Abstract
The variable clinical course of follicular lymphoma (FL) is determined by the molecular heterogeneity of tumor cells and complex interactions within the tumor microenvironment (TME). IL-4 producing follicular helper T cells (TFH) are critical components of the FL TME. Binding of IL-4 to IL-4R on FL cells activates JAK/STAT signaling. We identified STAT6 mutations (STAT6MUT) in 13% of FL (N = 33/258), all clustered within the DNA binding domain. Gene expression data and immunohistochemistry showed upregulation of IL-4/STAT6 target genes in STAT6MUT FL, including CCL17, CCL22, and FCER2 (CD23). Functionally, STAT6MUT was gain-of-function by serial replating phenotype in pre-B CFU assays. Expression of STAT6MUT enhanced IL-4 induced FCER2/CD23, CCL17 and CCL22 expression and was associated with nuclear accumulation of pSTAT6. RNA sequencing identified PARP14 -a transcriptional switch and co-activator of STAT6- among the top differentially upregulated genes in IL-4 stimulated STAT6MUT lymphoma cells and in STAT6MUT primary FL cells. Quantitative chromatin immunoprecipitation (qChIP) demonstrated binding of STAT6MUT but not STAT6WT to the PARP14 promotor. Reporter assays showed increased IL-4 induced transactivation activity of STAT6MUT at the PARP14 promotor, suggesting a self-reinforcing regulatory circuit. Knock-down of PARP14 or PARP-inhibition abrogated the STAT6MUT gain-of-function phenotype. Thus, our results identify PARP14 as a novel therapeutic target in STAT6MUT FL.
Similar content being viewed by others
Introduction
Follicular lymphoma (FL) is among the most common malignant lymphomas worldwide. Most patients present with advanced stage disease and are still considered incurable [1]. Although considered the prototype of indolent lymphoma, FL is a highly heterogenous disease and a subset of patients has early treatment failure, aggressive clinical course and remarkably short overall survival [2, 3]. Treatment of relapsed, refractory or transformed FL remains a major clinical challenge [4]. Molecular-targeted therapies hold promise to improve treatment outcome if tailored toward the individual disease biology of distinct patient subsets.
The molecular hallmark of FL is the rearrangement of chromosome 18q21 resulting in overexpression of anti-apoptotic BCL2 [5]. Dozens of additional recurrent gene mutations have been reported in FL, many of which are likely to have distinct implications on the biology and the clinical course of the disease (reviewed in [6]). In addition to tumor cell intrinsic alterations, features of the tumor microenvironment (TME) have also been shown to be associated with treatment outcome [7,8,9,10,11].
FL cells retain a remarkable dependency on the TME, orchestrating a tumor-permissive immune niche (reviewed in [12]). The FL TME is enriched with T follicular helper (TFH) cells which express higher levels of IL-4 compared to TFH cells from healthy lymph nodes [13, 14]. Binding of IL-4 to its receptor (IL-4R) on lymphoma cells recruits JAK1/3 and activates STAT6 by phosphorylation of Y641. This promotes the formation of STAT6 homodimers, which translocate to the nucleus and bind to regulatory elements of STAT6 target genes via the DNA-binding domain. In the presence of IL-4, transcriptional repressors are released (e.g., HDAC2 and HDAC3) and transcriptional coactivators are recruited (e.g., EP300, NCOA1 and NCOA2) in a process that involves PARP14, a member of the poly ADP-ribose polymerases (PARP) family, thereby forming the STAT6 enhanceosome that drives STAT6-dependent gene expression [9, 15,16,17].
Others and we have previously reported that STAT6 is recurrently and significantly mutated in FL and other B cell lymphomas [18,19,20,21,22,23]. In-vitro studies and crystal structure analyses have suggested that these mutations might enhance the ability of STAT6 to bind to canonical DNA binding sites of STAT6 [23, 24]. However, the underlying mechanism of how STAT6 mutations (STAT6MUT) contribute to FL biology remains incompletely understood. Moreover, potential therapeutic vulnerabilities have not yet been explored. Here we describe a therapeutically targetable PARP14-mediated self-reinforcing regulatory circuit that amplifies IL-4 induced STAT6-dependent gene expression in STAT6MUT lymphoma cells.
Materials and methods
Cell lines and reagents
OCI-Ly1and OCI-Ly8 cells were cultured in IMDM (PAN Biotech, Aidenbach, Germany). 293 T HEK cells and HeLa cells were cultured in Dulbecco’s Modified Eagle Medium. All cells were cultured with 10% FBS (PAN), 37 °C 5% CO2. Cell lines were authenticated by short tandem repeat analysis (Eurofins, Val Fleuri, Luxembourg) and tested negative for mycoplasma by PCR. Of note, we confirmed that OCI-Ly1 harbors a variant (G375R) in the STAT6 DNA-binding domain [23] that, to the best of our knowledge, has not been reported in any other cell line or primary tumor sample. Cells were stably transduced with a CMV-driven cDNA expression construct (pHAGE-CMV-MCS-IRES-ZsGreen; PlasmID, EvNO00061605) encoding for Flag-tagged mutant (MUT) STAT6 (D419G, D419N, N421K, or D519V) or wild type (WT) STAT6 (PlasmID, HsCD00365550) as previously described [7], and stimulated with human recombinant IL-4 (Miltenyi Biotec, Cologne, Germany) as indicated.
Human ex vivo FL-like co-culture model
Briefly, we isolated germinal center (GC) cells from human tonsils and immortalized them by transduction of BCL2 and BCL6 as previously described [25]. In addition, we stably expressed STAT6WT, STAT6D419G or EV control. These FL-like cells were grown on the follicular dendritic cell (FDC) feeder cell line YK6-CD40lg-IL21, which stably expresses CD40L and IL-21.
Immunoblotting
Western blot and immunoprecipitation (IP) experiments were performed as previously described [7]. Technical details and antibodies are listed in the supplementary methods. Subcellular fractions were prepared using the Qproteome Nuclear Protein Kit (Qiagen, Hilden, Germany).
