Abstract
Despite advances in chronic lymphocytic leukaemia (CLL) treatment, globally chemotherapy remains a central treatment modality, with chemotherapy trials representing an invaluable resource to explore disease-related/genetic features contributing to long-term outcomes. In 499 LRF CLL4 cases, a trial with >12 years follow-up, we employed targeted resequencing of 22 genes, identifying 623 mutations. After background mutation rate correction, 11/22 genes were recurrently mutated at frequencies between 3.6% (NFKBIE) and 24% (SF3B1). Mutations beyond Sanger resolution (<12% VAF) were observed in all genes, with KRAS mutations principally composed of these low VAF variants. Firstly, employing orthogonal approaches to confirm <12% VAF TP53 mutations, we assessed the clinical impact of TP53 clonal architecture. Whilst ≥ 12% VAF TP53mut cases were associated with reduced PFS and OS, we could not demonstrate a difference between <12% VAF TP53 mutations and either wild type or ≥12% VAF TP53mut cases. Secondly, we identified biallelic BIRC3 lesions (mutation and deletion) as an independent marker of inferior PFS and OS. Finally, we observed that mutated MAPK-ERK genes were independent markers of poor OS in multivariate survival analysis. In conclusion, our study supports using targeted resequencing of expanded gene panels to elucidate the prognostic impact of gene mutations.
Similar content being viewed by others
Introduction
The application of new technologies continues to reveal the biological basis for the clinical heterogeneity apparent within CLL [1,2,3]. In particular, next generation sequencing of large patient cohorts has led to the discovery of recurring genomic mutations that cluster into distinct biological signalling pathways. Mutations of specific genes including TP53 [4,5,6,7,8,9,10], ATM [9, 11,12,13,14], BIRC3 [9, 15, 16], SF3B1 [9, 17,18,19,20], NOTCH1 [1, 9, 15, 17, 20,21,22,23], RPS15 [2, 24], EGR2 [25, 26] and KRAS [27, 28] are associated with poorer outcome, especially shorter time to first treatment or overall survival (OS). However, numerous factors influence the clinical significance of a driver mutation in an individual patient. These include clinical status, immunogenetic background, clone size, the presence of biallelic abnormalities and co-existing driver mutations or copy number alterations (CNAs). The clinical importance of these potentially confounding factors is most easily established in context of large clinical trials with long follow-up and where data on numerous biomarkers are available. One such study is the phase III UK LRF CLL4 trial (NCT 58585610) that randomly assigned 777 patients to fludarabine (FDR) or fludarabine plus cyclophosphamide (FC) for six courses, or chlorambucil (CHL) for 12 courses, with the primary endpoint of OS, and secondary endpoints of response rates, progression-free survival, toxic effects and quality of life [29]. The trial demonstrated superior response rates and progression-free survival (PFS) for FC-treated patients compared with those patients treated with FDR or CHL. Previous genomic analysis of this trial has shown TP53 [8], SF3B1 [17], NOTCH1 (coding [17] and non-coding [21]), ATM plus del(11q) [12] and EGR2 [26] lesions to have prognostic significance in multivariate analysis (MVA) and of RPS15 [24] in univariate analysis. The importance of data from CLL4 may be questioned given the studies showing the superior efficacy of FC plus an anti-CD20 antibody (FCR) compared with chemotherapy alone, with the exception of patients with a NOTCH1 mutation [20], and emerging data suggesting the superiority of novel agents compared with chemotherapy-based regimens. However, the observation that TP53, SF3B1 and RPS15 mutations remain poor risk factors in the German CLL8 trial comparing FCR vs. FC [20] and the continuing global need for chemotherapy in CLL for the foreseeable future, indicate that genomic data from the UK CLL4 trial will continue to have clinical relevance.
Accordingly, we performed targeted resequencing on all available pre-treatment samples (n = 499) from the CLL4 trial to investigate the incidence, clinico-biological associations and prognostic impact of a panel of 22 genes recurrently mutated in CLL (study overview in Fig. S1). Important findings include the failure of <12% VAF TP53 mutations (1.97–11.18% variant allele frequency [VAF]) to influence PFS or OS, the importance of 11q deletions on PFS and OS in the context of ATM and BIRC3 mutations, and the reduced OS associated with mutations in the MAPK-ERK genes: BRAF, KRAS and NRAS.
Methods
Patients and molecular assays
We studied 499 patient samples taken at randomisation [29]. Patients were diagnosed using the iwCLL guidelines [30], with informed consent obtained in accordance with the declaration of Helsinki. This study was approved by national/regional research ethics committees. The average lymphocyte percentage of the total white cell count in pre-treatment blood samples was 83.8%. To confirm high tumour load, CD19/CD5 positivity from cases with available flow-cytometry data were compared with their matched average lymphocyte percentage (n = 233), with an agreement bias of −0.8% (Fig. S2). Our study cohort did not significantly differ from the entire trial cohort in terms of: treatment allocation, CNAs, age, gender, disease stage, ZAP70/CD38 expression, or IGHV status (Table S1). The assessment of established biomarkers was performed as described [31]. All published genetic and biological data on CLL4 patients for genes: TP53 [8], ATM [12, 13], BIRC3 [12], NOTCH1 [17] (+3′UTR [21]) and SF3B1 [17], and CNAs: 13q deletion, 17p deletion, 11q deletion and trisomy 12 (5%, 10%, 5% and 3% clone size cut-offs, respectively [31]) were integrated into this study, as well as telomere length [32] and levels of prolymphocytes [33].
Targeted resequencing, bioinformatics analysis, variant filtering and validation
Mutations in 22 genes were analysed in all 499 patients (TruSeq Custom Amplicon, Illumina, San Diego, CA, USA) (Table S2). Libraries were generated from 250 or 50 ng (dependent on the amount of available starting material) of DNA according to manufacturer’s instructions. The average sequencing yield after Illumina processing (MiSeq, paired-end, 2 × 150 bp) from 28 runs was 6.9 Gbp, with a mean read depth of >1000 × (range 502–7948) across all targeted genes, with only nine amplicons below a mean read depth of 1000 (range 502–987) (Fig. S3).
