Abstract
Using a Burkitt lymphoma-like gene expression signature, we recently defined a high-risk molecular high-grade (MHG) group mainly within germinal centre B-cell like diffuse large B-cell lymphomas (GCB-DLBCL), which was enriched for MYC/BCL2 double-hit (MYC/BCL2-DH). The genetic basis underlying MHG-DLBCL and their aggressive clinical behaviour remain unknown. We investigated 697 cases of DLBCL, particularly those with MYC/BCL2-DH (n = 62) by targeted sequencing and gene expression profiling. We showed that DLBCL with MYC/BCL2-DH, and those with BCL2 translocation, harbour the characteristic mutation signatures that are associated with follicular lymphoma and its high-grade transformation. We identified frequent MYC hotspot mutations that affect the phosphorylation site (T58) and its adjacent amino acids, which are important for MYC protein degradation. These MYC mutations were seen in a subset of cases with MYC translocation, but predominantly in those of MHG. The mutations were more frequent in double-hit lymphomas with IG as the MYC translocation partner, and were associated with higher MYC protein expression and poor patient survival. DLBCL with MYC/BCL2-DH and those with BCL2 translocation alone are most likely derived from follicular lymphoma or its precursor lesion, and acquisition of MYC pathogenic mutations may augment MYC function, resulting in aggressive clinical behaviour.
Similar content being viewed by others
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common lymphoma in adults, accounting for around 75% of aggressive lymphomas. The current standard treatment for DLBCL is immunochemotherapy, typically R-CHOP (rituximab plus cyclophosphamide, doxorubicin, vincristine and prednisone), with 60% of patients living 10 years or more [1]. Patients who fail R-CHOP treatment respond poorly to currently available alternative therapies, and mortality is highest within the first two years after diagnosis [1]. The ability to risk-stratify patients with a low probability of cure following R-CHOP at diagnosis, and to enter them into clinical trials investigating alternative therapies is a significant unmet clinical need. A number of biomarkers have been investigated, but only MYC translocation and to a lesser extent, cell of origin (COO), are used routinely or in a clinical trial [1].
MYC translocation occurs in ~10% of DLBCL, and is frequently (21–83%) accompanied by an additional BCL2 and/or BCL6 translocation, known as double-hit (DH) or triple-hit (TH) [2,3,4,5,6,7,8,9,10,11,12,13,14,15]. MYC/BCL2-DH DLBCL are generally aggressive and respond poorly to currently available therapies, with the majority of patients dying within two years of diagnosis, although a minority of cases experience a long term survival [15, 16]. The clinical outcomes for MYC/BCL6-DH DLBCL are less clear owing to the small number of cases investigated and their heterogeneous COO [14, 17, 18]. Patients with a single MYC translocation (MYC-SH) show variable clinical courses. A subset of these cases has TP53 mutations and displays a worse survival, similar to that of MYC/BCL2-DH [18, 19]. There remains a need to clarify the prognostic stratification of MYC translocation-bearing DLBCL.
Based on COO classification, DLBCL has been divided into two broad subgroups: activated B-cell like (ABC) and germinal centre B-cell like (GCB) subtypes, with the ABC-DLBCL showing enhanced NF-κB activation and generally a worse prognosis [20, 21]. Although the COO classification provides broad biologically distinct categories, there is an apparent heterogeneity in clinical outcome within each subtype. Further subclassification of these broad molecular subtypes has been investigated by several recent studies based on clustered genetic changes and/or gene expression signatures [22,23,24,25,26]. Among these recent advances, Sha et al. and Ennishi et al. have defined a clinically and biologically distinct high-risk subgroup within GCB-DLBCL using, respectively, a Burkitt lymphoma-like or MYC/BCL2-DH-founded gene expression signature [25,26,27]. This subgroup, termed molecular high-grade (MHG) in the study by Sha et al., is enriched in cases with MYC/BCL2-DH, and more importantly includes other poor prognosis cases without the double-hit, which are not readily identified by other methods [25, 28].
Intriguingly, although MHG-DLBCL is enriched in MYC/BCL2-DH, a proportion of cases with MYC/BCL2-DH do not have the MHG gene expression signature and these cases show no worse survival than that of conventional GCB-DLBCL (non-MHG) [25]. To understand the genetic basis underlying MHG-DLBCL and its aggressive clinical behaviour, we used targeted sequencing to investigate 697 cases of DLBCL, particularly enriched for those with MYC/BCL2-DH(TH) (n = 62).
