Abstract
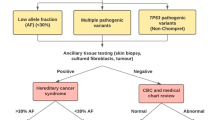
Myeloid malignancies associated with germline predisposition syndromes account for up to 10% of myeloid neoplasms. They are classified into three categories by the proposed 5th Edition of the World Health Organization Classification of Hematolymphoid Tumors: (1) neoplasms with germline predisposition without a pre-existing platelet disorder or organ dysfunction, (2) neoplasms with germline predisposition and pre-existing platelet disorder, or (3) neoplasms with germline predisposition and potential organ dysfunction. Recognizing these entities is critical because patients and affected family members benefit from interfacing with hematologists who specialize in these disorders and can facilitate tailored treatment strategies. However, identification of these syndromes in routine pathology practice is often challenging, as characteristic findings associated with these diagnoses at baseline are frequently absent, nonspecific, or impossible to evaluate in the setting of a myeloid malignancy. Here we review the formally classified germline predisposition syndromes associated with myeloid malignancies and summarize practical recommendations for pathologists evaluating a new myeloid malignancy diagnosis. Our intent is to empower clinicians to better screen for germline disorders in this common clinical setting. Recognizing when to suspect a germline predisposition syndrome, pursue additional ancillary testing, and ultimately recommend referral to a cancer predisposition clinic or hematology specialist, will ensure optimal patient care and expedite research to improve outcomes for these individuals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Murati A, Brecqueville M, Devillier R, Mozziconacci M-J, Gelsi-Boyer V, Birnbaum D. Myeloid malignancies: mutations, models and management. BMC Cancer. 2012;12:304.
Shallis RM, Wang R, Davidoff A, Ma X, Zeidan AM. Epidemiology of acute myeloid leukemia: recent progress and enduring challenges. Blood Rev. 2019;36:70–87.
SEER*Explorer. https://seer.cancer.gov/explorer/ Accessed 2 Aug 2022.
Volpe VO, Garcia-Manero G, Komrokji RS. Myelodysplastic syndromes: a new decade. Clin Lymphoma Myeloma Leuk. 2022;22:1–16.
Niemeyer CM. JMML genomics and decisions. Hematol Am Soc Hematol Educ Program. 2018;2018:307–12.
Gilad O, Dgany O, Noy-Lotan S, Krasnov T, Yacobovich J, Rabinowicz R, et al. Syndromes predisposing to leukemia are a major cause of inherited cytopenias in children. Haematologica. 2022;107:2081–95.
Feurstein S, Trottier AM, Estrada-Merly N, Pozsgai M, McNeely K, Drazer MW, et al. Germ line predisposition variants occur in myelodysplastic syndrome patients of all ages. Blood. 2022;140:2533–48.
Yang F, Long N, Anekpuritanang T, Bottomly D, Savage JC, Lee T, et al. Identification and prioritization of myeloid malignancy germline variants in a large cohort of adult patients with AML. Blood. 2022;139:1208–21.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia. 2022;36:1703–19.
Kanagal-Shamanna R. The emerging role of hematopathologists and molecular pathologists in detection, monitoring, and management of myeloid neoplasms with germline predisposition. Curr Hematol Malig Rep. 2021;16:336–44.
Weinberg OK, Kuo F, Calvo KR. Germline predisposition to hematolymphoid neoplasia. Am J Clin Pathol. 2019;152:258–76.
Tawana K, Brown AL, Churpek JE. Integrating germline variant assessment into routine clinical practice for myelodysplastic syndrome and acute myeloid leukaemia: current strategies and challenges. Br J Haematol. 2022;196:1293–310.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med J Am Coll Med Genet. 2015;17:405–24.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International consensus classification of myeloid neoplasms and acute leukemia: integrating morphological, clinical, and genomic data. Blood. 2022;140:1200–28. blood.2022015850
Tawana K, Wang J, Renneville A, Bödör C, Hills R, Loveday C, et al. Disease evolution and outcomes in familial AML with germline CEBPA mutations. Blood. 2015;126:1214–23.