At this depth subclonal mutations can be detected at the 2% level, assuming a minimum observation of four sequencing reads containing the variant base, a Q50 phred like base quality score (p(detected) = 99.999) and a cumulative binomial distribution for n read depth \([\frac{{N!}}{{n!\left( {N - n} \right)!}}p^n(1 - p)^{N - n}]\). In addition, six variants below 2% were included, since the number of sequencing reads in the variant base were more than ten times the assumed minimum observation (range 50–126), and the total read depth exceeded 2000 reads in all cases (range 2582–6389). Bioinformatic data processing of variants was conducted as previously described [14].
All mutations included in this study are listed in Table S3. As the CLL4 cohort lacked germ-line DNA, we only considered variants previously observed as somatically acquired in CLL [1, 2, 14] or annotated in COSMIC (v70) [34], except for specific circumstances regarding TP53, ATM, BIRC3 and NOTCH1. For TP53, additional mutations annotated in IARC were re-introduced [35]. Pathogenic ATM variants were included if; they were observed in AT families as pathogenic (LOVD [https://databases.lovd.nl/shared/genes/ATM]), they were evolutionary rare missense [36], or were somatically acquired in CLL [13] (Table S4). However, this variant strategy does not fully preclude ATM variants that exist in germ-line material. For BIRC3, only truncating mutations were included [9]. NOTCH1 PEST domain mutations not predicted to result in protein truncation were removed. All candidate variants were visually inspected in Integrated Genomics Viewer [37]. Genes were defined as recurrent using Tumour Portal (www.tumorportal.org/power), with the background mutation rate for CLL stated on the website, and the number of cases in the study (n = 499) inputted. Mutations were stratified using Sanger sequencing threshold of 12% [5, 9].
Thirty-one percent (194/623) of mutations were validated using orthogonal approaches, including Sanger (n = 120) and Ion Torrent (19 low-level TP53 mutations) sequencing, hybridization-based gene enrichment with subsequent sequencing (n = 27) and ddPCR (SF3B1 p.K700E [n = 11], NOTCH1 p.P2415fs [n = 19]). One hundred percent of variants were confirmed using this approach. An excellent agreement between TruSeq and orthogonal-derived VAFs was also observed, with an agreement bias of 0.02% (Fig. S4).
Statistical analysis
Fisher’s exact tests were performed for co-occurrence analysis between mutated genes and clinical features. PFS and OS were assessed from randomisation using Kaplan Meier (KM) and Log rank analysis. PFS was defined as time from randomisation to progression (i.e. relapse needing further treatment) or death, or to last follow-up date (Oct 2010; final CLL4 PFS update). OS was defined as time from randomisation to death or to last follow-up date for survivors (August 2016, final CLL4 OS update). Multivariate Cox Proportional Hazard models were generated for OS and PFS using backwards selection (P < 0.05), to test the confounding effect of multiple prognostic variables. The Bland–Altman test was used to test agreement between multiple factors, reporting the agreement bias, which is the mean difference between two measurements. All reported P values were two-sided and results were considered significant at the 5% level, using multiple hypothesis testing when appropriate (Benjamini and Hochberg method [38]). Statistical analysis was conducted using R v3.3.0, SPSS v23 (IBM), and Prism v6.0 g (GraphPad).
Results
Distribution of somatic mutations
We identified 623 mutations (mean = 1.25, min/max = 0/7 per patient) in 335 patients, 398 ≥ 12% VAF and 225 < 12% VAF, with 93% of the entire cohort harbouring ≥1 mutation or CNA (Fig. 1a). Ninety-seven patients without any established CNAs carried 124 mutations (mean = 1.28, min/max = 0/6 per patient), with 22 patients lacking any mutations or CNAs. After background correction (0.5/Mb, ≥3% recurrence, Table S5), 11/22 genes were recurrently mutated at frequencies between 3.6% (NFKBIE) and 24% (SF3B1), (Fig. 1a, Table S3, Fig. S5). 121 samples harboured 134 SF3B1 mutations; 46.3% were the p.K700E variant and 30.6% were other hotspot variants (p.K666X, p.H662X, pG740E, p.G742D). Two or more SF3B1 mutations were identified in 12 patients (Fig. S6), with six cases harbouring multiple SF3B1 mutations present with different VAFs, suggesting the presence of multiple mutated sub-clones. 69 NOTCH1 mutated patients were identified (13.8%), with 61 mutations in exon 34 (50/61 p.2514 fs) and 9 in the 3′UTR. Fifty-five patients carried 59 IARC-annotated TP53 mutations (exons 4–11, 88% in exons 5–8). Forty pathogenic ATM mutations were observed in 37 cases, without evidence of any mutational hotspots. BIRC3, POT, BRAF, XPO1 and KRAS were mutated in 7.2, 6, 6, 5.8 and 5.8% of cases, respectively. Thirty-eight cases harboured a mutation in BRAF, with 7 (18.4%) having the p.V600E variant (Fig. S7).
a Mutational landscape of CLL4. In the Waterfall plot, known recurrently mutated genes and copy number alterations are shown, hierarchically clustered by mutation frequency (vertical bar chart, right). The mutation burden captured by the study is shown in the bar chart above the heat map. Mutation types are depicted in the above key. The inset vertical bar chart represents the distribution of the number of mutated genes/CNAs per case. b Co-occurrence of all available clinico-biological features from the CLL4 clinical trial. The co-occurrence (red) or mutual exclusivity (green) is plotted per interaction in the graph based on the level of significance (from light to dark: P < 0.05, P < 0.01, Q > P [P < 0.05], Q > P [P < 0.01]).
Clinico-biological features of recurrently mutated genes
Next, we determined statistical associations between these gene mutations, and expansive clinico-biological features, using the Fisher’s Exact test (n = 1293 tests, Fig. 1b). One hundred twenty-six associations were observed, including 15 high- (FDR, Q > P [P < 0.05]), 35 medium- (P < 0.01) and 76 low-confidence associations (P < 0.05). Significant associations between mutations were found in only 10/171 possible associations, such as NOTCH1 + 3′UTR with BIRC3 (P = 0.02) and FBXW7 (P = 0.01), as well as BRAF with TP53 (P = 0.03) (Fig. 1b, Table S6).