Methods
Case selection and materials
Of the 697 DLBCL cases included, 400 were from the REMoDL-B trial (28 MYC translocation positive) and 297 (97 MYC translocation positive) from a UK population-based cohort [25, 28]. The vast majority of cases in the population-based cohort were from the Haematological Malignancy Research Network (HMRN) (www.hmrn.org), which tracks all haematological malignancies across 14 hospitals diagnosed by a centralised fully-integrated haematopathology laboratory [29]. Case selection in the present study was biased towards those with a MYC translocation, which, together with BCL2 and BCL6 translocations, was determined at the time of diagnosis (HMRN) or retrospectively collected from pathology records or by performing interphase fluorescent in situ hybridisation (FISH) on TMA (REMoDL-B) [18, 25]. Among the population-based cohort, five cases had a previous history of follicular lymphoma (FL), and a further 28 cases had a histological evidence of concurrent FL. For the REMoDL-B trial, patients with a previous history of low-grade lymphoma were excluded.
Formalin-fixed paraffin-embedded (FFPE) diagnostic tissue biopsy was available in each case and local ethical guidelines were followed for the use of these tissue materials for research with the approval of the ethics committees of the involved institutions (05-Q1604–10, 04-Q1205–125, 10-H0504–79).
DNA extraction and quality assessment
Haematoxylin and eosin stained FFPE tissue slides were reviewed and tumour rich areas (>40%) in consecutive sections were isolated by crude macrodissection in each specimen. DNA was extracted using the QIAamp DNA Micro Kit (QIAGEN, Crawley, UK) and quantified using a Qubit® Fluorometer (Life Technologies, UK). The quality of DNA samples was assessed by PCR of variably sized genomic fragments using a standardised protocol [30].
Targeted sequencing by HaloPlexHS enrichment and Illumina HiSeq sequencing
This was essentially carried out as described previously [31]. Briefly, 100 ng genomic DNA was subjected to targeted enrichment of 70 genes (Table S1), which are recurrently mutated in aggressive B-cell lymphomas using a customised HaloPlexHS probe library (Agilent Technologies). The HaloPlexHS probes incorporate molecular barcodes, hence allowing removal of PCR errors during data analysis. Library preparation was performed according to the manufacturer’s instructions for FFPE tissue samples. The pooled libraries were sequenced on an Illumina HiSeq4000 (2 × 150 bp end sequencing protocol) or HiSeq2500 (Rapid Run Mode 2 × 150 bp end sequencing protocol). As stipulated by our previous study, DNA samples amenable for PCR of ≥400 bp genomic fragments were investigated in a single replicate, while those amenable for PCR of 300 bp were analysed in duplicates, with reproducible variants being considered as a true change [31].
Variant calling and annotation
Sequence data analysis and variants calling were performed using a previously validated in-house protocol [31]. Briefly, SNV were called using UnifiedGenotyper with no downsampling [32]. As this was unable to call SNVs at <8% AAF reliably, MuTect2 was additionally employed for detection of hotspot mutations at low AAF values. Indel detection was separately carried out on the recalibrated bam files using Pindel v0.2.5 [33], which allowed detection of indels as low as 2% AAF. Variant calling files were concatenated to produce one library vcf each for the SNV and Indel pipelines, and then filtered using vcftools v0.1.15 and bedtools v2.25 for read depth, quality score and known PCR/sequence artefacts. Further filtering was carried out to remove variants in intronic regions outside essential splicing sites, SNPs with a minor allele frequency ≥0.1% (dbSNP database, 1000 Genomes Project, the ExAC exome sequencing database) and synonymous changes. In addition, missense variants predicted to be benign by at least seven out of nine functional prediction tools (SIFT, Polyphen2 HDIV, Polyphen2 HVAR, LRT, MutationTaster, MutationAssessor, FATHMM, SVM score and LR score) were excluded. The resulting variants were further scrutinised by reviewing the bam file to eliminate potential PCR and sequence artefacts. Only variants above the cut-off value (20 alternative allele depth for DNA samples amenable for PCR of ≥400 bp, 15 alternative allele depth in both replicates for DNA samples amenable for PCR of 300 bp) were considered to be a true change [31]. Finally, extensive search of COSMIC database and published literature was carried out to retain those known and confirmed to be somatic variants. The final mutation list can be found in Table S2.
Molecular subtyping by gene expression profiling
Whole-genome gene expression profiling was performed on mRNA extracted from FFPE diagnostic tissue specimens using the Illumina whole-genome DASL array [34]. Data analyses and COO classification were carried out using the “DLBCL automatic classifier” (DAC) [35]. The MHG group was identified based on a Burkitt lymphoma-like signature as defined in previous studies [25, 36].