Pabst T, Eyholzer M, Haefliger S, Schardt J, Mueller BU. Somatic CEBPA mutations are a frequent second event in families with germline CEBPA mutations and familial acute myeloid leukemia. J Clin Oncol. 2008;26:5088–93.
Tawana K, Rio-Machin A, Preudhomme C, Fitzgibbon J. Familial CEBPA-mutated acute myeloid leukemia. Semin Hematol. 2017;54:87–93.
Makishima H, Saiki R, Nannya Y, Korotev SC, Gurnari C, Takeda J, et al. Germline DDX41 mutations define a unique subtype of myeloid neoplasms. Blood. 2022. blood.2022018221.
Li P, Brown S, Williams M, White T, Xie W, Cui W, et al. The genetic landscape of germline DDX41 variants predisposing to myeloid neoplasms. Blood. 2022;140:716–55.
Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, et al. Inherited and Somatic Defects in DDX41 in Myeloid Neoplasms. Cancer Cell. 2015;27:658–70.
Churpek JE, Smith-Simmer K. DDX41-associated familial myelodysplastic syndrome and acute myeloid leukemia. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK574843/ (accessed 16 Aug2022).
Guha T, Malkin D. Inherited TP53 mutations and the Li-fraumeni syndrome. Cold Spring Harb Perspect Med. 2017;7:a026187.
Penkert J, Strüwe FJ, Dutzmann CM, Doergeloh BB, Montellier E, Freycon C, et al. Genotype-phenotype associations within the Li-Fraumeni spectrum: a report from the German Registry. J Hematol OncolJ Hematol Oncol. 2022;15:107.
Zebisch A, Lal R, Müller M, Lind K, Kashofer K, Girschikofsky M, et al. Acute myeloid leukemia with TP53 germ line mutations. Blood. 2016;128:2270–2.
Correa H. Li-fraumeni syndrome. J Pediatr Genet. 2016;5:84–88.
Tinat J, Bougeard G, Baert-Desurmont S, Vasseur S, Martin C, Bouvignies E, et al. 2009 version of the Chompret criteria for Li Fraumeni syndrome. J Clin Oncol J Am Soc Clin Oncol. 2009;27:e108–109. author reply e110.
Bougeard G, Renaux-Petel M, Flaman J-M, Charbonnier C, Fermey P, Belotti M, et al. Revisiting Li-Fraumeni syndrome from TP53 mutation carriers. J Clin Oncol J Am Soc Clin Oncol. 2015;33:2345–52.
Kratz CP, Freycon C, Maxwell KN, Nichols KE, Schiffman JD, Evans DG, et al. Analysis of the Li-Fraumeni Spectrum Based on an International Germline TP53 Variant Data Set: An International Agency for Research on Cancer TP53 Database Analysis. JAMA Oncol. 2021;7:1800–5.
Swaminathan M, Bannon SA, Routbort M, Naqvi K, Kadia TM, Takahashi K, et al. Hematologic malignancies and Li-Fraumeni syndrome. Cold Spring Harb Mol Case Stud. 2019;5:a003210.
Churpek JE, Lorenz R, Nedumgottil S, Onel K, Olopade OI, Sorrell A, et al. Proposal for the clinical detection and management of patients and their family members with familial myelodysplastic syndrome/acute leukemia predisposition syndromes. Leuk Lymphoma. 2013;54:28–35.
Schlegelberger B, Heller PG. RUNX1 deficiency (familial platelet disorder with predisposition to myeloid leukemia, FPDMM). Semin Hematol. 2017;54:75–80.
Bellissimo DC, Speck NA. RUNX1 mutations in inherited and sporadic leukemia. Front Cell Dev Biol. 2017;5:111.