Distribution of ≥12% VAF and <12% VAF mutations
Next, we classified mutations as Sanger positive (≥12% VAF) or negative (<12% VAF) by accounting for the impact of tumour purity on VAF. Initially, we studied 233 patients with tumour purity derived from CD5/CD19 flow cytometry. Raw VAFs were compared with purity-adjusted VAFs across all variants (n = 288), including <12% VAFs (n = 98), and showed an agreement bias of only 5% (Fig. S8A), which was even lower for <12% VAF mutations (agreement bias < 0.82%, Fig. S8B). Therefore, we analysed all raw VAFs, and observed three variant populations: those found at <12% VAF (1.49–11.56%, n = 225), those at larger subclonal or clonal levels (12.06–58.15%, n = 356), and those concomitant with deletion events (60.19–99.66%, n = 42) (Fig. 2a). SAMHD1 mutations were exclusively composed of ≥12% VAF (55.3% mean VAF), while ATM, MYD88, NOTCH1, SF3B1, TP53 and XPO1 mutations were found to be contain a significant majority of ≥12% VAF mutations. KRAS mutations were more likely to be composed of low VAF variants, with a mean VAF of 10.7% (two-way binomial test, False Discovery Rate [FDR], Q > P [P < 0.05]) (Fig. 2b).
a Distribution of mutation variant allele frequency. Scatter plot of all variants by read depth and VAF (red dots = < 12% VAF [left of dotted line], blue dots = > 12% VAF). b Distribution of ≥12% and <12% variants. Top: Proportion of ≥12% and <12% variants ranked by highest proportion of ≥12% VAF variants. Two-way binomial distribution used to test whether genes contained significantly more ≥12% VAF or <12% VAF mutations, with asterisks representing genes which retained significance after multiple hypothesis testing (Q > P [P < 0.05]). Bottom: VAF distribution of variants per gene. Variants with loss of the other allele (identified by FISH), shown in red for biallelic TP53, turquoise for biallelic ATM and pink for biallelic BIRC3.
Univariate impact of mutated genes on PFS and OS
Clinico-biological features and gene mutations associated with PFS and OS in univariate Cox Proportional Hazards analysis are shown in Fig. 3a. Gene mutations in TP53 (with/without del(17p), termed ‘TP53ab’) and EGR2 were associated with reduced PFS (Fig. 3a, Table 1 and Fig. S9). TP53ab, and recurrent mutations in SF3B1, NOTCH1 (+3′UTR), EGR2, RPS15, NFKBIE, BRAF, KRAS and NRAS were associated with reduced OS (Fig. 3a, Table 1 and Fig. S10). As expected, mutations in MYD88 were confined to IGHV-mutated (IGHV-M) cases, having no significant impact on OS in this subgroup of patients (Fig. S11). In addition, TP53 mutations were associated with poor response (Fig. 3b), NOTCH1 + 3′UTR mutations were associated with death from Richter’s syndrome (Fig. 3c), whilst TP53, SF3B1, NOTCH1 + 3′UTR, KRAS and EGR2 were significantly associated with < 10 year survival (Fig. 3d). Other significant associations are included in Figs. S12 and S13.
a Forest plot showing the hazard ratios of 26 significant variables for either overall survival (left; black) or progression-free survival (right; red) in univariate survival analysis. Variables sorted by the hazard ratio values for overall survival. b Bar chart showing the mutation frequency difference between TP53mut cases who achieved CR/NodPR or NR/PD. c Bar chart showing the NOTCH1 + 3′UTR mutation frequency in relation to Death from Richter’s syndrome. d Bar chart showing the mutation frequency in relation to patients termed ‘long-term survivors’ for TP53, SF3B1, NOTCH +3′UTR, KRAS and EGR2.
Clinical relevance of TP53 deletions and mutations
TP53 mutations below the threshold of Sanger sequencing have been associated with inferior survival in retrospective analysis of institutional cohorts [5, 9]. We observed 59 TP53 mutations in 55 patients (Fig. 4a); all of those tested (n = 51) were confirmed using orthogonal approaches (Table S3). These <12% TP53 mutated cases were enriched for BRAF and FBXW7 mutations (Table S7). TP53 mutations could be further subdivided into those with < 12% VAF (n = 16) or ≥12% VAF (n = 43), with no difference in the site or type of TP53 mutation between subgroups (Fig. 4a). After including 17p FISH data, 58 TP53ab patients were identified, divided into cases with sole 17p deletions (n = 3), isolated TP53 mutations (n = 27) or both (n = 23). Five TP53 mutated cases lacked FISH data.
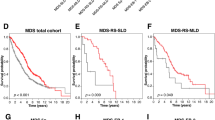
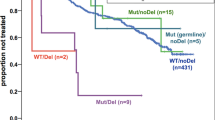
a Mutation Lolliplot displaying the TP53 mutations observed in CLL4, stratified by Sanger sequencing threshold. b Mutated genes/CNVs per TP53mut subgroup. One-way ANOVA conducted vs. TP53wt cases. c Examples of In-going and out-going edges drawn from each TP53mut subgroup, with patient ID number and IGHV status defined above each graph. d OS pairwise KM plot comparing ≥ 12% VAF TP53mut cases (red), <12% VAF TP53mut cases(green), and TP53wt cases (black). e PFS pairwise KM plot comparing ≥ 12% VAF TP53mut cases (red), <12% VAF TP53mut cases(green), and TP53wt cases (black). Inset table in D&E displays pairwise log rank P values between each variable vs. wild type.
Next, we assessed the genomic complexity of TP53mut cases. Both <12% VAF and ≥12% VAF TP53mut groups had increased mutation/CNA frequency in comparison to TP53wt cases (both P < 0.001) (Fig. 4b). To further understand the complexity of these two patient subgroups, we inferred the evolutionary history of TP53mut cases as previously described in CLL [2]. Both <12% VAF and ≥12% VAF cases exhibited the same heterogeneous pattern of co-existing mutations, where TP53 mutations were present at higher, or lower VAFS than concomitant driver mutations (Table S8, Figs. 4c and S14).
