Abstract
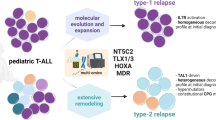
TAL1+ T-cell acute lymphoblastic leukemia (T-ALL) is a distinct subtype of leukemia with poor outcomes. Through the cooperation of co-activators, including RUNX1, GATA3, and MYB, the TAL1 oncoprotein extends the immature thymocytes with autonomy and plays an important role in the development of T-ALL. However, this process is not yet well understood. Here, by investigating the transcriptome and prognosis of T-ALL from multiple cohorts, we found that S1PR3 was highly expressed in a subset of TAL1+ T-ALL (S1PR3hi TAL1+ T-ALL), which showed poor outcomes. Through pharmacological and genetic methods, we identified a specific survival-supporting role of S1P-S1PR3 in TAL1+ T-ALL cells. In T-ALL cells, TAL1-RUNX1 up-regulated the expression of S1PR3 by binding to the enhancer region of S1PR3 gene. With hyperactivated S1P-S1PR3, T-ALL cells grew rapidly, partly by activating the KRAS signal. Finally, we assessed S1PR3 inhibitor TY-52156 in T-ALL patient-derived xenografts (PDXs) mouse model. We found that TY-52156 attenuated leukemia progression efficiently and extended the lifespan of S1PR3hi TAL1+ T-ALL xenografts. Our findings demonstrate that S1PR3 plays an important oncogenic role in S1PR3hi TAL1+ T-ALL and may serve as a promising therapeutic target.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in the paper are presented in the paper and/or the Supplementary Materials. The raw RNA-seq data reported in this paper, including T-ALL samples, HSPCs (lin−CD34+), and thymocytes derived from non-hematological malignant donors, were deposited in the Genome Sequence Archive database under accession number HRA002558.
References
Belver L, Ferrando A. The genetics and mechanisms of T cell acute lymphoblastic leukaemia. Nat Rev Cancer. 2016;16:494–507.
Homminga I, Pieters R, Langerak AW, de Rooi JJ, Stubbs A, Verstegen M, et al. Integrated transcript and genome analyses reveal NKX2-1 and MEF2C as potential oncogenes in T cell acute lymphoblastic leukemia. Cancer Cell. 2011;19:484–97.
Liu Y, Easton J, Shao Y, Maciaszek J, Wang Z, Wilkinson MR, et al. The genomic landscape of pediatric and young adult T-lineage acute lymphoblastic leukemia. Nat Genet. 2017;49:1211–8.
Ferrando AA, Neuberg DS, Staunton J, Loh ML, Huard C, Raimondi SC, et al. Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell. 2002;1:75–87.
Cordo V, van der Zwet JCG, Cante-Barrett K, Pieters R, Meijerink JPP. T-cell Acute Lymphoblastic Leukemia: A Roadmap to Targeted Therapies. Blood Cancer Discov. 2021;2:19–31.
Correia NC, Arcangeli ML, Pflumio F, Barata JT. Stem Cell Leukemia: how a TALented actor can go awry on the hematopoietic stage. Leukemia 2016;30:1968–78.
Wang D, Zhu G, Wang N, Zhou X, Yang Y, Zhou S, et al. SIL-TAL1 rearrangement is related with poor outcome: a study from a Chinese institution. PLoS ONE. 2013;8:e73865.
D’Angio M, Valsecchi MG, Testi AM, Conter V, Nunes V, Parasole R, et al. Clinical features and outcome of SIL/TAL1-positive T-cell acute lymphoblastic leukemia in children and adolescents: a 10-year experience of the AIEOP group. Haematologica 2015;100:e10–3.
Gerby B, Tremblay CS, Tremblay M, Rojas-Sutterlin S, Herblot S, Hebert J, et al. SCL, LMO1 and Notch1 reprogram thymocytes into self-renewing cells. PLoS Genet. 2014;10:e1004768.
Sanda T, Lawton LN, Barrasa MI, Fan ZP, Kohlhammer H, Gutierrez A, et al. Core transcriptional regulatory circuit controlled by the TAL1 complex in human T cell acute lymphoblastic leukemia. Cancer Cell. 2012;22:209–21.
Mansour MR, Abraham BJ, Anders L, Berezovskaya A, Gutierrez A, Durbin AD, et al. Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science. 2014;346:1373–7.
Kwiatkowski N, Zhang T, Rahl PB, Abraham BJ, Reddy J, Ficarro SB, et al. Targeting transcription regulation in cancer with a covalent CDK7 inhibitor. Nature 2014;511:616–20.
Choi A, Illendula A, Pulikkan JA, Roderick JE, Tesell J, Yu J, et al. RUNX1 is required for oncogenic Myb and Myc enhancer activity in T-cell acute lymphoblastic leukemia. Blood 2017;130:1722–33.
Ward AF, Braun BS, Shannon KM. Targeting oncogenic Ras signaling in hematologic malignancies. Blood 2012;120:3397–406.