Al-Harbi S, Aljurf M, Mohty M, Almohareb F, Ahmed SOA. An update on the molecular pathogenesis and potential therapeutic targeting of AML with t(8;21)(q22;q22.1);RUNX1-RUNX1T1. Blood Adv. 2020;4:229–38.
Brown AL, Arts P, Carmichael CL, Babic M, Dobbins J, Chong C-E, et al. RUNX1-mutated families show phenotype heterogeneity and a somatic mutation profile unique to germline predisposed AML. Blood Adv. 2020;4:1131–44.
Luddy RE, Champion LA, Schwartz AD. A fatal myeloproliferative syndrome in a family with thrombocytopenia and platelet dysfunction. Cancer. 1978;41:1959–63.
Stockley J, Morgan NV, Bem D, Lowe GC, Lordkipanidzé M, Dawood B, et al. Enrichment of FLI1 and RUNX1 mutations in families with excessive bleeding and platelet dense granule secretion defects. Blood. 2013;122:4090–3.
Jalagadugula G, Mao G, Kaur G, Goldfinger LE, Dhanasekaran DN, Rao AK. Regulation of platelet myosin light chain (MYL9) by RUNX1: implications for thrombocytopenia and platelet dysfunction in RUNX1 haplodeficiency. Blood. 2010;116:6037–45.
Kanagal-Shamanna R, Loghavi S, DiNardo CD, Medeiros LJ, Garcia-Manero G, Jabbour E, et al. Bone marrow pathologic abnormalities in familial platelet disorder with propensity for myeloid malignancy and germline RUNX1 mutation. Haematologica. 2017;102:1661–70.
Gaidzik VI, Teleanu V, Papaemmanuil E, Weber D, Paschka P, Hahn J, et al. RUNX1 mutations in acute myeloid leukemia are associated with distinct clinico-pathologic and genetic features. Leukemia. 2016;30:2160–8.
Noris P, Perrotta S, Seri M, Pecci A, Gnan C, Loffredo G, et al. Mutations in ANKRD26 are responsible for a frequent form of inherited thrombocytopenia: analysis of 78 patients from 21 families. Blood. 2011;117:6673–80.
Pippucci T, Savoia A, Perrotta S, Pujol-Moix N, Noris P, Castegnaro G, et al. Mutations in the 5’ UTR of ANKRD26, the ankirin repeat domain 26 gene, cause an autosomal-dominant form of inherited thrombocytopenia, THC2. Am J Hum Genet. 2011;88:115–20.
Sullivan MJ, Palmer EL, Botero JP. ANKRD26-related thrombocytopenia and predisposition to myeloid neoplasms. Curr Hematol Malig Rep. 2022. https://doi.org/10.1007/s11899-022-00666-4.
Al Daama SA, Housawi YH, Dridi W, Sager M, Otieno FG, Hou C, et al. A missense mutation in ANKRD26 segregates with thrombocytopenia. Blood. 2013;122:461–2.
Di Paola J, Porter CC. ETV6-related thrombocytopenia and leukemia predisposition. Blood. 2019;134:663–7.
Melazzini F, Palombo F, Balduini A, De Rocco D, Marconi C, Noris P, et al. Clinical and pathogenic features of ETV6-related thrombocytopenia with predisposition to acute lymphoblastic leukemia. Haematologica. 2016;101:1333–42.
Di Paola J, Fisher MH. ETV6-related thrombocytopenia and platelet dysfunction. Platelets. 2021;32:141–3.
Noetzli L, Lo RW, Lee-Sherick AB, Callaghan M, Noris P, Savoia A, et al. Germline mutations in ETV6 are associated with thrombocytopenia, red cell macrocytosis and predisposition to lymphoblastic leukemia. Nat Genet. 2015;47:535–8.
Nishii R, Baskin-Doerfler R, Yang W, Oak N, Zhao X, Yang W, et al. Molecular basis of ETV6-mediated predisposition to childhood acute lymphoblastic leukemia. Blood. 2021;137:364–73.