Pre-B CFU assay
Emu-BCL2 (B6.Cg-Tg(BCL2)36Wehi/J) mice were sacrificed and femurs were flushed with phosphate buffered saline (PBS). 1 × 106 bone marrow cells were retrovirally transduced with STAT6WT, STAT6D419G or empty vector (EV) (all cloned into pMSCV-IRES-GFP) as previously [7] described, incubated for 4 h (5% CO2, 37 °C) and plated onto methylcellulose (0.3 × 106 cells/mL) that supports the growth of pre-B colony-forming-units (CFUs) (M3630, Stem Cell Technologies, Vancouver, British Columbia, Canada). Pre-B CFUs were counted according to the manufacturer’s guidelines. Cells from each plate were washed off and replated entirely onto fresh M3630 methylcellulose media every 7 days.
RNA sequencing
OCI-Ly1 cells stably expressing either STAT6WT (N = 9) or STAT6MUT (D419G, D419N, N421K, each N = 3) were stimulated with IL-4 (10 ng/ml for 20 min), washed and replated in IL-4 free media. RNA sequencing libraries were prepared from mRNA isolated at 2, 4, and 8 h (Direct-zol RNA MiniPrep Plus, Zymo Research, Irvine, California, USA), respectively, using the prime-seq protocol as previously described and sequenced on an Illumina HiSeq 1500 system [26]. Raw sequence data was processed following the Drop-seq data pipeline. Differential gene expression analysis was performed using DESeq2 package [27]. Raw and processed RNA sequencing data have been made available via the Gene-Expression-Omnibus platform under the accession number GSE208031.
We re-analyzed previously published single cell RNA sequencing data [28] from eight primary FL with known STAT6 genotypes using the R package Seurat (version 3) [29].
Proximity ligation assay (PLA)
Briefly, 2 µm cytoblock sections of OCI-Ly8 cells (STAT6WT vs STAT6MUT with or without IL-4 stimulation [10 ng/ml for 20 min]) were co-stained with mouse anti-STAT6 (LSBio, LS-B6154, 1:800) and rabbit anti-PARP14 (Sigma, HPA012063, 1:100). Images were acquired on a Vectra Polaris imaging system using inForm automated image analysis software (Akoya). PLA spots (TexasRed channel) per cell were counted in five representative view fields (40×) using HALO Image Analysis Platform version 3.2.
Luciferase reporter assay
PARP14 was cloned into pGL3 basic vector as detailed in the supplement following the manufacturer’s guidelines (Promega, Madison, Wisconsin, USA). The PARP14-pGL3 vector (200 ng) was co-transfected with pRL-CMV Renilla luciferase control reporter (40 ng, Promega) and 10 or 50 ng STAT6 expression vector (pHAGE CMV-STAT6MUT vs pHAGE-CMV-STAT6WT) into 293 T HEK cells using ViaFectTM (Promega). After 24 h, cells were stimulated with IL-4 (10 ng/ml) for 6 h and analyzed by Dual-Glo® Luciferase Assay (Promega). The PARP14 knock-down experiment was performed in HeLa cells since these express high levels of endogenous PARP14. Two shRNA constructs were used to knock-down PARP14 (SHC016-1EA, TRCN0000053158, TRCN0000053159, Sigma Aldrich, St. Louis, Missouri, USA), a non-targeting shRNA construct was used as control (see supplementary methods). PJ34 (Selleckchem, Munich, Germany) was used as a PARP inhibitor (50 µM, 30 min prior to IL-4 stimulation). Viability was measured by Vi-Cell XR, cell viability analyzer (Beckman Coulter).
Quantitative chromatin immunoprecipitation (qChIP)
qChIP was performed in OCI-Ly8 expressing STAT6WT, STAT6D419G or EV stimulated with IL-4 (10 ng/ml for 24 h). Cells were washed in PBS and double crosslinked with EGS (Sigma Aldrich) and formaldehyde (1%, Thermo Fisher Scientific, Waltham, Massachusetts, USA). 15 µg of sheared chromatin was used for each IP of 3x Flag-tagged STAT6 (STAT6WT or STAT6D419G), H3 or IgG as previously described [30].
Additional methods are provided in the supplement.
Results
STAT6 mutations cluster within the DNA binding domain
In a cohort of 258 patients with advanced stage FL, we identified 35 STAT6 mutations in 33 diagnostic biopsies (13%; Fig. 1A). All mutations clustered within the DNA binding domain, mostly at D419 (N = 16, 43%). In addition, 3 patients harbored a known (yet rare) polymorphic D419N variant at this position (rs11172102, Fig. 1A). The variant allele frequencies (VAF) of two of these cases were suggestive of somatically acquired mutations (Supplementary Fig. 1A). Two FL harbored two mutations each, both located within the DNA binding domain, respectively. Manual review of the sequencing reads spanning the mutations demonstrated that these mutations were located in cis on the same allele (Supplementary Fig. 2A).
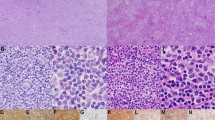
A Lolliplot of STAT6 mutations in 258 human FL biopsies. B Unsupervised hierarchical clustering of gene expression data from 106 human FL, annotated for STAT6 genotype. C Same data used for gene set enrichment analysis (GSEA) for established IL-4 signatures comparing STAT6WT and STAT6MUT. D Volcano plot of differentially expressed genes between STAT6WT and STAT6MUT FL (N = 106, log2FC = ±0.65, p value = 0.05). E FCER2 (CD23) gene expression in human FL with wild type STAT6 (STAT6WT) vs any STAT6 mutation (STAT6MUT) vs STAT6 mutations at position D419 (STAT6D419). F Representative immunohistochemistry (IHC) stains for CD23 and pSTAT6 in human FL with STAT6WT (left) and STAT6MUT (STAT6D419G, right).