Kortum RL, Rouquette-Jazdanian AK, Samelson LE. Ras and extracellular signal-regulated kinase signaling in thymocytes and T cells. Trends Immunol. 2013;34:259–68.
von Lintig FC, Huvar I, Law P, Diccianni MB, Yu AL, Boss GR. Ras activation in normal white blood cells and childhood acute lymphoblastic leukemia. Clin Cancer Res. 2000;6:1804–10.
Hartzell C, Ksionda O, Lemmens E, Coakley K, Yang M, Dail M, et al. Dysregulated RasGRP1 responds to cytokine receptor input in T cell leukemogenesis. Sci Signal. 2013;6:ra21.
Ksionda O, Melton AA, Bache J, Tenhagen M, Bakker J, Harvey R, et al. RasGRP1 overexpression in T-ALL increases basal nucleotide exchange on Ras rendering the Ras/PI3K/Akt pathway responsive to protumorigenic cytokines. Oncogene 2016;35:3658–68.
Wen Z, Yun G, Hebert A, Kong G, Ranheim EA, Finn R, et al. Nras Q61R/+ and Kras-/- cooperate to downregulate Rasgrp1 and promote lympho-myeloid leukemia in early T-cell precursors. Blood 2021;137:3259–71.
Van Thillo Q, De Bie J, Seneviratne JA, Demeyer S, Omari S, Balachandran A, et al. Oncogenic cooperation between TCF7-SPI1 and NRAS(G12D) requires beta-catenin activity to drive T-cell acute lymphoblastic leukemia. Nat Commun. 2021;12:4164.
Kindler T, Cornejo MG, Scholl C, Liu J, Leeman DS, Haydu JE, et al. K-RasG12D-induced T-cell lymphoblastic lymphoma/leukemias harbor Notch1 mutations and are sensitive to gamma-secretase inhibitors. Blood 2008;112:3373–82.
Karra L, Romero-Moya D, Ksionda O, Krush M, Gu Z, Mues M, et al. Increased baseline RASGRP1 signals enhance stem cell fitness during native hematopoiesis. Oncogene 2020;39:6920–34.
Cartier A, Hla T. Sphingosine 1-phosphate: Lipid signaling in pathology and therapy. Science. 2019;366:eaar5551.
Ogle ME, Olingy CE, Awojoodu AO, Das A, Ortiz RA, Cheung HY, et al. Sphingosine-1-Phosphate Receptor-3 Supports Hematopoietic Stem and Progenitor Cell Residence Within the Bone Marrow Niche. Stem Cells. 2017;35:1040–52.
Cyster JG, Schwab SR. Sphingosine-1-phosphate and lymphocyte egress from lymphoid organs. Annu Rev Immunol. 2012;30:69–94.
Vorbach S, Grunder A, Zhou F, Koellerer C, Jutzi JS, Simoni M, et al. Enhanced expression of the sphingosine-1-phosphate-receptor-3 causes acute myelogenous leukemia in mice. Leukemia 2020;34:721–34.
Xie SZ, Kaufmann KB, Wang W, Chan-Seng-Yue M, Gan OI, Laurenti E, et al. Sphingosine-1-phosphate receptor 3 potentiates inflammatory programs in normal and leukemia stem cells to promote differentiation. Blood Cancer Discov. 2021;2:32–53.
Pyne NJ, Pyne S. Sphingosine 1-phosphate and cancer. Nat Rev Cancer. 2010;10:489–503.
Shu Y, Wang Y, Lv WQ, Peng DY, Li J, Zhang H, et al. ARRB1-Promoted NOTCH1 Degradation Is Suppressed by OncomiR miR-223 in T-cell Acute Lymphoblastic Leukemia. Cancer Res. 2020;80:988–98.
Cramer-Morales K, Nieborowska-Skorska M, Scheibner K, Padget M, Irvine DA, Sliwinski T, et al. Personalized synthetic lethality induced by targeting RAD52 in leukemias identified by gene mutation and expression profile. Blood 2013;122:1293–304.
Wang L, Chen F, Liu R, Shi L, Zhao G, Yan Z. Gene expression and immune infiltration in melanoma patients with different mutation burden. BMC Cancer. 2021;21:379.
Salomone S, Waeber C. Selectivity and specificity of sphingosine-1-phosphate receptor ligands: caveats and critical thinking in characterizing receptor-mediated effects. Front Pharm. 2011;2:9.
Sanna MG, Wang SK, Gonzalez-Cabrera PJ, Don A, Marsolais D, Matheu MP, et al. Enhancement of capillary leakage and restoration of lymphocyte egress by a chiral S1P1 antagonist in vivo. Nat Chem Biol. 2006;2:434–41.
Koide Y, Hasegawa T, Takahashi A, Endo A, Mochizuki N, Nakagawa M, et al. Development of novel EDG3 antagonists using a 3D database search and their structure-activity relationships. J Med Chem. 2002;45:4629–38.
Murakami A, Takasugi H, Ohnuma S, Koide Y, Sakurai A, Takeda S, et al. Sphingosine 1-phosphate (S1P) regulates vascular contraction via S1P3 receptor: investigation based on a new S1P3 receptor antagonist. Mol Pharm. 2010;77:704–13.