Interphase fluorescence in situ hybridisation (FISH)
Chromosome translocation status was available from routine haematopathological diagnosis or previous studies for MYC, BCL2 and BCL6 in 550, 233 and 218 cases, respectively [18, 25]. In the REMoDL-B and HMRN cohort, MYC translocation was screened with Dako MYC break-apart probe, and those showing no evidence of MYC translocation but with MHG phenotype were further investigated with Abbott MYC break-apart and MYC/IGH dual fusion probe. In the remaining cases from other UK hospitals, MYC translocation was investigated with Abbott MYC break-apart probe.
Interphase FISH was performed for BCL2 and BCL6 translocation in 433 and 366 cases in the present study. In cases with MYC translocation, additional FISH was performed with MYC/IGH (Abbott) (if not yet done), then MYC/IGK and MYC/IGL (Cytocell) dual fusion probes in those without any evidence of MYC and IGH fusion. All FISH was carried out on tissue microarray or whole tissue section as described previously [18, 25].
MYC immunohistochemistry
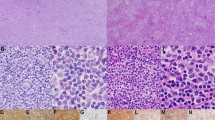
Data on MYC protein expression by immunohistochemistry on tissue microarrays (TMA) slides were available from our previous study [25]. The immunostained slides were scanned and MYC protein expression was quantified and presented as percentage of positive nuclear staining in lymphoma cells using IHC Profiler Image-J software according to the instructions for nuclear protein targets (https://imagej.net/) [37].
Statistical analysis
The probability of MYC hotspot mutations occurring by chance was assessed by Poisson regression. Associations among chromosomal rearrangements, mutations and clinical variables were analysed using Fisher’s exact test. Survival analysis was carried out using Cox proportional hazards and likelihood ratio tests in R (https://cran.r-project.org). All quoted P values are two-sided.
Results
Among the 697 cases investigated by targeted sequencing, 553 were investigated for chromosomal translocations by interphase FISH; and MYC, BCL2 and BCL6 translocations were found in 125, 136 and 97 cases, respectively, with MYC/BCL2/BCL6-TH in 11, MYC/BCL2-DH in 51 (BCL6 translocation data unknown in 8) and MYC/BCL6-DH in 22 (Figs. 1 and 2).
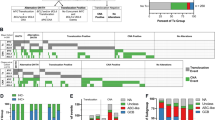
a Heatmap illustration of mutation distribution according to chromosome translocation status; Where data available, evidence of previous or concurrent follicular lymphoma is indicated. b DLBCL with BCL2 translocation, particularly those with MYC/BCL2-DH, harbour the cardinal mutation signature of FL, and also the mutation profile associated with its high-grade transformation [38,39,40,41]. Representative mutation data in FL and transformed FL are from the study by Kridel et al. [41], with EZH2 mutation considered as the core changes associated with FL [38,39,40,41]. c Comparison of BCL2 mutation profile between BCL2 translocation positive DLBCL in the present study and FL in the study by Huet et al. [42]. d DLBCL with BCL2 translocation often have a previous or concurrent follicular lymphoma. MYC/BCL2-DH MYC/BCL2 double-hit, TH MYC/BCL2/BCL6 triple-hit, SH single-hit, tr +ve/−ve translocation positive/negative, FL follicular lymphoma.
The mutation profile of DLBCL with MYC/BCL2-DH or BCL2-SH suggests their derivation from follicular lymphoma
In general, DLBCL with MYC/BCL2/BCL6-TH and MYC/BCL2-DH had a similar mutation profile, and were characterised by a higher mutation load and more frequent mutations in BCL2, KMT2D, CREBBP, EZH2 and TNFRSF14 than those with isolated MYC translocation (Figs. 2a, S1 and S2). Interestingly, these mutations are the cardinal features of FL [38,39,40,41,42], and were similarly seen in DLBCL with BCL2-SH (Fig. 2b). The BCL2 mutation profile was almost identical between BCL2 translocation positive DLBCL and FL (Fig. 2c) [42]. These findings suggest that DLBCL with a BCL2 translocation may be derived from a FL or its precursor lesion. In support of this suggestion, DLBCL with MYC/BCL2-DH(TH) and those with BCL2-SH also harboured an additional mutation profile (MYC, GNA13, TP53, P2RY8, PIM1, CCND3, B2M, EBF1 and S1PR2), which was associated with high-grade transformation of FL as shown by several previous studies (Fig. 2b) [38,39,40,41]. Furthermore, both groups (28 and 50%, respectively) frequently presented with either a previous or concurrent FL (Fig. 2d). Intriguingly, DLBCL with MYC-BCL2-DH(TH) were more often associated with a previous, but not concurrent FL than those with BCL2-SH. Nonetheless, 65% of BCL2 translocation positive cases lacked documented evidence of FL at diagnosis (Fig. 2d). With the exception of MYC mutations, there was no significant difference in the mutation profile between BCL2 translocation positive cases with and without FL (Fig. S3). MYC mutation was more frequent in BCL2 translocation positive cases without FL, and this was primarily due to a high frequency of MYC translocation in this group.