STAT6 MUT FL are enriched for IL-4 linked gene expression
To identify gene expression patterns associated with STAT6MUT, we re-analyzed the available genome-wide RNA profiling data from 106 diagnostic biopsies from our cohort, comprising 17 and 89 cases with and without STAT6MUT, respectively (details provided in Supplementary Methods). Unsupervised hierarchical clustering distinguished two groups that were enriched for the respective STAT6 genotypes (MUT vs WT, p < 0.0001; Fig. 1B). Gene set enrichment analysis (GSEA) demonstrated that STAT6MUT cases were significantly enriched for two previously described IL-4 gene expression signatures [31, 32] (Fig. 1C). The top differentially expressed genes included known IL-4 regulated genes such as CCL17, CCL22, and FCER2 (Fig. 1D), which encodes for the low-affinity cell surface receptor for IgE (FcεRII), alias CD23.
We validated increased FCER2 gene expression in STAT6MUT FL in another 138 diagnostic biopsies from the GLSG cohort with known mutation profile and available digital multiplexed gene expression data (nCounter, NanoString) (Fig. 1E). To validate enhanced CD23 protein expression and STAT6 pathway activation in STAT6MUT FL, we performed immunohistochemistry (IHC) of primary patient samples. We indeed observed increased CD23 expression in STAT6MUT FL compared to STAT6WT FL (Fig. 1F, left panel). Moreover, we confirmed that pSTAT6 positive lymphoma cells formed prominent clusters in STAT6MUT FL in the vicinity of IL-4 producing TFH cells (Fig. 1F, right panel) as previously described [33].
DNA binding site mutations are gain-of-function and require IL-4 for increased STAT6 activation
To functionally test whether STAT6 DNA binding site mutations are gain-of-function, we utilized a modified pre-B colony-formation-unit (CFU) assay and analyzed the serial replating capacity. Specifically, bone marrow cells from mice expressing a BCL2-transgene in B-cells only (Emu-BCL2) were retrovirally transduced with STAT6WT, STAT6D419G or EV control and plated on methylcellulose supplemented with cytokines that support the growth of pre-B CFUs plus additional mouse IL-4 (mIL-4). Beyond passage 3, only STAT6D419G conferred a serial replating phenotype (Fig. 2A).
A Hematopoietic stem and progenitor cells (HSPCs) from Emu-BCL2 mouse cells transduced with either STAT6WT, STAT6D419G or empty vector (EV) and serially replated on cytokine-supplemented methylcellulose (MethoCult, M3630) that supports the growth of mouse pre-B colony-forming units (CFUs) plus additional mouse IL-4 (mIL-4) (N = 3, mean ± SD). B FCER2 mRNA levels (by qPCR) in OCI-Ly1 and OCI-Ly8 cells expressing either STAT6WT, STAT6D419G or EV control after IL-4 stimulation (10 ng/mL, 24 h; 2−dCt values relative to STAT6WT, N = 3, mean ± SD). C CD23 cell surface expression (by FACS) on OCI-Ly1 and OCI-Ly8 expressing either STAT6WT, STAT6D419G or EV control after IL-4 stimulation (10 ng/mL, 24 h; geometric mean, N = 3, mean ± SD). D Soluble CD23 (sCD23) levels (by ELISA) in cell culture supernatants of OCI-Ly1 and OCI-Ly8 cells expressing either STAT6WT, STAT6D419G or EV control after IL-4 stimulation (10 ng/mL, 72 h by ELISA (N = 3, mean ± SD). E Immunoblots of subcellular fractions (cytoplasmic vs nuclear) of OCI-Ly1 and F OCI-Ly8 cells expressing either STAT6WT or STAT6D419G. “−“ indicates no IL-4 stimulation, “+” indicates IL-4 stimulation (10 ng/mL for 20 min), and “P” indicates IL-4 pulse stimulation (IL-4 10 ng/mL for 20 min, then wash & withdrawal of IL-4 and incubation for another 8 h in fresh media without IL-4). G Representative immunohistochemistry (IHC) stain for phosphorylated STAT6 (pSTAT6) in OCI-Ly1 cells expressing STAT6WT or STAT6D419G with or without IL-4 pulse stimulation.
To study the molecular mechanisms of this IL-4 induced gain-of-function phenotype, we stably expressed STAT6MUT (D419G, D419N, N421K, or D519V), STAT6WT or EV control in OCI-Ly1 and OCI-Ly8 cells, two B cell lymphoma cell lines that harbor the FL hallmark translocation t(14;18). Upon IL-4 stimulation, we observed significantly increased FCER2 gene expression in cells stably expressing STAT6D419G (Fig. 2B), STAT6D419N, STAT6N421K, or STAT6D519V (Supplementary Fig. 3A) as compared to STAT6WT. Likewise, IL-4 induced expression of CD23 on cell surfaces and soluble CD23 (sCD23) in cell supernatants was significantly increased in STAT6MUT vs STAT6WT cells (Fig. 2C, D). Importantly, we did not see any differences of STAT6MUT vs STAT6WT in the absence of IL-4 (Fig. 2B–D, Supplementary Fig. 3A–C).
DNA binding site mutations promote aberrant IL-4 induced nuclear accumulation of pSTAT6
To study the mechanism of increased IL-4 induced transcriptional activation of STAT6MUT, we determined pSTAT6 levels in cytoplasmic and nuclear fractions. Following IL-4 stimulation, we observed higher nuclear pSTAT6 levels in cells expressing STAT6D419G (Fig. 2E, F) or STAT6D419N (Supplementary Fig. 4A) as compared to STAT6WT. Accordingly, cytoplasmic fractions were more depleted of pSTAT6 in STAT6MUT vs STAT6WT cells. Total STAT6 and pSTAT6 levels in whole cell lysates were not different (Supplementary Fig. 4B, C).