Lastly, we assessed the clinical impact of <12% VAF and ≥12% VAF TP53mut subgroups in pairwise KM analysis. ≥12% VAF TP53mut were associated with reduced PFS and OS compared with cases with wild-type TP53 (≥12% TP53mut = OS: median = 2.18 years vs. 6.11 years, P < 0.001, PFS: median = 0.5 years vs. 2.17 years, P < 0.001). In contrast, we could not demonstrate a significant difference between the <12% VAF TP53mut cases and either the wild type or ≥12% VAF TP53mut patients for PFS or OS (<12% TP53mut = OS: median = 4.21 years vs. 6.11 years, P = 0.12, PFS: median = 1.92 years vs. 2.17 years, P = 0.196) (Fig. 4d, e). These observations held true in 17p deletion stratified analysis (Fig. S15), confirming the importance of TP53mut clone size on survival in this cohort. Stratified < 12% VAF vs. ≥ 12% VAF analysis for other genes with sufficient mutated cases in this cohort can be found in Figs. S16 and S17.
Biallelic BIRC3 deleted patients infer reduced OS in comparison to sole 11q deleted patients
Although neither ATM nor BIRC3 mutations, regardless of their VAF (Figs. S16 and S17), were associated with reduced PFS or OS in univariate survival analysis (Figs. S12 and S13), it has previously been demonstrated that the impact of these mutations may be dependent on the presence of a concomitant 11q deletion [12, 39]. Therefore, we performed an integrated analysis of the clinical impact of ATM and BIRC3 mutations in the context of 11q deleted CLL. ATM Mutations spanned the entire gene, whilst those targeting BIRC3 were restricted to the CARD domain, as previously shown [9, 11,12,13, 39] (Figs. 5a and S7). Importantly, ATM and BIRC3 mutations were mutually exclusive in our series (Fig. 5b), suggesting that these mutations may define subgroups of 11q deleted CLL. Deletions of 11q were identified using a FISH probe, which encompasses the ATM but not the BIRC3 locus. Accordingly, concomitant BIRC3 loss was defined from previously published SNP6.0 data [12], or where additional DNA was available (n = 21), using shallow WGS (positive cases presented in Fig. S18). Cases (n = 135) were then categorised into five distinct subgroups: sole 11q deletion (n = 71), biallelic ATM abnormalities (abs) (n = 12), biallelic BIRC3 abs (n = 9), sole ATM mutations (n = 24) and sole BIRC3 mutations (n = 19).
a Mutation Lolliplot of ATM (upper) and BIRC3 (lower) mutations observed in CLL4. b Heat map of ATM and BIRC3 mutated cases stratified by 11q deletion status. c OS pairwise KM plot comparing mutated ATM (left) and BIRC3 (right) in the context of 11q deletion. d PFS pairwise KM plot comparing mutated ATM (left) and BIRC3 (right) in the context of 11q deletion. Inset table in C&D displays pairwise log rank P values between each variable vs. wild type for combined pairwise KM analysis of ATM and BIRC3 in the context of 11q deletion.
After omitting ten cases with co-existing TP53ab [12], we conducted pairwise KM analysis for these five groups compared with cases with no 11q abnormality. (Figs. 5c, d and S19). For both PFS and OS, sole 11q deletion (PFS: median = 1.4 years vs. 2.5 years, P < 0.0001, OS: median = 4.8 years vs. 6.4 years, P = 0.002), as well as biallelic ATM (PFS: median = 1 year vs. 2.5 years, P = 0.001, OS: median = 4.2 years vs. 6.4 years, P = 0.049) and biallelic BIRC3 (PFS: median = 1 year vs. 2.5 years, P < 0.0001, OS: median = 3.3 years vs. 6.4 years, P = 0.001), were associated with a significantly reduced survival.
The outcome of cases with biallelic abs was then compared with those with del(11q) only. There were no significant differences in PFS (biallelic ATM vs. 11q = 1 year vs. 1.4 years, P = 0.336; biallelic BIRC3: 1 year vs. 1.4 years, P = 0.178); however cases with biallelic BIRC3 abs had a significantly reduced OS, whilst cases with biallelic ATM abs did not significantly differ in median survival times compared with sole 11q deleted cases (biallelic ATM vs. 11q = 4.2 years vs. 4.8 years, P = 0.493; biallelic BIRC3: 3.3 years vs. 4.8 years, P = 0.03). This suggests that biallelic loss of BIRC3 represents the subgroup of 11q deleted CLL with the worst outcome following initial treatment with chemotherapy.
MAPK-ERK pathway members: BRAF, KRAS and NRAS, all infer poor OS in CLL4
Mutations in MAPK-ERK genes, BRAF (38 mutations/30 cases), KRAS (34/29) and NRAS (11/10), were principally composed of specific hotspot variants (BRAF: p.G469A/E, KRAS: p.G13D, NRAS: p.Q61K/R) (Fig. S7), and the majority of MAPK-ERK mutated cases (87%) only harboured a mutation in one of these genes (Fig. 6a). Interestingly, MAPK-ERK mutated patients displayed an increased frequency of mutated genes and CNVs per case vs. MAPK-ERK wild-type patients (Fig. S20). In univariate analysis, each mutation was associated with a shorter median OS than wild type: BRAF (OS median: 3.92 years vs. 6 years, P = 0.009), KRAS (OS median: 3.83 years vs. 5.89 years, P < 0.001), and NRAS (OS median: 4.24 years vs. 5.88 years, P = 0.01) (Fig. 6b–d). Stratified <12% VAF vs. ≥ 12% VAF analysis indicated that the outcome of KRAS mutated cases was independent of VAF while shorter OS in BRAF mutated cases was associated with < 12% VAF (Fig. S16; Table S9). Taken together, MAPK-ERK mutations exhibited inferior OS compared with wild-type cases (OS median: 3.83 years vs. 6.10 years, P < 0.001), and were negatively associated with long-term survival (Odds ratio = 0.19, P = 0.0003) (Fig. 6e), with only 4/60 mutated cases defined as long-term survivors. Furthermore, MAPK-ERK mutated patients were more likely to carry IGHV-U genes (IGHV-U Odds Ratio = 4.29, P < 0.0001; IGHV homology >99% Odds ratio = 3.51, P = 0.0002), and significantly less likely to harbour del(13q) as a sole aberration (Odds ratio = 0.23, P < 0.0001, Table S10).
a Heat map of BRAF (blue), KRAS (green), NRAS (red) and co-mutated genes of MAPK-ERK mutated cases (black). Cases wild type for each gene represented by grey bars. Overall survival univariate KM plots for BRAF (b), KRAS (c), NRAS (d), and a combined variable of APK-ERK (e). Coloured line represents mutated cases, black line represents wild-type cases.