Spinner MA, Sanchez LA, Hsu AP, Shaw PA, Zerbe CS, Calvo KR, et al. GATA2 deficiency: a protean disorder of hematopoiesis, lymphatics, and immunity. Blood. 2014;123:809–21.
Katsumura KR, Bresnick EH. Group the GFM. The GATA factor revolution in hematology. Blood. 2017;129:2092–102.
Hsu AP, Johnson KD, Falcone EL, Sanalkumar R, Sanchez L, Hickstein DD, et al. GATA2 haploinsufficiency caused by mutations in a conserved intronic element leads to MonoMAC syndrome. Blood. 2013;121:3830–7. S1-7.
Johnson KD, Hsu AP, Ryu M-J, Wang J, Gao X, Boyer ME, et al. Cis-element mutated in GATA2-dependent immunodeficiency governs hematopoiesis and vascular integrity. J Clin Invest. 2012;122:3692–704.
Bresnick EH, Jung MM, Katsumura KR. Human GATA2 mutations and hematologic disease: how many paths to pathogenesis? Blood Adv. 2020;4:4584–92.
Ping N, Sun A, Song Y, Wang Q, Yin J, Cheng W, et al. Exome sequencing identifies highly recurrent somatic GATA2 and CEBPA mutations in acute erythroid leukemia. Leukemia. 2017;31:195–202.
Calvo KR, Vinh DC, Maric I, Wang W, Noel P, Stetler-Stevenson M, et al. Myelodysplasia in autosomal dominant and sporadic monocytopenia immunodeficiency syndrome: diagnostic features and clinical implications. Haematologica. 2011;96:1221–5.
Narumi S. Discovery of MIRAGE syndrome. Pediatr Int J Jpn Pediatr Soc. 2022;64:e15283.
Tanase-Nakao K, Olson TS, Narumi S MIRAGE Syndrome. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK564655/ (accessed 12 Sep2022).
Bluteau O, Sebert M, Leblanc T, Peffault de Latour R, Quentin S, Lainey E, et al. A landscape of germ line mutations in a cohort of inherited bone marrow failure patients. Blood. 2018;131:717–32.
Schwartz JR, Ma J, Lamprecht T, Walsh M, Wang S, Bryant V, et al. The genomic landscape of pediatric myelodysplastic syndromes. Nat Commun. 2017;8:1557.
Narumi S, Amano N, Ishii T, Katsumata N, Muroya K, Adachi M, et al. SAMD9 mutations cause a novel multisystem disorder, MIRAGE syndrome, and are associated with loss of chromosome 7. Nat Genet. 2016;48:792–7.
Buonocore F, Kühnen P, Suntharalingham JP, Del Valle I, Digweed M, Stachelscheid H, et al. Somatic mutations and progressive monosomy modify SAMD9-related phenotypes in humans. J Clin Invest 2017;127:1700–13.
Ahmed IA, Farooqi MS, Vander Lugt MT, Boklan J, Rose M, Friehling ED, et al. Outcomes of hematopoietic cell transplantation in patients with germline SAMD9/SAMD9L mutations. Biol Blood Marrow Transpl J Am Soc Blood Marrow Transpl. 2019;25:2186–96.
Chen D-H, Below JE, Shimamura A, Keel SB, Matsushita M, Wolff J, et al. Ataxia-pancytopenia syndrome is caused by missense mutations in SAMD9L. Am J Hum Genet. 2016;98:1146–58.
Gorcenco S, Komulainen-Ebrahim J, Nordborg K, Suo-Palosaari M, Andréasson S, Krüger J, et al. Ataxia-pancytopenia syndrome with SAMD9L mutations. Neurol Genet. 2017;3:e183.
Nagamachi A, Matsui H, Asou H, Ozaki Y, Aki D, Kanai A, et al. Haploinsufficiency of SAMD9L, an endosome fusion facilitator, causes myeloid malignancies in mice mimicking human diseases with monosomy 7. Cancer Cell. 2013;24:305–17.