To determine the STAT6 signaling kinetics, we stimulated OCI-Ly1 and OCI-Ly8 cells with IL-4 for 20 min only, and then cultured the cells for an additional 8 h in the absence of IL-4 (“pulse stimulation” (P)), modeling the transient exposure of FL cells to IL-4 within the dynamic TME. Following IL-4 pulse stimulation, we detected higher levels of nuclear pSTAT6 in STAT6MUT cells as compared to STAT6WT cells (Fig. 2E, F, Supplementary Fig. 4C). Similar results were obtained when these cells were analyzed for pSTAT6 by IHC (Fig. 2G). IL-4 pulse simulation also led to significantly enhanced expression of FCER2 mRNA and membrane-bound CD23 in STAT6MUT vs STAT6WT cells (Supplementary Fig. 3D, E). Thus, DNA binding site mutations in STAT6 lead to increased accumulation of pSTAT6 within the nucleus, and increased transcription and expression of STAT6 target genes.
PARP14 is strongly upregulated in IL-4 stimulated STAT6MUT cells
We performed RNA sequencing of OCI-Ly1 cells expressing STAT6MUT (D419G, D419N, or N421K, N = 3 each) or STAT6WT (N = 9) at different time points following IL-4 stimulation (2, 4, and 8 h, respectively; Supplementary Fig. 5A). We selected OCI-Ly1 cells because of their marked responsiveness to IL-4. Principal components analysis revealed tight clustering of STAT6D419G, STAT6D419N, and STAT6N421K, mirroring the highly similar gain-of-function phenotype across the different mutations. This justified pooling of data from all STAT6MUT cells for comparative analyses. In contrast, STAT6WT was clearly separated from STAT6MUT by PC2 (Supplementary Fig. 5B). Transcriptional changes over time were mostly represented by shifts in PC1 (Supplementary Fig. 5B). The expression levels of 54 genes were significantly different across all analyzed time points. Differential gene expression was skewed toward upregulation in STAT6MUT vs STAT6WT, both by fold change and number of genes (Fig. 3A). Supplementary Fig. 5C shows the top differentially expressed genes for each time point and unsupervised clustering robustly separated STAT6MUT from STAT6WT cells. Reassuringly, we found FCER2 to be differentially upregulated at the 8 h time point (log2 fold change 0.65, p = 0.0007). Other known STAT6 target genes such as IRF4 and SOCS1 were also significantly upregulated in STAT6MUT vs STAT6WT cells at one or more time points. Of note, Poly(ADP-Ribose) Polymerase Family Member 14 (PARP14) was among the top most differentially upregulated genes across all time points (Fig. 3A, Supplementary Fig. 5C). We confirmed that IL-4 induced increased PARP14 expression by qPCR (Fig. 3B) and by Western blot (Fig. 3C) in STAT6MUT cells, whereas levels were not different in STAT6WT cells. PARP14 has been reported to function as a transcriptional switch for STAT6-dependent gene activation [16]. Specifically, in the presence of IL-4 the catalytic activity of PARP14 facilitates STAT6 binding to the promoter, and release of transcriptionally repressive HDACs. We could indeed confirm direct interaction of PARP14 and STAT6 in IL-4 stimulated lymphoma cells by IP of 3xFlag-tagged STAT6 and immunoblotting for PARP14 (Fig. 3D). Furthermore, we performed proximity ligation assays (PLA) in OCI-Ly8 cells and showed significantly increased STAT6-PARP14 interaction upon IL-4 stimulation (Supplementary Fig. 6) and in IL-4 stimulated STAT6D419G cells as compared to STAT6WT cells (Fig. 3E).
A Volcano plot of differentially expressed genes from whole transcriptome sequencing of OCI-Ly1 cells expressing STAT6WTor STAT6MUT (D419G, D419N, N421K) across all time points (i.e., 2, 4, and 8 h, respectively). Dashed lines indicate log2FC ±1.0, adj. p value 0.0001. B PARP14 expression validated by quantitative PCR (qPCR) in OCI-Ly1 cells. C Immunoblot analysis of PARP14 protein expression in OCI-Ly1 and OCI-Ly8 expressing STAT6WT or STAT6D419G after IL-4 stimulation (10 ng/mL, 24 h). D Immunoprecipitation of 3xFlag-tagged STAT6 and immunoblotting for PARP14 and STAT6 (3xFlag) in OCI-LY8 after IL-4 stimulation (10 ng/mL, 24 h). E Proximity ligation assay (PLA) of STAT6 and PARP14. Representative images of OCI-Ly8 cells expressing STAT6WT or STAT6D419G after IL-4 stimulation (left); TexasRed channel used to detect red amplification signal. Box plots display the percentage of TexasRed positive cells per total cell number for each analyzed microscopic view field (N = 5, right). F Single-cell RNA sequencing analysis. Violin plot showing PARP14 expression in tumor cells from 8 FL patients with wild type STAT6 (STAT6WT, N = 6) vs any STAT6 mutation (MUT, N = 2) vs STAT6 mutation at position D419 (D419MUT, N = 1).
PARP14 is significantly upregulated in STAT6 MUT tumor cells in human FL
To test whether that PARP14 is indeed upregulated in STAT6MUT human FL, we analyzed single cell RNA sequencing data from human FL biopsies with known mutation status (N = 8, Supplementary Fig. 7) from Haebe et al. [28]. In fact, tumor cells from FLs harboring any STAT6MUT (N = 2), and STAT6D419G in particular (N = 1), had significantly higher PARP14 expression compared to STAT6WT tumor cells (Fig. 3F). Of note, this difference in PARP14 expression was only seen in a subpopulation of tumor cells, consistent with our hypothesis that this phenotype is restricted to FL cells that harbor the STAT6 mutation and are exposed to TME-derived IL-4, i.e., localized in spatial proximity to IL-4 producing TFH cells.
Validation in a human ex vivo FL-like co-culture system
We wanted to validate our findings in an ex vivo model that more closely resembles human FL. Therefore, we immortalized human tonsil-derived GC B cells by stably expressing BCL2 and BCL6, along with either STAT6WT, STAT6D419G or EV (Fig. 4A, B). Importantly, these cells maintain a GC phenotype (Fig. 4C) and absolutely require FDC support plus IL21 and CD40L (YK6-CD40lg-IL21) for sustained growth, mirroring the TME-dependence of FL.