Multivariate modelling identifies TP53ab, biallelic BIRC3, SF3B1, EGR2 and MAPK-ERK gene mutations as independent markers of inferior OS
Finally, we constructed comprehensive multivariate Cox Proportional Hazards models for PFS and OS (Table 2), which included those clinical and genetic variables significant in univariate analysis, as well as biallelic ATM and BIRC3 as they emerged from our stratified 11q deletion analysis, and short telomeres based on our previous paper on the topic [32]. A backwards selection approach was applied, until all variables within the model had a P value < 0.05. For PFS, the final model was constructed from 225 patients and 210 events (274 were excluded due to missing data) and showed that TP53ab (HR = 4.98, P < 0.001), biallelic BIRC3 (HR = 3.83, P = 0.004), short telomeres (HR = 1.96, P < 0.001), sole 11q deletion (HR = 1.82, P = 0.003), and increased prolymphocytes (HR = 1.51, P = 0.033) were independent markers of PFS. For OS, the final model was constructed from 391 patients and 323 events (108 observations were excluded due to missing data). TP53ab (HR = 4.25, P < 0.001), biallelic BIRC3 (HR = 2.76, P = 0.004), mutations in EGR2 (HR = 2.19, P = 0.015), MAPK-ERK genes (HR = 1.68, P = 0.002), SF3B1 (HR = 1.54, P = 0.001), as well as IGHV-U genes (HR = 1.83, P < 0.001) and Binet stage B&C (HR = 1.45, P = 0.008), were all observed as independent markers of OS. This data extends on our univariate survival analysis, showing that cases with biallelic BIRC3 deletions exhibit reduced PFS and OS, and that mutations in the MAPK-ERK pathway lead to reduced OS.
Discussion
We report targeted resequencing analysis of 22 genes known to be recurrently mutated in CLL in the UK CLL4 clinical trial. CLL4 represents an ideal candidate for such an analysis, with expansive clinical and biological description [8, 12, 13, 17, 21, 24, 26, 29, 31,32,33, 40, 41] and protracted clinical follow-up. Our study confirms previous studies incorporating samples from this patient cohort showing the impact of TP53ab on PFS and OS in MVA, SF3B1, EGR2 [25, 26], RPS15 [1, 24] and NFKBIE [25, 28, 42] mutations on OS in univariate analysis, with SF3B1 and EGR2 mutations retained as independent markers of OS in MVA.
The literature suggests that patients with MAPK-ERK mutations represent a biologically distinct subgroup, where MAPK-ERK mutations are frequently mutually exclusive, are enriched for trisomy 12, unmutated IGHV genes and other adverse biological markers (e.g. CD38, ZAP-70, CD49d), and are linked to inferior time to first treatment in retrospective cohorts [41, 43,44,45]. We now show the MAPK-ERK genes, BRAF, KRAS and NRAS (collectively representing 12.2% of patients) are also independently associated with short OS in a cohort of patients requiring treatment. Vendramini et al. showed a similar frequency of mutations in these genes (14%) [44], while Giménez et al. found that 5.5% of CLL cases harbours functionally deleterious mutations in 11 genes involved in the MAPK-ERK pathway [45], the latter likely reflects the early-stage composition of the cohort. In support of the biological impact of these mutated genes in CLL, (1) Analysis of mutated patients exhibit an enrichment of gene sets associated with transcriptional activation of the MAPK-ERK pathway [44], (2) preliminary in vitro analysis suggests cells from these patients are prone to killing with ERK inhibitors [45], (3) BRAF mutations accelerated disease progression in Eµ-TCL1 mice [46], (4) mutant BRAF has been implicated in venetoclax resistance [47], and (5) KRAS mutated cases associated with poor response to chemoimmunotherapy [27] and lenalidomide [48].
Screening for TP53ab using FISH and Sanger sequencing has known prognostic value [6, 8, 20, 31], and predicts for resistance to chemoimmunotherapy [49]. TP53 mutations that present at low VAFs, below the detection limit of conventional Sanger sequencing may also be positively selected by chemotherapy, and also predict inferior survival, at least in retrospective, institutional cohorts [3, 5, 9]. The TP53 Network of ERIC provide expansive guidelines on the most suitable approach for TP53 mutational analysis, but also conclude that the clinical importance of low-level TP53 clones remains an unresolved issue, requiring validation in clinical trials [49]. We demonstrated inferior PFS and OS only for those patients with ≥12% VAF TP53 mutations, but we could not demonstrate inferior survival associated with cases harbouring <12% VAF TP53 mutations, the inference perhaps is that these cases represent an intermediate-risk group. Given the unexpected nature of this finding, we also conducted stratified 17p deleted survival analysis, identifying the same result for <12% VAF TP53 mutations without 17p deletion. Furthermore, we proceeded to show that our observation was not associated with any differences in the type of TP53 mutation, their co-existence with other more clonal prognostically-important gene mutations or biological features, nor the enrichment of any specific treatment. As a consequence, we feel that our observation is technically sound, and warrants confirmation in further studies.