Nagata Y, Narumi S, Guan Y, Przychodzen BP, Hirsch CM, Makishima H, et al. Germline loss-of-function SAMD9 and SAMD9L alterations in adult myelodysplastic syndromes. Blood. 2018;132:2309–13.
Tummala H, Walne A, Dokal I. The biology and management of dyskeratosis congenita and related disorders of telomeres. Expert Rev Hematol. 2022;15:685–96.
Alter BP, Giri N, Savage SA, Rosenberg PS. Cancer in the National Cancer Institute inherited bone marrow failure syndrome cohort after fifteen years of follow-up. Haematologica. 2018;103:30–9.
Kratz CP, Izraeli S. Down syndrome, RASopathies, and other rare syndromes. Semin Hematol. 2017;54:123–8.
Patnaik MM, Lasho TL. Genomics of myelodysplastic syndrome/myeloproliferative neoplasm overlap syndromes. Hematol Am Soc Hematol Educ Program. 2020;2020:450–9.
Dh G, Re F, Rh L, Br K, Pl W, Kj J. Neurofibromatosis type 1. Nat Rev Dis Primer. 2017; 3. https://doi.org/10.1038/nrdp.2017.4.
Roberts AE, Allanson JE, Tartaglia M, Gelb BD. Noonan syndrome. Lancet Lond Engl. 2013;381:333–42.
Tartaglia M, Gelb BD, Zenker M. Noonan syndrome and clinically related disorders. Best Pr Res Clin Endocrinol Metab. 2011;25:161–79.
Leardini D, Messelodi D, Muratore E, Baccelli F, Bertuccio SN, Anselmi L, et al. Role of CBL mutations in cancer and non-malignant phenotype. Cancers. 2022;14:839.
Niemeyer CM, Flotho C. Juvenile myelomonocytic leukemia: who’s the driver at the wheel? Blood. 2019;133:1060–70.
Kratz CP, Niemeyer CM, Castleberry RP, Cetin M, Bergsträsser E, Emanuel PD, et al. The mutational spectrum of PTPN11 in juvenile myelomonocytic leukemia and Noonan syndrome/myeloproliferative disease. Blood. 2005;106:2183–5.
Kratz CP, Niemeyer CM. Juvenile myelomonocytic leukemia. Hematol Amst Neth. 2005;10:100–3.
Niemeyer CM. RAS diseases in children. Haematologica. 2014;99:1653–62.
Boucher AC, Caldwell KJ, Crispino JD, Flerlage JE. Clinical and biological aspects of myeloid leukemia in Down syndrome. Leukemia. 2021;35:3352–60.
Roberts I, Alford K, Hall G, Juban G, Richmond H, Norton A, et al. GATA1-mutant clones are frequent and often unsuspected in babies with Down syndrome: identification of a population at risk of leukemia. Blood. 2013;122:3908–17.
Goemans BF, Noort S, Blink M, Wang Y-D, Peters STCJ, van Wouwe JP, et al. Sensitive GATA1 mutation screening reliably identifies neonates with Down syndrome at risk for myeloid leukemia. Leukemia. 2021;35:2403–6.
Pine SR, Guo Q, Yin C, Jayabose S, Druschel CM, Sandoval C. Incidence and clinical implications of GATA1 mutations in newborns with Down syndrome. Blood. 2007;110:2128–31.
Cunniff C, Bassetti JA, Ellis NA. Bloom’s syndrome: clinical spectrum, molecular pathogenesis, and cancer predisposition. Mol Syndromol. 2017;8:4–23.
Hafsi W, Badri T, Rice AS. Bloom Syndrome. In: StatPearls. StatPearls Publishing: Treasure Island (FL), 2022 http://www.ncbi.nlm.nih.gov/books/NBK448138/ Accessed 22 Nov 2022.
Thomas ERA, Shanley S, Walker L, Eeles R. Surveillance and treatment of malignancy in Bloom syndrome. Clin Oncol R Coll Radio G B. 2008;20:375–9.