A Schematic of a fully human FL-like model system: human tonsil-derived germinal center (GC) B cells expressing BCL2 and BCL6 as well as STAT6WT or STAT6D419G or EV are co-cultured with follicular dendritic cell (FDC) feeder cells that express CD40L and IL-21 (YK6-CD40lg-IL-21). B Western Blot of FL-like cells, whole cell lysates. C Flow cytometry of GC markers on FL-like cells with indicated STAT6 genotypes. D Western Blot of FL-like cells, cellular and nuclear fraction. E qPCR analysis of FCER2, CCL17, and CCL21, as well as F PARP14 in FL-like cells with indicated STAT6 genotypes with and without IL-4 stimulation.
Using this FL-like co-culture we could indeed confirm our previous findings, including increased IL-4 induced nuclear levels of pSTAT6 and PARP14 in STAT6D419G cells compared to STAT6WT cells (Fig. 4D), significantly increased IL-4 induced gene expression of the known STAT6 target genes FCER2, CCL17, and CCL22 (Fig. 4E), as well as significantly increased PARP14 expression only in IL-4 stimulated STAT6D419G cells (Fig. 4F).
PARP14 per se is a novel target of mutant STAT6
We hypothesized that PARP14 per se could be a novel (aberrant) target gene of STAT6MUT, but not of STAT6WT. To directly assess and quantify its binding to the PARP14 promotor, we performed cross-linked chromatin IP for 3x Flag of IL-4 stimulated lymphoma cells (OCI-Ly8) stably expressing either 3x Flag-tagged STAT6MUT (D419G) or 3xFlag-tagged STAT6WT followed by quantitative PCR for the PARP14 promotor region. This indeed demonstrated increased binding of STAT6D419G to the PARP14 promoter, whereas STAT6WT was not different compared to EV control (Fig. 5A). We identified several putative STAT6 binding sites in the PARP14 promotor by bioinformatic prediction (Fig. 5B). To functionally validate our finding of increased binding of STAT6MUT to the PARP14 promotor, we cloned a 621 bp fragment containing two potential STAT6 binding sites in close proximity to the PARP14 start codon into a luciferase reporter construct. The reporter construct was then expressed in HEK 293 T cells, along with either STAT6D419G or STAT6WT. We used HEK 293 T cells as they express the IL-4 receptor but no endogenous STAT6. In the presence of IL-4, STAT6D419G indeed showed significantly increased transactivation activity compared to STAT6WT (Fig. 5C) in a dose-dependent manner.
A Cross-linked chromatin immunoprecipitation (ChIP) of 3xFlag-tagged STAT6WT or STAT6D419G in OCI-LY8 cells after IL-4 stimulation (10 ng/mL, 24 h) followed by quantitative PCR for the PARP14 promoter region (qChIP). B Schematic of the PARP14 promotor region, indicating predicted / putative STAT6 binding sites and the 641 bp region that was cloned into pGL3. C PARP14 promoter luciferase assay (pGL3) with increasing amounts of co-transfected STAT6WT or STAT6D419G in 293 T cells in the presence of IL-4 (24 h after transfection, 10 ng/mL for 6 h). Shown are fold changes (FC) of luciferase activity (relative light unit, RLU) normalized to 1 ng STAT6WT (N = 6, mean ± SD). D CD23 cell surface expression (by FACS) on OCI-Ly1 and E OCI-Ly8, each expressing either STAT6WT or STAT6D419G with shRNA-mediated knock-down of PARP14 (sh4) or a non-targeting (scrambled) control (scr), respectively. Immunoblot of respective cells as indicated below. F CD23 cell surface expression by FACS on OCI-Ly1 and OCI-L8 expressing STAT6WT or STAT6D419G with or without IL-4 stimulation (10 mg/mL, 24 h) and treatment with the PARP inhibitor PJ34 (50 µM, 15 min prior to IL-4 stimulation) or vehicle, respectively. (N = 3, mean ± SD). Cell viability of cells as indicated below (N = 3, mean ± SD).
Inhibition of PARP blocks the mutant STAT6 gain-of-function phenotype
We wanted to exploit the potential of targeting PARP14 to block the STAT6MUT phenotype. Since 293T cells do not express endogenous PARP14 we used HeLa cells. We first expressed two different shRNAs to knock-down PARP14 and again quantified STAT6-activated transcriptional activity by luciferase assay. In fact, knock-down of PARP14 significantly reduced transactivation activity of both STAT6D419G and STAT6D419N, correlating with the efficiency of PARP14 knock-down (Supplementary Fig. 8). In contrast, STAT6WT showed no baseline transcriptional activity and was unaffected by PARP14 knock-down (Supplementary Fig. 8).
Next, we knocked-down PARP14 in OCI-Ly1 and OCI-Ly8 stably expressing either STAT6WT, STAT6D419G or EV. In both cell lines shRNA-mediated knock-down of PARP14 (sh4) resulted in significant reduction of IL-4 induced CD23 expression compared to scrambled control (scr) and completely abrogated the STAT6MUT gain-of-function phenotype (Fig. 5D, E).
Finally, we sought to inhibit PARP14 pharmacologically. For this, OCI-Ly1 and OCI-Ly8 cells stably expressing STAT6MUT or STAT6WT were treated with the PARP inhibitor PJ34, in the absence or presence of IL-4, and assayed for STAT6 activation by CD23 expression. PARP inhibition completely blocked IL-4 induced expression of CD23 in cells expressing STAT6D419G down to levels observed with STAT6WT (Fig. 5F). Vice versa, we did not observe a significant effect of PJ34 treatment in the absence of IL-4 or in cells expressing STAT6WT.
Discussion
Re-education of the TME has been identified as a hallmark of FL, actually enabling its pathogenesis and progression [34]. Furthermore, the composition of the TME has been shown to impact clinical outcome of patients with FL [10, 11, 35, 36]. However, the molecular mechanisms that orchestrate and maintain this re-education process in genetically defined subsets of FL often remain elusive, yet they hold great potential as therapeutic targets.