There remains disagreement regarding the relative clinical significance of deletion and mutation of the BIRC3 and ATM genes, both mapping to the long arm of chromosome 11. The ATM gene is mutated in 30–40% of 11q deleted patients [11, 13], where it results in biallelic inactivation of ATM, driving an impaired DNA damage response [50]. The prognostic impact of ATM mutations is controversial in unselected cohorts [9], with the strongest impact when the wild-type allele is lost. In our study, whilst we triaged ATM mutations based on their putative pathogenicity, several are reported in both somatic (i.e. COSMIC) and germline (i.e. dbSNP, EXAC, ClinVar) databases, lending uncertainty to their prognostic impact. The sequencing of matched germ-line material would provide additional clarity, but was not possible due to the historical nature of CLL4. Preliminary studies support a pathogenetic role of BIRC3 [16, 39], more recent studies provide less certainty. For example, in the RESONATE clinical trial [51] and the large retrospective study coordinated by ERIC [52], BIRC3 mutations were not linked to inferior PFS or TTFT, respectively. Another comparator would be the RESONATE2 trial, which compared first line treatment with Ibrutinib vs. chlorambucil [53]. The 24 month PFS for 11q deleted patients in the Ibrutinib arm was 97%. Further studies are required to determine if the long-term outcome of biallelic BIRC3 cases is equally good under modern small molecule inhibition. In our previous CLL4 analysis, we demonstrated that BIRC3 dysfunction (defined as deletion AND/OR mutations of BIRC3) did not impact survival in 11q deleted CLL, while biallelic ATM lesions remained informative [12]. However, this analysis utilised Sanger sequencing, and hence only identified a small number of BIRC3 mutations. Our current study, therefore aimed to expand the analysis with a larger patient cohort with significantly improved technology. This approach permitted the identification of a meaningful number of cases with loss and mutation of BIRC3. As neither ATM nor BIRC3 mutations were linked to survival in univariate analysis, we performed a stratified analysis in 11q deleted cases. In so doing, we show that biallelic BIRC3 cases have a further reduction in survival in comparison to sole 11q deleted cases and were found to be independent prognostic markers for PFS and OS in MVA. Finally, ATM and BIRC3 mutated cases without 11q deletion have a similar survival to wild-type cases.
In conclusion, our study makes three main contributions to the field. We show an expansive analysis of the impact of clinico-biological disease features on the clinical importance of important gene mutations, including SF3B1, EGR2 and the MAPK-ERK genes. Our analysis suggests that <12% VAF TP53 mutations are an intermediate survival group. Finally, we show that biallelic BIRC3 aberrations identify a novel patient subgroup with poor survival, inferior to those with 11q deletions alone. Taken together, we demonstrate that a more expansive genomic screening approach provides additional clinical information, thereby helping to establish the precise importance of genetic alterations in the context of other established and emerging biomarkers. Furthermore, our work will facilitate the development of international standards for the detection and interpretation of somatic mutations in CLL.
References
Puente XS, Beà S, Valdés-Mas R, Villamor N, Gutiérrez-Abril J, Martín-Subero JI, et al. Non-coding recurrent mutations in chronic lymphocytic leukaemia. Nature. 2015;526:519–24.
Landau DA, Tausch E, Taylor-weiner AN, Stewart C, Reiter JG, Bahlo J, et al. Mutations driving CLL and their evolution in progression and relapse. Nature. 2015;526:525–30.
Nadeu F, Martín-García D, López-Guillermo A, Navarro A, Colado E, Campo E, et al. Clinical impact of the subclonal architecture and mutational complexity in chronic lymphocytic leukemia. Leukemia. 2017;32:645–53.
Minervini CF, Cumbo C, Orsini P, Brunetti C, Anelli L, Zagaria A, et al. TP53 gene mutation analysis in chronic lymphocytic leukemia by nanopore MinION sequencing. Diagn Pathol. 2016;11:96.
Rossi D, Khiabanian H, Spina V, Ciardullo C, Bruscaggin A, Famà R, et al. Clinical impact of small TP53 mutated subclones in chronic lymphocytic leukemia. Blood. 2014;123:2139–48.
Zenz T, Eichhorst B, Busch R, Denzel T, Habe S, Winkler D, et al. TP53 mutation and survival in chronic lymphocytic leukemia. J Clin Oncol. 2010;28:4473–9.
Malcikova J, Stano-Kozubik K, Tichy B, Kantorova B, Pavlova S, Tom N, et al. Detailed analysis of therapy-driven clonal evolution of TP53 mutations in chronic lymphocytic leukemia. Leukemia. 2015;29:877–85.
Gonzalez D, Martinez P, Wade R, Hockley S, Oscier D, Matutes E, et al. Mutational status of the TP53 gene as a predictor of response and survival in patients with chronic lymphocytic leukemia: results from the LRF CLL4 trial. J Clin Oncol. 2011;29:2223–9.
Nadeu F, Delgado J, Royo C, Baumann T, Stankovic T, Pinyol M, et al. Clinical impact of clonal and subclonal TP53, SF3B1, BIRC3, NOTCH1 and ATM mutations in chronic lymphocytic leukemia. Blood. 2016;127:2122–30.
Pekova S, Mazal O, Cmejla R, Hardekopf DW, Plachy R, Zejskova L, et al. A comprehensive study of TP53 mutations in chronic lymphocytic leukemia: analysis of 1287 diagnostic and 1148 follow-up CLL samples. Leuk Res. 2011;35:889–98.
Austen B, Powell JE, Alvi A, Edwards I, Hooper L, Starczynski J, et al. Mutations in the ATM gene lead to impaired overall and treatment-free survival that is independent of IGVH mutation status in patients with B-CLL. Blood. 2005;106:3175–82.
Rose-Zerilli MJJ, Forster J, Parker H, Parker A, Rodri AÉ, Chaplin T, et al. ATM mutation rather than BIRC3 deletion and/or mutation predicts reduced survival in 11q-deleted chronic lymphocytic leukemia: Data from the UK LRF CLL4 trial. Haematologica. 2014;99:736–42.
Skowronska A, Parker A, Ahmed G, Oldreive C, Davis Z, Richards S, et al. Biallelic ATM inactivation significantly reduces survival in patients treated on the United Kingdom leukemia research fund chronic lymphocytic leukemia 4 trial. J Clin Oncol. 2012;30:4524–32.
