 2006;20:549–55.
Westers TM, Cremers EM, Oelschlaegel U, Johansson U, Bettelheim P, Matarraz S, et al. Immunophenotypic analysis of erythroid dysplasia in myelodysplastic syndromes. A report from the IMDSFlow working group. Haematologica. 2017;102:308–19.
Ogata K, Nakamura K, Yokose N, Tamura H, Tachibana M, Taniguchi O, et al. Clinical significance of phenotypic features of blasts in patients with myelodysplastic syndrome. Blood. 2002;100:3887–96.
van de Loosdrecht AA, Westers TM, Westra AH, Drager AM, van der Velden VH, Ossenkoppele GJ. Identification of distinct prognostic subgroups in low- and intermediate-1-risk myelodysplastic syndromes by flow cytometry. Blood. 2008;111:1067–77.
Porwit A, van de Loosdrecht AA, Bettelheim P, Brodersen LE, Burbury K, Cremers E, et al. Revisiting guidelines for integration of flow cytometry results in the WHO classification of myelodysplastic syndromes-proposal from the International/European LeukemiaNet Working Group for Flow Cytometry in MDS. Leukemia. 2014;28:1793–8.
Xu F, Wu L, He Q, Zhang Z, Chang C, Li X. Immunophenotypic analysis of erythroid dysplasia and its diagnostic application in myelodysplastic syndromes. Int Med J. 2012;42:401–11.
Mathis S, Chapuis N, Debord C, Rouquette A, Radford-Weiss I, Park S, et al. Flow cytometric detection of dyserythropoiesis: a sensitive and powerful diagnostic tool for myelodysplastic syndromes. Leukemia. 2013;27:1981–7.
Eidenschink Brodersen L, Menssen AJ, Wangen JR, Stephenson CF, de Baca ME, Zehentner BK, et al. Assessment of erythroid dysplasia by “difference from normal” in routine clinical flow cytometry workup. Cytom Part B, Clin Cytom. 2015;88:125–35.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, et al. Proposals for the classification of the myelodysplastic syndromes. Br J Haematol. 1982;51:189–99.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization classification of haematolymphoid tumours: myeloid and histiocytic/dendritic neoplasms. Leukemia. 2022;36:1703–19.
Greenberg P, Cox C, LeBeau MM, Fenaux P, Morel P, Sanz G, et al. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood. 1997;89:2079–88.
Greenberg PL, Tuechler H, Schanz J, Sanz G, Garcia-Manero G, Sole F, et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood. 2012;120:2454–65.
Machherndl-Spandl S, Suessner S, Danzer M, Proell J, Gabriel C, Lauf J, et al. Molecular pathways of early CD105-positive erythroid cells as compared with CD34-positive common precursor cells by flow cytometric cell-sorting and gene expression profiling. Blood Cancer J. 2013;3:e100.
Pannu KK, Joe ET, Iyer SB. Performance evaluation of QuantiBRITE phycoerythrin beads. Cytometry. 2001;45:250–8.
Kosulin K, Rauch M, Ambros PF, Potschger U, Chott A, Jager U, et al. Screening for adenoviruses in haematological neoplasia: High prevalence in mantle cell lymphoma. Euro J Cancer. 2014;50:622–7.
Ebner K, Suda M, Watzinger F, Lion T. Molecular detection and quantitative analysis of the entire spectrum of human adenoviruses by a two-reaction real-time PCR assay. J Clin Microbiol. 2005;43:3049–53.
Dietel M, Hafner N, Jansen L, Durst M, Runnebaum IB. Novel splice variant CAR 4/6 of the coxsackie adenovirus receptor is differentially expressed in cervical carcinogenesis. J Mol Med. 2011;89:621–30.
Hoermann G, Blatt K, Greiner G, Putz EM, Berger A, Herrmann H, et al. CD52 is a molecular target in advanced systemic mastocytosis. FASEB J Off Publ Fed Am Soc Exp Biol. 2014;28:3540–51.
Liu J, Zhang J, Ginzburg Y, Li H, Xue F, De Franceschi L, et al. Quantitative analysis of murine terminal erythroid differentiation in vivo: novel method to study normal and disordered erythropoiesis. Blood. 2013;121:e43–9.
Zhou T, Kinney MC, Scott LM, Zinkel SS, Rebel VI. Revisiting the case for genetically engineered mouse models in human myelodysplastic syndrome research. Blood. 2015;126:1057–68.
Wu G, Cheng, Zhang C. Membrane protein CAR promotes hematopoietic regeneration upon stress. Haematologica. 2021;106:2180–90.
Valent P, Busche G, Theurl I, Uras IZ, Germing U, Stauder R, et al. Normal and pathological erythropoiesis in adults: from gene regulation to targeted treatment concepts. Haematologica. 2018;103:1593–603.
Anders M, Vieth M, Rocken C, Ebert M, Pross M, Gretschel S, et al. Loss of the coxsackie and adenovirus receptor contributes to gastric cancer progression. Br J Cancer. 2009;100:352–9.
Matsumoto K, Shariat SF, Ayala GE, Rauen KA, Lerner SP. Loss of coxsackie and adenovirus receptor expression is associated with features of aggressive bladder cancer. Urology. 2005;66:441–6.
Huang KC, Altinoz M, Wosik K, Larochelle N, Koty Z, Zhu L, et al. Impact of the coxsackie and adenovirus receptor (CAR) on glioma cell growth and invasion: requirement for the C-terminal domain. Int J Cancer. 2005;113:738–45.