Here, we add novel and clinically relevant aspects to seminal studies that established TFH-derived IL-4 [13, 37, 38] and STAT6 mutations [23] as drivers in FL. Specifically, we describe a therapeutically targetable PARP14-mediated self-reinforcing regulatory circuit that amplifies IL-4 induced transcriptional activity of STAT6MUT (Fig. 6).
STAT6 mutations amplify IL-4 induced STAT6-dependent gene activation via an intracellular self-reinforcing regulatory microcircuit that involves aberrantly increased PARP14 levels in IL-4 stimulated STAT6MUT FL cells. Increased STAT6-dependent gene expression involves cytokines (e.g., CCL17 and CCL22) which contribute to the re-education of the tumor microenvironment, including increased recruitment of IL-4 producing T follicular helper (TFH) cells. Details see text. Yellow star indicates STAT6 mutation. Figure created with BioRender.com.
Conceptually, we propose that this IL-4 driven re-education process of the TME is composed of at least two self-reinforcing regulatory circuits, an intracellular and an extracellular route (Fig. 6). The extracellular route has been well described and involves cytokines like CCL17 and CCL22 in the FL TME [39]. Both chemokines are produced by FL cells in response to IL-4 stimulation (and CD40L) and promote recruitment of Tregs and further accumulation of IL-4 producing TFH cells [39]. Our data shows that these cytokines are indeed among the top upregulated genes in primary patient biopsies of STAT6MUT FL, supporting the concept that STAT6 mutations amplify the IL-4 driven re-education process of the FL TME. In this work, we focused on the molecular mechanisms of the intracellular route (the microcircuit) and identified several aspects of clinical-translational relevance.
First, the gain-of-function phenotype of STAT6MUT was strictly dependent on the presence of IL-4 stimulation. Hence, the biological and clinical relevance of STAT6 mutations in patients may only be adequately evaluable in the context of the TME composition, specifically including TFH abundance and distribution and IL-4 levels. This could explain why the STAT6 mutation status per se has not yet been found to be associated with differences in the clinical course of the disease or treatment outcome [21]. In fact, when we performed an exploratory analysis of our previously reported cohort of patients, who received standard immunochemotherapies (R-CHOP or R-CVP) for advanced stage FL [21], we observed only a trend toward shorter failure-free survival for patients with STAT6MUT FL, but this did not reach statistical difference (data not shown). We hypothesize that combined biomarkers, e.g., integrating the TFH abundance/TME composition and/or IL-4 levels with STAT6 mutation status, will help to further refine patient stratification [36].
Interestingly, an alternative mechanism of constitutive IL-4 signaling and STAT6 activation independent of external IL-4 stimulation has been reported for another B cell lymphoma subtype, primary mediastinal B cell lymphoma (PMBL). Specifically, up to a quarter of PMBL harbor gain-of-function mutations in the IL-4R alpha chain, which, of note, frequently co-occur with additional mutations in the JAK/STAT pathway, including STAT6 [40], further supporting the concept that STAT6 mutations function primarily as amplifiers of this signaling cascade. While we did not find IL-4R mutations in our previously reported cohort of patients with untreated FL (N = 112, GLSG2000 cohort) [21], this may have to be re-addressed in larger cohorts including patients with variant histologies that indicate less dependence on the TME, such as diffuse FL or histologically transformed FL [18,19,20, 41, 42].
Furthermore, we notice that STAT6D419N, which is annotated as a missense germline polymorphism (rs11172102) exerts a gain-of-function phenotype similar to all other somatic STAT6 mutations affecting the DNA binding domain. In the absence of matched germline control, we cannot ultimately distinguish between rare germline polymorphisms and somatically acquired mutations in each of these cases. Importantly however, irrespective of its acquisition, our data indicates that this amino acid change in the DNA binding domain similarly contributes to the distinct biology of STAT6MUT FL. Thus, caution should be taken when sequencing data is analyzed using standard pipelines, which frequently filter out putative or confirmed germline variants.
Finally and importantly, we identified PARP14 as a novel STAT6MUT-specific target gene. PARP14 was previously found to associate with STAT6 [43] and function as a transcriptional switch and activator of STAT6-dependent gene expression in the presence of IL-4 [44]. Our data support a model in which increased PARP14 levels drive a self-reinforcing microcircuit that promotes the assembly of the STAT6 enhanceosome complex in IL-4 stimulated STAT6MUT lymphoma cells, thereby further amplifying STAT6-dependent gene activation. Of note, the catalytic activity of PARP14 has been shown to be essential for the transcriptional enhancement function of PARP14 [15]. In fact, we show that pharmacological inhibition of the PARP enzymatic activity was able to completely abrogate the STAT6MUT gain-of-function phenotype.
Our study also raises some interesting successive hypotheses that should be addressed in future studies. It is, for instance, intriguing to speculate that increased PARP14 levels may have additional functional consequences. As such, PARP14 has been implicated in mediating IL-4 induced attenuation of caspase-3 activation [45], i.e., increased PARP14 levels could contribute to protection against apoptosis and provide a survival advantage to STAT6MUT lymphoma cells. Furthermore, our study provides proof-of-principle that the identification and functional characterization of aberrant targets of STAT6MUT, potentially not limited to PARP14, as well as aberrant targets of other recurrently mutated transcription factors holds promise for developing individualized treatment strategies.
In summary, we show that STAT6 mutations amplify IL-4 induced STAT6-dependent gene activation via a self-reinforcing regulatory circuit that involves aberrantly increased PARP14 levels, and therefore represents a novel therapeutic target in STAT6MUT FL.
References
Hiddemann W, Cheson BD. How we manage follicular lymphoma. Leukemia 2014;28:1388–95.
Casulo C, Byrtek M, Dawson KL, Zhou X, Farber CM, Flowers CR, et al. Early Relapse of Follicular Lymphoma After Rituximab Plus Cyclophosphamide, Doxorubicin, Vincristine, and Prednisone Defines Patients at High Risk for Death: An Analysis From the National LymphoCare Study. J Clin Oncol : Off J Am Soc Clin Oncol. 2015;33:2516–22.