Guièze R, Robbe P, Clifford R, De Guibert S, Pereira B, Timbs A, et al. Presence of multiple recurrent mutations confers poor trial outcome of relapsed/refractory CLL. Blood. 2015;126:2110–7.
Fabbri G, Rasi S, Rossi D, Trifonov V, Khiabanian H, Ma J, et al. Analysis of the chronic lymphocytic leukemia coding genome: role of NOTCH1 mutational activation. J Exp Med. 2011;208:1389–401.
Rossi D, Rasi S, Spina V, Bruscaggin A, Monti S, Ciardullo C, et al. Integrated mutational and cytogenetic analysis identifies new prognostic subgroups in chronic lymphocytic leukemia. Blood. 2013;121:1403–12.
Oscier DG, Rose-Zerilli MJJ, Winkelmann N, Gonzalez de Castro D, Gomez B, Forster J, et al. The clinical significance of NOTCH1 and SF3B1 mutations in the UK LRF CLL4 trial. Blood. 2012;121:468–75.
Jeromin S, Weissmann S, Haferlach C, Dicker F, Bayer K, Grossmann V, et al. SF3B1 mutations correlated to cytogenetics and mutations in NOTCH1, FBXW7, MYD88, XPO1 and TP53 in 1160 untreated CLL patients. Leukemia. 2014;28:108–17.
Quesada V, Conde L, Villamor N, Ordóñez GR, Jares P, Bassaganyas L, et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet. 2012;44:47–52.
Stilgenbauer S, Schnaiter A, Paschka P, Zenz T, Rossi M, Döhner K, et al. Gene mutations and treatment outcome in chronic lymphocytic leukemia: Results from the CLL8 trial. Blood. 2014;123:3247–54.
Larrayoz M, Rose-Zerilli MJJ, Kadalayil L, Parker H, Blakemore S, Forster J, et al. Non-coding NOTCH1 mutations in chronic lymphocytic leukemia; their clinical impact in the UK CLL4 trial. Leukemia. 2016;31:510–4.
Rossi D, Rasi S, Fabbri G, Spina V, Fangazio M, Forconi F, et al. Mutations of NOTCH1 are an independent predictor of survival in chronic lymphocytic leukemia. Blood. 2016;119:521–9.
Puente XS, Pinyol M, Quesada V, Conde L, Ordóñez GR, Villamor N, et al. Whole-genome sequencing identifies recurrent mutations in chronic lymphocytic leukaemia. Nature. 2011;475:101–5.
Ljungstrom V, Cortese D, Young E, Pandzic T, Mansouri L, Plevova K, et al. Whole expme sequencing in relapsig chronic lymphocytic leukemia: clinical impact of recurrent RPS15 mutations. Blood. 2016;127:1007–16.
Damm F, Mylonas E, Cosson A, Yoshida K, Della Valle V, Mouly E, et al. Acquired initiating mutations in early hematopoietic cells of CLL patients. Cancer Discov. 2014;4:1088–101.
Young E, Noerenberg D, Mansouri L, Ljungström V, Frick M, Sutton L-A, et al. EGR2 mutations define a new clinically aggressive subgroup of chronic lymphocytic leukemia. Leukemia. 2017;31:1547–54.
Herling CD, Klaumünzer M, Rocha CK, Altmüller J, Thiele H, Bahlo J, et al. Complex karyotypes and KRAS and POT1 mutations impact outcome in CLL after chlorambucil-based chemotherapy or chemoimmunotherapy. Blood. 2016;128:395–404.
Doménech E, Gómez-López G, Gzlez-Peña D, López M, Herreros B, Menezes J, et al. New mutations in chronic lymphocytic leukemia identified by target enrichment and deep sequencing. PLoS ONE. 2012;7:2–7.
Catovsky D, Richards S, Matutes E, Oscier D, Dyer M, Bezares R, et al. Assessment of fludarabine plus cyclophosphamide for patients with chronic lymphocytic leukaemia (the LRF CLL4 Trial): a randomised controlled trial. Lancet. 2007;370:230–9.
Hallek M, Cheson BD, Catovsky D, Caligaris-Cappio F, Dighiero G, Döhner H, et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia: a report from the International Workshop on Chronic Lymphocytic Leukemia updating the National Cancer Institute-Working Group 1996 guidelines. Blood. 2008;111:5446–56.
Oscier D, Wade R, Davis Z, Morilla A, Best G, Richards S, et al. Prognostic factors identified three risk groups in the LRF CLL4 trial, independent of treatment allocation. Haematologica. 2010;95:1705–12.
Strefford JC, Kadalayil L, Forster J, Rose-Zerilli MJJ, Parker A, Lin TT, et al. Telomere length predicts progression and overall survival in chronic lymphocytic leukemia: Data from the UK LRF CLL4 trial. Leukemia. 2015;29:2411–4.
Oscier D, Else M, Matutes E, Morilla R, Strefford JC, Catovsky D. The morphology of CLL revisited: the clinical significance of prolymphocytes and correlations with prognostic/molecular markers in the LRF CLL4 trial. Br J Haematol. 2016;174:767–75.
Forbes SA, Beare D, Boutselakis H, Bamford S, Bindal N, Tate J, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res. 2017;45:D777–83.
Petitjean A, Mathe E, Kato S, Ishioka C, Tavtigian S, Hainaut P, et al. Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat. 2007;28:622–9.
Tavtigian SV, Oefner PJ, Babikyan D, Hartmann A, Healey S, Calvez-kelm FLe, et al. Rare, evolutionarily unlikely missense substitutions in ATM confer increased risk of breast cancer. Am J Hum Genet. 2009;85:427–46.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrated genomics viewer. Nat Biotechnol. 2011;29:24–26.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc. 1995;57:289–300.
Rossi D, Fangazio M, Rasi S, Vaisitti T, Monti S, Cresta S, et al. Disruption of BIRC3 associates with fludarabine chemorefractoriness in TP53 wild-type chronic lymphocytic leukemia. Blood. 2012;119:2854–62.