As expected, DLBCLs with MYC/BCL2-DH(TH) were either MHG (56%) or GCB (38%), the remaining cases being unclassifiable (6%) (Fig. 3). Similarly, the majority of DLBCL with BCL2-SH were GCB (74%), with the remaining cases distributed among MHG (14%), ABC (4%) and unclassified (8%). It is worth noting that the three cases of ABC-DLBCL with BCL2-SH lacked both EZH2 and GNA13 mutations that were nearly exclusively seen in GCB (MHG)-DLBCL.
MHG-DLBCL with MYC translocation are enriched with MYC mutations that enhance its stability and transforming capacity
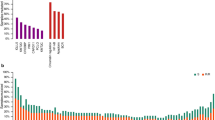
Our previous study showed that among patients with a MYC translocation, MHG-DLBCL had a significantly worse survival than GCB-DLBCL (non-MHG) [25]. To understand the genetic basis underlying MHG-DLBCL and its aggressive clinical behaviour, we compared the mutation profile among MHG, GCB and ABC subtype, and also between MHG and GCB within MYC/BCL2 double-hit groups (Fig. 4). This revealed a significantly higher frequency of MYC mutations in the MHG group (Fig. 4b).
a Comparison of mutation profile among MHG, GCB and ABC subtypes. The panel includes only the genes (n = 57) that had a mutation frequency ≥5% in at least one molecular subtype to make the figure legible. b Mutation comparison between MHG and GCB within the MYC/BCL2-DH(TH) group. Only genes showing a significant or apparent difference are included in the figure panel, with BCL2 mutation included as a reference. Fisherʼs exact test was used to analyse the difference of mutation frequency between various groups with statistical significance indicated.
MYC is a known target of somatic hypermutation machinery, and as expected many of the MYC mutations were in the RCY-motif (R = A/G, Y = C/T), with their extent attenuating when further downstream from the promoter (Fig. 5a). In comparison with synonymous mutations, there was a significant enrichment of non-synonymous changes in codons 57, 58 and 59 (Fig. 5a). In addition, an in-frame deletion of codons 56–60 was seen in one case. These mutations are likely to be functional, pathogenic and selected during lymphoma development as they affect the phosphorylation site (T58) and its neighbouring amino acids, which are important for MYC protein degradation [43]. Several lymphoma derived MYC mutants, including P57S and T58A, have been previously shown to dramatically increase the half-life of MYC protein, and also confer increased transforming capacity [44, 45].
a Distribution of MYC variants with hotspot mutations clustered at the phosphorylation site (T58) and its neighbouring amino acids that are important for MYC degradation. *One case shows an in-frame deletion of codons 56–60 and all other mutations are missense changes. The codons 57, 58 and 59 hotspot mutations and the in-frame deletion are likely pathogenic and selected during lymphoma development. MYC variants are annotated according to transcript ENST00000377970.6 in keeping with the literature. b MYC hotspot mutations in codons 57, 58 and 59 are seen in a subset of cases with MYC translocation, more frequent in those with MYC/BCL2-DH, but are significantly enriched in MHG-DLBCL. c MYC hotspot mutations are more commonly seen in MYC/BCL2-DH DLBCL with IG gene as the MYC translocation partner, with a considerable overlap with MHG phenotype. d DLBCL with MYC mutation in codons 57–59 show a significantly higher MYC protein expression than those with MYC translocation, but lacking these pathogenic mutations. The MYC protein expression was investigated by immunohistochemistry, quantified using IHC Profiler Image-J software and presented as percentage of positive nuclear staining in lymphoma cells. Unpaired t-test was used to compare the two groups. MHG molecular high-grade, GCB germinal centre B-cell like, ABC activated B-cell like UNC, unclassified.