Pong RC, Roark R, Ou JY, Fan J, Stanfield J, Frenkel E, et al. Mechanism of increased coxsackie and adenovirus receptor gene expression and adenovirus uptake by phytoestrogen and histone deacetylase inhibitor in human bladder cancer cells and the potential clinical application. Cancer Res. 2006;66:8822–8.
Pong RC, Lai YJ, Chen H, Okegawa T, Frenkel E, Sagalowsky A, et al. Epigenetic regulation of coxsackie and adenovirus receptor (CAR) gene promoter in urogenital cancer cells. Cancer Res. 2003;63:8680–6.
Morton PE, Hicks A, Nastos T, Santis G, Parsons M. CAR regulates epithelial cell junction stability through control of E-cadherin trafficking. Sci Rep. 2013;3:2889.
Valent P, Orazi A, Steensma DP, Ebert BL, Haase D, Malcovati L, et al. Proposed minimal diagnostic criteria for myelodysplastic syndromes (MDS) and potential pre-MDS conditions. Oncotarget. 2017;8:73483–500.
Matarraz S, Lopez A, Barrena S, Fernandez C, Jensen E, Flores-Montero J, et al. Bone marrow cells from myelodysplastic syndromes show altered immunophenotypic profiles that may contribute to the diagnosis and prognostic stratification of the disease: a pilot study on a series of 56 patients. Cytom Part B Clin Cytom. 2010;78:154–68.
Chu SC, Wang TF, Li CC, Kao RH, Li DK, Su YC, et al. Flow cytometric scoring system as a diagnostic and prognostic tool in myelodysplastic syndromes. Leuk Res. 2011;35:868–73.
Alhan C, Westers TM, Cremers EM, Cali C, Witte BI, Ossenkoppele GJ, et al. High flow cytometric scores identify adverse prognostic subgroups within the revised international prognostic scoring system for myelodysplastic syndromes. Br J Haematol. 2014;167:100–9.
Alhan C, Westers TM, Cremers EM, Cali C, Witte BI, Ossenkoppele GJ, et al. The myelodysplastic syndromes flow cytometric score: a three-parameter prognostic flow cytometric scoring system. Leukemia. 2016;30:658–65.
Yan H, Ali A, Blanc L, Narla A, Lane JM, Gao E, et al. Comprehensive phenotyping of erythropoiesis in human bone marrow: Evaluation of normal and ineffective erythropoiesis. Am J Hematol. 2021;96:1064–76.
Acknowledgements
We like to thank Nadine Witzeneder, Barbara Peter, Martin Danzer, Harald Herrmann, and Hans-Ulrich Klein for their skillful technical assistance.
Funding
This study was supported by The Austrian Science Fund (FWF) SFB grants F4701-B20, F4704-B20, P30625-B28, by a grant of the Upper Austrian Cancer Aid Fund and by a stem cell grant of the Medical University of Vienna, Austria.
Author information
Authors and Affiliations
Contributions
KB and SMS designed the study and performed key laboratory experiments. LK, SG, AS and EK performed mouse experiments. WRS provided patients and statistical analyses. HUK contributed bioinformatics. SS, MD, and JP performed molecular and gene array studies. JL performed flow cytometry experiments. GE and IS performed transduction- and cell sorting experiments. GH provided laboratory data and molecular studies. NH provided modified pBK-CMV vectors containing different CXADR splice variants. MCB, UP, OZ, and AW contributed patients and logistic support. CG and GW performed molecular studies and provided logistic support. TL performed the adenovirus peptide-binding assay. UG contributed patients and logistic support. PV, PBS and PB designed the study and wrote the manuscript. All authors contributed by writing parts of the manuscript and by critically reading the paper. All authors approved the final version of the document.
Corresponding author
Ethics declarations
Competing interests
The authors declare no completing interests in this study. Conflict of interest outside the study are: P.V. received research grants from Celgene, Novartis, Incyte, Pfizer, and AOP Orphan, and honoraria from Celgene, Novartis, Pfizer, Incyte, and AOP Orphan. S.M.S received honoraria from Celgene, Novartis, Pfizer, Abbvie, Amgen, Jazz, Gilead, Servier and Incyte. P.B. was supported by Alexion outside the study. T.L. received research grants from Incyte and Novartis, and honoraria from Incyte, Pfizer, Angelini, Amgen and Chimerix. O.Z. was supported by Pfizer, Takeda, Amgen, Novartis, BMS, Genzyme, Sanofi, Pierre Fabre and Roche, and he is part of advisory boards of Amgen, Pfizer and Novartis. U.G. received research support from Novartis and Celgene, and honoraria from Celgene, Jazz, Novartis and Jansen. G.W. received research grants from Abbvie, Amgen, AstraZeneca, BMS, Böhringer Ingelheim, Gilead, Incyte, Lilly, Mundipharma, Novartis, Pfizer, Roche and Tesaro. G.H. received research grants from Novartis, he received honoraria and he is part of advisory boards of Novartis, Roche, Beckman Coulter, Pfizer, Celgene, Bristol-Myers Squibb. L.K. was supported by a DOCmed Fellowship of the Austrian Academy of Sciences (OeAW) and a research grant of the Vienna Comprehensive Cancer Center (CCC). P.B.S. reports grants and personal fees from Roche, Celgene, and AbbVie; and personal fees from Amgen, Takeda, AbbVie, and Janssen Cilag outside the submitted work.The other authors declare that they have no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bauer, K., Machherndl-Spandl, S., Kazianka, L. et al. CAR virus receptor mediates erythroid differentiation and migration and is downregulated in MDS. Leukemia 37, 2250–2260 (2023). https://doi.org/10.1038/s41375-023-02015-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-02015-7