Parker H, Rose-Zerilli MJJ, Larrayoz M, Clifford R, Edelmann J, Blakemore S, et al. Genomic disruption of the histone methyltransferase SETD2 in chronic lymphocytic leukaemia. Leukemia. 2016;30:2179–86.
Pandzic T, Larsson J, He L, Kundu S, Ban K, Akhtar-Ali M, et al. Transposon mutagenesis reveals fludarabine resistance mechanisms in chronic lymphocytic leukemia. Clin Cancer Res. 2016;22:6217–27.
Mansouri L, Sutton L-A, Ljungström V, Bondza S, Arngården L, Bhoi S, et al. Functional loss of IκBε leads to NF-κB deregulation in aggressive chronic lymphocytic leukemia. J Exp Med. 2015;212:833–43.
Leeksma AC, Taylor J, Dubois J, Dietrich S, de Boer F, Zelenetz A, et al. Clonal diversity predicts adverse outcome in chronic lymphocytic leukemia. Leukemia. 2018;33:390–402.
Vendramini E, Bomben R, Pozzo F, Benedetti D, Bittolo T, Rossi FM, et al. KRAS, NRAS, and BRAF mutations are highly enriched in trisomy 12 chronic lymphocytic leukemia and are associated with shorter treatment-free survival. Leukemia. 2019;12:10–14.
Giménez N, Valero JG, López-Otín C, Payer AR, Puente XS, Martínez-Trillos A, et al. Mutations in RAS-BRAF-MAPK-ERK pathway define a specific subgroup of patients with adverse clinical features and provide new therapeutic options in chronic lymphocytic leukemia. Haematologica. 2018;104:576–86.
Tsai Y-T, Sass EJ, Lozanski G, Byrd JC, Harrington BK, Jaynes F, et al. BRAF V600E accelerates disease progression and enhances immune suppression in a mouse model of B-cell leukemia. Blood Adv. 2017;1:2147–60.
Herling CD, Abedpour N, Weiss J, Schmitt A, Jachimowicz RD, Merkel O, et al. Clonal dynamics towards the development of venetoclax resistance in chronic lymphocytic leukemia. Nat Commun. 2018;9:727.
Takahashi K, Wierda WG, Keating M, Kim E, Thompson P, Burger JA, et al. Clinical implications of cancer gene mutations in patients with chronic lymphocytic leukemia treated with lenalidomide. Blood. 2018;131:1820–32.
Malcikova J, Tausch E, Rossi D, Sutton LA, Soussi T, Zenz T, et al. ERIC recommendations for TP53 mutation analysis in chronic lymphocytic leukemia - Update on methodological approaches and results interpretation. Leukemia. 2018;32:1070–80.
Austen B, Skowronska A, Baker C, Powell JE, Gardiner A, Oscier D, et al. Mutation status of the residual ATM allele is an important determinant of the cellular response to chemotherapy and survival in patients with chronic lymphocytic leukemia containing an 11q deletion. J Clin Oncol. 2007;25:5448–57.
Brown JR, Delgado J, Jaeger U, Montillo M, Hillmen P, Kipps TJ, et al. Extended follow-up and impact of high-risk prognostic factors from the phase 3 RESONATE study in patients with previously treated CLL/SLL. Leukemia. 2017;32:83–91.
Baliakas P, Hadzidimitriou A, Sutton LA, Rossi D, Minga E, Villamor N, et al. Recurrent mutations refine prognosis in chronic lymphocytic leukemia. Leukemia. 2015;29:329–36.
Barr PM, Robak T, Owen C, Tedeschi A, Bairey O, Bartlett NL, et al. Sustained efficacy and detailed clinical follow-up of first-line ibrutinib treatment in older patients with chronic lymphocytic leukemia: extended phase 3 results from RESONATE-2. Haematologica. 2018;103:1502–10.
Acknowledgements
The authors would like to gratefully acknowledge all the patients and clinicians than contributed to the UK CLL4 trial. The CLL4 trial was funded by a core grant from Bloodwise. ME was supported by the Arbib Charitable Fund. This study was supported by grants from Bloodwise (12036, 11052), Kay Kendall Leukaemia Fund (873), Cancer Research UK (C2750/A23669, C34999/A18087, ECMC C24563/A15581) and the Oxford Partnership Comprehensive Biomedical Research Centre with funding from the Department of Health’s National Institute of Health Research (NIHR) Biomedical Research Centre funding scheme. The views expressed in this publication are those of the authors and not necessarily those of the Department of Health. Thank you to Miss Kate Latham for contributing to DNA extraction of CLL4 patient samples.
Author information
Authors and Affiliations
Contributions
SJB, MJJR-Z, RC, AS and JCS designed the research. SJB, MJJR-Z and JCS analysed the data. SJB, MJJR-Z, DGO and JCS and wrote the paper. SJB, RC, HP, PA, ES-D. ML, ZD, LK PR, DV, JF, AB, RM, DC, ME, DB, HMC, DGO, RJW, AJS, MSC, MJJR-Z and AS performed the research and/or contributed patient samples and associated data. All authors read and agreed to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Blakemore, S.J., Clifford, R., Parker, H. et al. Clinical significance of TP53, BIRC3, ATM and MAPK-ERK genes in chronic lymphocytic leukaemia: data from the randomised UK LRF CLL4 trial. Leukemia 34, 1760–1774 (2020). https://doi.org/10.1038/s41375-020-0723-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-020-0723-2
This article is cited by
-
ERIC recommendations for TP53 mutation analysis in chronic lymphocytic leukemia—2024 update
Leukemia (2024)
-
NFKBIE mutations are selected by the tumor microenvironment and contribute to immune escape in chronic lymphocytic leukemia
Leukemia (2024)
-
An inflammatory response-related gene signature can predict the prognosis and impact the immune infiltration of multiple myeloma
Clinical and Experimental Medicine (2024)
-
Different prognostic impact of recurrent gene mutations in chronic lymphocytic leukemia depending on IGHV gene somatic hypermutation status: a study by ERIC in HARMONY
Leukemia (2023)
-
Ultrafast prediction of somatic structural variations by filtering out reads matched to pan-genome k-mer sets
Nature Biomedical Engineering (2022)