These MYC hotspot mutations were seen in a subset of DLBCL with MYC translocation, more frequently in those with MYC/BCL2-DH, and the majority (74%) were MHG (Fig. 5b). In contrast, cases with MYC non-synonymous mutations in other codons did not show any association with molecular subtype although occurred predominantly in those with MYC/IG translocation (Fig. 5b, c). MYC hotspot mutations were significantly more frequent in cases with MYC/IG (41%) than those with MYC/non-IG translocation (8%), and together had a considerable overlap with MHG phenotype (Fig. 5c). Cases with MYC hotspot mutations had a significantly higher MYC protein expression than those with other MYC mutations (Fig. 5d) as shown by immunohistochemistry and quantitative analysis of the scanned immunostained slides [25]. Finally, MYC pathogenic mutations had a potential adverse effect on patient survival (Fig. 6). Even within the MHG group, cases with MYC pathogenic mutations had significantly worse overall survival than those without these mutations in the REMoDL-B trial, and a similar trend was also seen in HMRN’s population-based cohort (Fig. 6a, b). In a separate analysis within cases with MYC/BCL2-DH irrespective of their MHG status, cases with MYC pathogenic mutations also had significantly worse overall survival than those without these mutations in the REMoDL-B trial, albeit not in HMRN’s population-based cohort (Fig. S4). In multivariable analysis adjusting for MHG and MYC/BCL2-DH, MYC pathogenic mutations retained statistical significance in the REMoDL-B group, albeit not in the HMRN cohort.
a Differential impact on survival between MYC mutations in codons 57–59, and others. b MHG-DLBCL with MYC mutations in codons 57–59 show the worst overall survival in comparison with GCB-DLBCL. c Single variable Cox proportional hazards regression analysis of progression-free survival in GCB-DLBCL. *In multivariable analysis adjusting for MHG and MYC/BCL2-DH, MYC pathogenic mutations in codons 57–59 retain statistical significance in the REMoDL-B cohort, albeit not in the HMRN cohort.
A low allelic frequency (4–5% AAF) of the above MYC pathogenic mutations was seen in three cases, likely representing a subclonal change. Among these three cases, two had molecular subtyping data and one was MHG.
DLBCL with MYC/BCL6-DH or BCL6-SH, or MYC-SH are heterogeneous in their mutation profile and molecular subtype
There was no apparent mutation signature, nor biased molecular subtype associated with BCL6 translocation with exception of BCL10 and NOTCH2 mutations which were significantly enriched in cases with MYC/BCL6-DH or BCL6-SH (Figs. 2a, 3 and S5). Similarly, there was no specific mutation signature in DLBCL with MYC-SH with the exception of high frequent MYC mutations. Cases with MYC-SH varied in their molecular subtype, nonetheless MHG (39%) and GCB (29%) accounted for the majority. Interestingly, 6 of the 11 MHG-DLBCL with MYC-SH had TP53 mutations.
Discussion
Using integrated analyses of chromosome translocation, somatic mutation profiling of a panel of 70 genes that are recurrently mutated in aggressive B-cell lymphoma, and molecular subtype in a large cohort of DLBCL, the present study made two novel observations.
First, we have provided several strands of evidence indicating that DLBCL with MYC/BCL2-DH and those with BCL2-SH are most likely derived from a low-grade FL or its precursor lesion. These include finding: (1) a cardinal mutation signature (BCL2, KMT2D, CREBBP, EZH2 and TNFRSF14) associated with FL development; (2) a mutation profile (MYC, GNA13, TP53, P2RY8, PIM1, CCND3, B2M, EBF1 and S1PR2) associated with FL high-grade transformation; and (3) frequent presence of a previous or concurrent FL in DLBCL with BCL2 translocation.
We acknowledge the limitation of the relatively small gene panel used in the present study, and the above speculation needs to be confirmed by more comprehensive genetic profiling. Nonetheless, the speculation is supported by the finding that transformed FLs also show frequent MYC hotspot mutation [41]. In support of our study, the mutation signature associated with FL was also the characteristic change in high-grade B-cell lymphoma with MYC and BCL2 translocations [46], and in the EZB or C3 genetic subgroup, which are enriched by BCL2 translocation [23, 24].
Intriguingly, DLBCL with MYC/BCL2-DH had strong mutation signatures associated with FL, but less frequent evidence of a concurrent FL than those with BCL2-SH (28% vs 50%). Given the highly proliferative nature of DLBCL with MYC/BCL2-DH, these high-grade lymphoma cells may frequently efface the low-grade FL lesion, potentially leading to its underdetection. In addition, a single lymph node is commonly biopsied for histological diagnosis, increasingly needle core rather than excision biopsies. This would underestimate the true incidence of FL in patients with DLBCL. Alternatively, DLBCL with MYC/BCL2-DH may be derived from a precursor lesion, such as a common mutated precursor cell population. In fact, transformed FLs are more commonly (66–83%) derived from a common mutated precursor cell (CPC) population, in a process of divergent evolution [38,39,40,41]. The tissue compartment containing the CPC population is likely to be in situ follicular neoplasia (ISFN), the precursor lesion of FL, albeit this remains to be confirmed in future investigations. It is pertinent to speculate that BCL2 translocation positive DLBCL could be similarly derived from an ISFN lesion, regardless of any evidence for parallel FL development.