Niraj J, Färkkilä A, D’Andrea AD. The fanconi anemia pathway in cancer. Annu Rev Cancer Biol. 2019;3:457–78.
Glanz A, Fraser FC. Spectrum of anomalies in Fanconi anaemia. J Med Genet. 1982;19:412–6.
West AH, Churpek JE. Old and new tools in the clinical diagnosis of inherited bone marrow failure syndromes. Hematol Am Soc Hematol Educ Program. 2017;2017:79–87.
Mehta PA, Ebens C Fanconi Anemia. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK1401/ Accessed 14 Sep 2022.
Nelson A, Myers K Shwachman-Diamond Syndrome. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK1756/ Accessed 14 Sep 2022.
Bezzerri V, Cipolli M. Shwachman-diamond syndrome: molecular mechanisms and current perspectives. Mol Diagn Ther. 2019;23:281–90.
Reilly CR, Shimamura A. Predisposition to myeloid malignancies in Shwachman-Diamond syndrome: Biological insights and clinical advances. Blood. 2022. blood.2022017739
Furutani E, Liu S, Galvin A, Steltz S, Malsch MM, Loveless SK, et al. Hematologic complications with age in Shwachman-Diamond syndrome. Blood Adv. 2022;6:297–306.
Da Costa L, Leblanc T, Mohandas N. Diamond-Blackfan anemia. Blood. 2020;136:1262–73.
Sieff C Diamond-Blackfan Anemia. In: Adam MP, Everman DB, Mirzaa GM, Pagon RA, Wallace SE, Bean LJ et al. (eds). GeneReviews®. University of Washington, Seattle: Seattle (WA), 1993 http://www.ncbi.nlm.nih.gov/books/NBK7047/ Accessed 14 Sep 2022.
Means RT. Pure red cell aplasia. Blood. 2016;128:2504–9.
Vlachos A, Rosenberg PS, Atsidaftos E, Alter BP, Lipton JM. Incidence of neoplasia in diamond blackfan anemia: a report from the diamond blackfan anemia registry. Blood. 2012;119:3815–9.
Rosenberg PS, Zeidler C, Bolyard AA, Alter BP, Bonilla MA, Boxer LA, et al. Stable long-term risk of leukaemia in patients with severe congenital neutropenia maintained on G-CSF therapy. Br J Haematol. 2010;150:196–9.
Dale DC, Bolyard AA, Schwinzer BG, Pracht G, Bonilla MA, Boxer L, et al. The severe chronic neutropenia international registry: 10-year follow-up report. Support Cancer Ther. 2006;3:220–31.
Acknowledgements
We would like to thank Dr. Catherine Leith, Dr. Erik Ranheim, and Dr. David Yang for helpful discussions and comments.
Funding
This work was supported by National Institute of Health (K08 DK127244 and T32 HL007899) and The Hartwell Foundation to DRM, and by the Evans Foundation to JEC.
Author information
Authors and Affiliations
Contributions
DRM, EFR, VLH, RKS, and JEC reviewed the literature; DRM, EFR, and JEC developed and wrote the manuscript; JDR, VLH, RKS, ATP, and ALS provided feedback and edited manuscript.
Corresponding author
Ethics declarations
Competing interests
JEC receives honoraria from UpToDate, Inc related to an educational article on hereditary hematologic malignancies. VHL is an employee and shareholder of Exact Sciences. The authors otherwise declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Reinig, E.F., Rubinstein, J.D., Patil, A.T. et al. Needle in a haystack or elephant in the room? Identifying germline predisposition syndromes in the setting of a new myeloid malignancy diagnosis. Leukemia 37, 1589–1599 (2023). https://doi.org/10.1038/s41375-023-01955-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-023-01955-4
This article is cited by
-
Genomic testing for germline predisposition to hematologic malignancies
Blood Research (2024)