Jurinovic V, Kridel R, Staiger AM, Szczepanowski M, Horn H, Dreyling MH, et al. Clinicogenetic risk models predict early progression of follicular lymphoma after first-line immunochemotherapy. Blood 2016;128:1112–20.
Carbone A, Roulland S, Gloghini A, Younes A, von Keudell G, López-Guillermo A, et al. Follicular lymphoma. Nat Rev Dis Prim. 2019;5:83.
Kridel R, Sehn LH, Gascoyne RD. Pathogenesis of follicular lymphoma. J Clin Investig. 2012;122:3424–31.
Weigert O, Weinstock DM. The promises and challenges of using gene mutations for patient stratification in follicular lymphoma. Blood 2017;130:1491–8.
Bararia D, Hildebrand JA, Stolz S, Haebe S, Alig S, Trevisani CP, et al. Cathepsin S Alterations Induce a Tumor-Promoting Immune Microenvironment in Follicular Lymphoma. Cell Rep. 2020;31:107522.
Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC, et al. Prediction of Survival in Follicular Lymphoma Based on Molecular Features of Tumor-Infiltrating Immune. Cells 2004;351:2159–69.
Hellmuth JC, Louissaint A Jr., Szczepanowski M, Haebe S, Pastore A, Alig S, et al. Duodenal-type and nodal follicular lymphomas differ by their immune microenvironment rather than their mutation profiles. Blood 2018;132:1695–702.
Kiaii S, Clear AJ, Ramsay AG, Davies D, Sangaralingam A, Lee A, et al. Follicular lymphoma cells induce changes in T-cell gene expression and function: potential impact on survival and risk of transformation. J Clin Oncol : Off J Am Soc Clin Oncol. 2013;31:2654–61.
Tobin JWD, Keane C, Gunawardana J, Mollee P, Birch S, Hoang T, et al. Progression of Disease Within 24 Months in Follicular Lymphoma Is Associated With Reduced Intratumoral Immune Infiltration. J Clin Oncol : Off J Am Soc Clin Oncol. 2019;37:3300–9.
Nath K, Tsang H, Gandhi MKJAOL. Tumor microenvironment of follicular lymphoma. Ann. Lymphoma 2021;2021:5.
Amé-Thomas P, Le Priol J, Yssel H, Caron G, Pangault C, Jean R, et al. Characterization of intratumoral follicular helper T cells in follicular lymphoma: role in the survival of malignant B cells. Leukemia 2012;26:1053–63.
Pangault C, Ame-Thomas P, Ruminy P, Rossille D, Caron G, Baia M, et al. Follicular lymphoma cell niche: identification of a preeminent IL-4-dependent TFH-B cell axis. Leukemia 2010;24:2080–9.
Goenka S, Cho SH, Boothby M. Collaborator of Stat6 (CoaSt6)-associated poly(ADP-ribose) polymerase activity modulates Stat6-dependent gene transcription. J Biol Chem. 2007;282:18732–9.
Mehrotra P, Riley JP, Patel R, Li F, Voss L, Goenka S. PARP-14 functions as a transcriptional switch for Stat6-dependent gene activation. J Biol Chem. 2011;286:1767–76.
Takeda K, Tanaka T, Shi W, Matsumoto M, Minami M, Kashiwamura S, et al. Essential role of Stat6 in IL-4 signalling. Nature 1996;380:627–30.
Morin RD, Assouline S, Alcaide M, Mohajeri A, Johnston RL, Chong L, et al. Genetic Landscapes of Relapsed and Refractory Diffuse Large B-Cell Lymphomas. Clin Cancer Res : Off J Am Assoc Cancer Res. 2016;22:2290–300.
Okosun J, Bodor C, Wang J, Araf S, Yang CY, Pan C, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014;46:176–81.
Pasqualucci L, Khiabanian H, Fangazio M, Vasishtha M, Messina M, Holmes AB, et al. Genetics of follicular lymphoma transformation. Cell Rep. 2014;6:130–40.
Pastore A, Jurinovic V, Kridel R, Hoster E, Staiger AM, Szczepanowski M, et al. Integration of gene mutations in risk prognostication for patients receiving first-line immunochemotherapy for follicular lymphoma: a retrospective analysis of a prospective clinical trial and validation in a population-based registry. Lancet Oncol. 2015;16:1111–22.
Ritz O, Guiter C, Castellano F, Dorsch K, Melzner J, Jais JP, et al. Recurrent mutations of the STAT6 DNA binding domain in primary mediastinal B-cell lymphoma. Blood 2009;114:1236–42.
Yildiz M, Li H, Bernard D, Amin NA, Ouillette P, Jones S, et al. Activating STAT6 mutations in follicular lymphoma. Blood 2015;125:668–79.
Li J, Rodriguez JP, Niu F, Pu M, Wang J, Hung L-W. et al. Structural basis for DNA recognition by STAT6. Proc Natl Acad Sci U S A. 2016;113:13015–20.
Caeser R, Gao J, Di Re M, Gong C, Hodson DJ. Genetic manipulation and immortalized culture of ex vivo primary human germinal center B cells. Nat Protoc. 2021;16:2499–519.
Janjic A, Wange LE, Bagnoli JW, Geuder J, Nguyen P, Richter D, et al. Prime-seq efficient and powerful bulk RNA sequencing. Genome Biol. 2022;23:88. https://doi.org/10.1186/s13059-022-02660-8.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Haebe S, Shree T, Sathe A, Day G, Czerwinski DK, Grimes SM, et al. Single-cell analysis can define distinct evolution of tumor sites in follicular lymphoma. Blood 2021;137:2869–80.
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM 3rd, et al. Comprehensive Integration of Single-Cell Data. Cell 2019;177:1888–902.e21.
Deliard S, Zhao J, Xia Q, Grant SF. Generation of high quality chromatin immunoprecipitation DNA template for high-throughput sequencing (ChIP-seq). J Visualized Exp. 2013;74:50286.