Hnisz D, Weintraub AS, Day DS, Valton AL, Bak RO, Li CH, et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 2016;351:1454–8.
Leong WZ, Tan SH, Ngoc PCT, Amanda S, Yam AWY, Liau WS, et al. ARID5B as a critical downstream target of the TAL1 complex that activates the oncogenic transcriptional program and promotes T-cell leukemogenesis. Genes Dev. 2017;31:2343–60.
Calo E, Wysocka J. Modification of enhancer chromatin: what, how, and why? Mol Cell. 2013;49:825–37.
Moore AR, Rosenberg SC, McCormick F, Malek S. RAS-targeted therapies: is the undruggable drugged? Nat Rev Drug Discov. 2020;19:533–52.
Hunter JC, Manandhar A, Carrasco MA, Gurbani D, Gondi S, Westover KD. Biochemical and Structural Analysis of Common Cancer-Associated KRAS Mutations. Mol Cancer Res. 2015;13:1325–35.
Maeda S, Shiimura Y, Asada H, Hirata K, Luo F, Nango E, et al. Endogenous agonist-bound S1PR3 structure reveals determinants of G protein-subtype bias. Sci Adv. 2021;7:eabf5325.
Martz CA, Ottina KA, Singleton KR, Jasper JS, Wardell SE, Peraza-Penton A, et al. Systematic identification of signaling pathways with potential to confer anticancer drug resistance. Sci Signal. 2014;7:ra121.
Singh A, Greninger P, Rhodes D, Koopman L, Violette S, Bardeesy N, et al. A gene expression signature associated with “K-Ras addiction” reveals regulators of EMT and tumor cell survival. Cancer Cell. 2021;39:441–2.
Schwab SR, Pereira JP, Matloubian M, Xu Y, Huang Y, Cyster JG. Lymphocyte sequestration through S1P lyase inhibition and disruption of S1P gradients. Science 2005;309:1735–9.
Breart B, Ramos-Perez WD, Mendoza A, Salous AK, Gobert M, Huang Y, et al. Lipid phosphate phosphatase 3 enables efficient thymic egress. J Exp Med. 2011;208:1267–78.
Gordon WR, Roy M, Vardar-Ulu D, Garfinkel M, Mansour MR, Aster JC, et al. Structure of the Notch1-negative regulatory region: implications for normal activation and pathogenic signaling in T-ALL. Blood 2009;113:4381–90.
Giambra V, Jenkins CE, Lam SH, Hoofd C, Belmonte M, Wang X, et al. Leukemia stem cells in T-ALL require active Hif1alpha and Wnt signaling. Blood 2015;125:3917–27.
Yan Z, Xia J, Cao Z, Zhang H, Wang J, Feng T, et al. Multi-omics integration reveals potential stage-specific druggable targets in T-cell acute lymphoblastic leukemia. Genes & Diseases (in press).
Fischer AM, Katayama CD, Pages G, Pouyssegur J, Hedrick SM. The role of erk1 and erk2 in multiple stages of T cell development. Immunity 2005;23:431–43.
Coffield VM, Helms WS, Jiang Q, Su L. Galpha13 mediates a signal that is essential for proliferation and survival of thymocyte progenitors. J Exp Med. 2004;200:1315–24.
Acknowledgements
We thank the National Clinical Research Center for Child Health and Disorders for the use of their Shared Services (Flow Cytometry Laboratory, Animal Facility, and Experimental Pathology Laboratory) for completing this research. This study was supported by the National Natural Science Foundation of China (#81870126, #82070167, and #82270160 to LZ; #81900190 to YS) and The Chongqing Science and Technology Bureau Major Project (cstc2020jcyj-msxmX0782 to YS). We are grateful to Professors Bing Liu, Yufeng Shi, and Qing Lu for helping us to revise our paper and Figures. Finally, we are greatly indebted to the T-ALL patients and their families for their support and willingness to participate in the study.
Author information
Authors and Affiliations
Contributions
Conceptualization: LZ, YS, and JY. Methodology: DZ, TTJ, DYM, WQL, JZ, and YS. Investigation: MYG, HBW, ZYL, HYS, LMZ, and LZ. Data curation: HYZ, TTJ, and JY. Formal analysis: HYL, GF, DYP, BJY, and ZJY. Visualization: FIT, ZDL, LS, ZYC, and ST. Writing: YS, LZ, JY, TCH, ZJY, and HZ. All the authors have read and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, D., Jiang, T., Ma, D. et al. S1P-S1PR3-RAS promotes the progression of S1PR3hi TAL1+ T-cell acute lymphoblastic leukemia that can be effectively inhibited by an S1PR3 antagonist. Leukemia 37, 1982–1993 (2023). https://doi.org/10.1038/s41375-023-02000-0
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41375-023-02000-0
This article is cited by
-
Sphingosine 1-phosphate signalling in cancer stem cells
Oncogenesis (2025)