Second, we have identified frequent MYC hotspot mutations that affect the phosphorylation site (T58) and its adjacent amino acids, which are critical for FBXW7 mediated proteasome degradation (Fig. 5a) [43]. Such lymphoma derived MYC mutants (T58A, P57S) have been shown to increase the half-life of MYC protein from 30 to 110 min, and also confer increased transforming capacity, but are defective in promoting apoptosis due to failure to activate BIM [44, 45, 47]. Thus, these hotspot mutations are likely to be pathogenic and selected during lymphoma development.
Although these MYC hotspot mutations were seen in a subset of cases with MYC translocation, they were predominantly in MHG. This may explain, at least in part, the heterogeneous molecular subtype and clinical outcome of DLBCL with MYC translocation, including those with MYC/BCL2-DH. MYC hotspot mutations (pathogenic changes) were significantly more frequent in cases with MYC/IG than those with MYC/non-IG translocation (41% vs 8%, P = 0.005), potentially explaining in part why these cases showed a worse prognosis than those with non-IG gene as the MYC partner [48, 49]. Cases with MYC pathogenic mutations were significantly associated with higher MYC protein expression as assessed by immunohistochemistry and quantitative imaging analysis. This could potentially explain in part the variability (20–100%) of MYC protein expression in tumour cells of MYC translocation positive DLBCL [50]. The above findings are also consistent with the observation that the adverse prognosis of the MYC/BCL2-DH group is primarily due to those with IG as MYC translocation partner [48, 51].
It is worth noting that the above MYC hotspot mutations were also seen at a subclonal level, albeit in only a few cases. This finding raises an interesting question about whether tumour cells carrying these mutations are more resistant to therapies, and could thus be enriched in resistant or relapsed disease. If this is the case, the clinical impact of these MYC mutations may be under-estimated in the present study, as the investigation was exclusively based on diagnostic tissue biopsies.
More recently, a second hotspot of tumour-associated MYC mutations was identified in codons 243–249 through meta-analysis of mutation data from Burkitt lymphoma [52]. These mutants (T244A, P245A) phenocopy the aforementioned mutations in their enhanced MYC protein stability, transforming capacity, and also defective BIM activation [52]. We did not observe any hotspot mutations in this region, but this could be due to the relatively small number of cases in the present study, or different mutation spectra between Burkitt and DLBCL. Mutation in codon 138 has also been suggested to enhance MYC protein stability and thus regarded as pathogenic change in DLBCL [53, 54]. Nonetheless, mutation in codon 138 was frequently accompanied by the change in codon 58, and its independent impact remains uncertain due to a very limited number of mutant cases identified [54].
In the 2017 WHO lymphoma classification, MYC/BCL6-DH DLBCL are included in the double-hit category, without any distinction from those with MYC/BCL2/BCL6-TH or MYC/BCL2-DH [55]. We show here that DLBCL with MYC/BCL6-DH are significantly different in their mutation profile and molecular subtype from those with MYC/BCL2/BCL6-TH or MYC/BCL2-DH. This observation is further supported by a recent study albeit based on a smaller cohort [46]. In fact, MYC/BCL6-DH DLBCL are highly heterogeneous in their molecular subtypes, indicating their diverse COO, notwithstanding the high prevalence of NOTCH2 mutations [23, 24]. BCL6 translocation is recurrently seen in both follicular and marginal zone lymphoma [56,57,58,59]. It would be interesting to explore whether MYC/BCL6-DH DLBCL also result from high-grade transformation of a low-grade lesion such as follicular or marginal zone B-cell lymphoma or their precursor lesions, and their heterogeneous molecular subtypes reflect their inherent features from their derived low-grade lesion. These heterogeneities, in addition to the small numbers of cases available for each study, may explain the disparate clinical outcomes reported for MYC/BCL6-DH DLBCL [14, 17, 18]. In light of this, MYC/BCL6-DH DLBCL should not be regarded as a single group, and their biology and clinical management need to be explored in the context of their respective molecular subtype, rather than within the double-hit category.
In summary, DLBCL with MYC/BCL2-DH harbour the characteristic mutation signatures that are associated with FL development and its high-grade transformation, suggesting their derivation from FL or its precursor lesion, probably following acquisition of a MYC translocation. Our study also identifies the novel association of MHG-DLBCL with MYC hotspot mutations that enhance its stability and transforming capacity, and further highlight the pathogenic role of these mutations and their clinical significance, beyond transcriptional deregulation as a result of translocation.
References
Davies A. Tailoring front-line therapy in diffuse large B-cell lymphoma: who should we treat differently? Hematol Am Soc Hematol Educ Program. 2017;2017:284–94.
Aukema SM, Siebert R, Schuuring E, van Imhoff GW, Kluin-Nelemans HC, Boerma EJ, et al. Double-hit B-cell lymphomas. Blood. 2011;117:2319–31.