Lu X, Nechushtan H, Ding F, Rosado MF, Singal R, Alizadeh AA, et al. Distinct IL-4-induced gene expression, proliferation, and intracellular signaling in germinal center B-cell-like and activated B-cell-like diffuse large-cell lymphomas. Blood 2005;105:2924–32.
Schaefer CF, Anthony K, Krupa S, Buchoff J, Day M, Hannay T, et al. PID: the Pathway Interaction Database. Nucleic Acids Res. 2009;37:D674–9.
Pangault C, Amé-Thomas P, Ruminy P, Rossille D, Caron G, Baia M, et al. Follicular lymphoma cell niche: identification of a preeminent IL-4-dependent T(FH)-B cell axis. Leukemia 2010;24:2080–9.
Scott DW, Gascoyne RD. The tumour microenvironment in B cell lymphomas. Nat Rev Cancer. 2014;14:517–34.
Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC, et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med. 2004;351:2159–69.
Mondello P, Fama A, Larson MC, Feldman AL, Villasboas JC, Yang Z-Z, et al. Lack of intrafollicular memory CD4 + T cells is predictive of early clinical failure in newly diagnosed follicular lymphoma. Blood Cancer J. 2021;11:130.
Calvo KR, Dabir B, Kovach A, Devor C, Bandle R, Bond A, et al. IL-4 protein expression and basal activation of Erk in vivo in follicular lymphoma. Blood 2008;112:3818–26.
Pandey S, Mourcin F, Marchand T, Nayar S, Guirriec M, Pangault C, et al. IL-4/CXCL12 loop is a key regulator of lymphoid stroma function in follicular lymphoma. Blood 2017;129:2507–18.
Rawal S, Chu F, Zhang M, Park HJ, Nattamai D, Kannan S, et al. Cross talk between follicular Th cells and tumor cells in human follicular lymphoma promotes immune evasion in the tumor microenvironment. J Immunol (Baltim, Md :1950). 2013;190:6681–93.
Viganò E, Gunawardana J, Mottok A, Van Tol T, Mak K, Chan FC, et al. Somatic IL4R Mutations in Primary Mediastinal Large B-cell lymphoma lead to constitutive JAK-STAT signaling activation. Blood. 2018;131:2036–2046.
Martín-Moro F, Marquet-Palomanes J, Piris-Villaespesa M, Lopez-Jiménez J, García-Cosío M. Diffuse follicular lymphoma variant with a typical diagnostic pattern and an unusually aggressive clinical presentation. Int J Hematol. 2020;112:136–8.
Xian RR, Xie Y, Haley LM, Yonescu R, Pallavajjala A, Pittaluga S, et al. CREBBP and STAT6 co-mutation and 16p13 and 1p36 loss define the t(14;18)-negative diffuse variant of follicular lymphoma. Blood Cancer J. 2020;10:69.
Goenka S, Boothby M.Selective potentiation of Stat-dependent gene expression by collaborator of Stat6 (CoaSt6), a transcriptional cofactor. Proc Natl Acad Sci U S A. 2006;103:4210–5.
Goenka S, Kaplan MH. Transcriptional regulation by STAT6. Immunologic Res. 2011;50:87–96.
Cho SH, Goenka S, Henttinen T, Gudapati P, Reinikainen A, Eischen CM, et al. PARP-14, a member of the B aggressive lymphoma family, transduces survival signals in primary B cells. Blood 2009;113:2416–25.
Acknowledgements
We acknowledge the iFlow Core Facility of the university hospital Munich (INST 409/225-1 FUGG) for assistance with the generation of flow cytometry data. We are very grateful to Dan Hodson (University of Cambridge, UK) for providing expertize and support with ex vivo modelling of lymphoma from primary human B cells.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
MM, WK, VP, DB, MSS, UZS, and OW designed the study. MM performed immunoblotting, RNA seq, FACS, ELISA and analysis of patient data. EO performed cloning and site-specific mutagenesis experiments. WE, CZ, and L.W. planned, performed and analyzed RNA sequencing experiments. SH, TS, and RL provided and analyzed ssRNA seq data. CDS, WK, and MH developed the FL like GC model and performed qPCR, immunoblotting, and FACS analysis. ES and MH performed immunoblotting. WK performed CFU, qChIP and IP experiments. JAH, MA, and LA assisted with IP experiments and interpretation of the data. LA and MR performed PLA assays. MA, GO, HH, AMS, WH, AR contributed primary patient material and performed IHC stains and analyses. WH supervised the clinical research. MS, JR, and WKla performed the digital multiplexed gene expression and immunohistochemical analyses. OW oversaw all aspects of planning, analyzing and interpretation of experimental and clinical data. MM, WK, VP, and OW wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
This research was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation), Project-ID WE 4679/2-1 (OW), SCHM2440/8-1 (MSS), and the Max-Eder Program of the Deutsche Krebshilfe e.V. (DKH, German Cancer Aid), Project-ID 110659/70112997 (OW). OW is supported by an Else-Kröner Excellence Fellowship from the Else-Kröner-Fresenius Stiftung (Project-ID 2021_EKES.13). LA is supported by the Else Kröner-Fresenius Clinician Scientist Program: Cancer immunotherapy, and the Munich Clinician Scientist Program (MCSP). LA and JAH are supported by the German Consortium for Translational Cancer Research (DKTK) School of Oncology. MS, Amgen, Speakers Bureau. The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mentz, M., Keay, W., Strobl, C.D. et al. PARP14 is a novel target in STAT6 mutant follicular lymphoma. Leukemia 36, 2281–2292 (2022). https://doi.org/10.1038/s41375-022-01641-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-022-01641-x
This article is cited by
-
STAT6 mutations enriched at diffuse large B-cell lymphoma relapse reshape the tumor microenvironment
International Journal of Hematology (2024)
-
STAT6 promoting oxalate crystal deposition-induced renal fibrosis by mediating macrophage-to-myofibroblast transition via inhibiting fatty acid oxidation
Inflammation Research (2023)