Barrans S, Crouch S, Smith A, Turner K, Owen R, Patmore R, et al. Rearrangement of MYC is associated with poor prognosis in patients with diffuse large B-cell lymphoma treated in the era of rituximab. J Clin Oncol. 2010;28:3360–5.
Obermann EC, Csato M, Dirnhofer S, Tzankov A. Aberrations of the MYC gene in unselected cases of diffuse large B-cell lymphoma are rare and unpredictable by morphological or immunohistochemical assessment. J Clin Pathol. 2009;62:754–6.
Copie-Bergman C, Gaulard P, Leroy K, Briere J, Baia M, Jais JP, et al. Immuno-fluorescence in situ hybridization index predicts survival in patients with diffuse large B-cell lymphoma treated with R-CHOP: a GELA study. J Clin Oncol. 2009;27:5573–9.
Savage KJ, Johnson NA, Ben Neriah S, Connors JM, Sehn LH, Farinha P, et al. MYC gene rearrangements are associated with a poor prognosis in diffuse large B-cell lymphoma patients treated with R-CHOP chemotherapy. Blood. 2009;114:3533–7.
Burotto M, Berkovits A, Dunleavy K. Double hit lymphoma: from biology to therapeutic implications. Expert Rev Hematol. 2016;9:669–78.
Visco C, Tzankov A, Xu-Monette ZY, Miranda RN, Tai YC, Li Y, et al. Patients with diffuse large B-cell lymphoma of germinal center origin with BCL2 translocations have poor outcome, irrespective of MYC status: a report from an international DLBCL rituximab-CHOP consortium program study. Haematologica. 2013;98:255–63.
Johnson NA, Slack GW, Savage KJ, Connors JM, Ben Neriah S, Rogic S, et al. Concurrent expression of MYC and BCL2 in diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol. 2012;30:3452–9.
Horn H, Ziepert M, Becher C, Barth TF, Bernd HW, Feller AC, et al. MYC status in concert with BCL2 and BCL6 expression predicts outcome in diffuse large B-cell lymphoma. Blood. 2013;121:2253–63.
Hu S, Xu-Monette ZY, Tzankov A, Green T, Wu L, Balasubramanyam A, et al. MYC/BCL2 protein coexpression contributes to the inferior survival of activated B-cell subtype of diffuse large B-cell lymphoma and demonstrates high-risk gene expression signatures: a report from the international DLBCL rituximab-CHOP consortium program. Blood. 2013;121:4021–31.
Cunningham D, Hawkes EA, Jack A, Qian W, Smith P, Mouncey P, et al. Rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisolone in patients with newly diagnosed diffuse large B-cell non-Hodgkin lymphoma: a phase 3 comparison of dose intensification with 14-day versus 21-day cycles. Lancet. 2013;381:1817–26.
Green TM, Young KH, Visco C, Xu-Monette ZY, Orazi A, Go RS, et al. Immunohistochemical double-hit score is a strong predictor of outcome in patients with diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol. 2012;30:3460–7.
Tzankov A, Xu-Monette ZY, Gerhard M, Visco C, Dirnhofer S, Gisin N, et al. Rearrangements of MYC gene facilitate risk stratification in diffuse large B-cell lymphoma patients treated with rituximab-CHOP. Mod Pathol. 2014;27:958–71.
Rosenthal A, Younes A. High grade B-cell lymphoma with rearrangements of MYC and BCL2 and/or BCL6: Double hit and triple hit lymphomas and double expressing lymphoma. Blood Rev. 2017;31:37–42.
Friedberg JW. How I treat double-hit lymphoma. Blood. 2017;130:590–6.
Ye Q, Xu-Monette ZY, Tzankov A, Deng L, Wang X, Manyam GC, et al. Prognostic impact of concurrent MYC and BCL6 rearrangements and expression in de novo diffuse large B-cell lymphoma. Oncotarget. 2016;7:2401–16.
Clipson A, Barrans S, Zeng N, Crouch S, Grigoropoulos NF, Liu H, et al. The prognosis of MYC translocation positive diffuse large B-cell lymphoma depends on the second hit. J Pathol Clin Res. 2015;1:125–33.
Schiefer AI, Kornauth C, Simonitsch-Klupp I, Skrabs C, Masel EK, Streubel B, et al. Impact of single or combined genomic alterations of TP53, MYC, and BCL2 on survival of patients with diffuse large B-cell lymphomas: a retrospective cohort study. Med (Baltim). 2015;94:e2388.
Alizadeh AA, Eisen MB, Davis RE, Ma C, Lossos IS, Rosenwald A, et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature. 2000;403:503–11.
Wright G, Tan B, Rosenwald A, Hurt EH, Wiestner A, Staudt LM. A gene expression-based method to diagnose clinically distinct subgroups of diffuse large B cell lymphoma. Proc Natl Acad Sci USA. 2003;100:9991–6.
Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell. 2017;171:481–94.
Chapuy B, Stewart C, Dunford AJ, Kim J, Kamburov A, Redd RA, et al. Molecular subtypes of diffuse large B cell lymphoma are associated with distinct pathogenic mechanisms and outcomes. Nat Med. 2018;24:679–90.
Schmitz R, Wright GW, Huang DW, Johnson CA, Phelan JD, Wang JQ, et al. Genetics and pathogenesis of diffuse large B-cell lymphoma. N. Engl J Med. 2018;378:1396–407.
Sha C, Barrans S, Cucco F, Bentley MA, Care MA, Cummin T, et al. Molecular high-grade B-cell lymphoma: defining a poor-risk group that requires different approaches to therapy. J Clin Oncol. 2019;37:202–12.
Ennishi D, Jiang A, Boyle M, Collinge B, Grande BM, Ben Neriah S, et al. Double-hit gene expression signature defines a distinct subgroup of germinal center B-cell-like diffuse large B-cell lymphoma. J Clin Oncol. 2019;37:190–201.
Chan WC. Using gene expression profiling to move beyond MYC/BCL2 rearrangements in high-grade lymphoma. J Clin Oncol. 2019;37:175–7.
Painter D, Barrans S, Lacy S, Smith A, Crouch S, Westhead D et al. Cell-of-origin in diffuse large B-cell lymphoma: findings from the UK's population-based Haematological Malignancy Research Network. Br J Haematol. 2018;185:781–84.
Smith A, Howell D, Crouch S, Painter D, Blase J, Wang HI et al. Cohort profile: the haematological malignancy research network (HMRN); a UK population-based patient cohort. Int J Epidemiol. 2018;47:700.
Wang M, Escudero-Ibarz L, Moody S, Zeng N, Clipson A, Huang Y, et al. Somatic mutation screening using archival formalin-fixed, paraffin-embedded tissues by fluidigm multiplex PCR and illumina sequencing. J Mol Diagn. 2015;17:521–32.
Cucco F, Clipson A, Kennedy H, Sneath TJ, Wang M, Barrans S, et al. Mutation screening using formalin-fixed paraffin-embedded tissues: a stratified approach according to DNA quality. Lab Invest. 2018;98:1084–92.
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet. 2011;43:491–8.
Ye K, Schulz MH, Long Q, Apweiler R, Ning Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics. 2009;25:2865–71.
Barrans SL, Crouch S, Care MA, Worrillow L, Smith A, Patmore R, et al. Whole genome expression profiling based on paraffin embedded tissue can be used to classify diffuse large B-cell lymphoma and predict clinical outcome. Br J Haematol. 2012;159:441–53.
Care MA, Barrans S, Worrillow L, Jack A, Westhead DR, Tooze RM. A microarray platform-independent classification tool for cell of origin class allows comparative analysis of gene expression in diffuse large B-cell lymphoma. PLoS ONE. 2013;8:e55895.
Sha C, Barrans S, Care MA, Cunningham D, Tooze RM, Jack A, et al. Transferring genomics to the clinic: distinguishing Burkitt and diffuse large B cell lymphomas. Genome Med. 2015;7:64.
Varghese F, Bukhari AB, Malhotra R, De A. IHC profiler: an open source plugin for the quantitative evaluation and automated scoring of immunohistochemistry images of human tissue samples. PLoS ONE. 2014;9:e96801.
Okosun J, Bodor C, Wang J, Araf S, Yang CY, Pan C, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014;46:176–81.
Pasqualucci L, Khiabanian H, Fangazio M, Vasishtha M, Messina M, Holmes AB, et al. Genetics of follicular lymphoma transformation. Cell Rep. 2014;6:130–40.
Bouska A, Zhang W, Gong Q, Iqbal J, Scuto A, Vose J, et al. Combined copy number and mutation analysis identifies oncogenic pathways associated with transformation of follicular lymphoma. Leukemia. 2017;31:83–91.
Kridel R, Chan FC, Mottok A, Boyle M, Farinha P, Tan K, et al. Histological transformation and progression in follicular lymphoma: a clonal evolution study. PLoS Med. 2016;13:e1002197.
Huet S, Szafer-Glusman E, Tesson B, Xerri L, Fairbrother WJ, Mukhyala K, et al. BCL2 mutations do not confer adverse prognosis in follicular lymphoma patients treated with rituximab. Am J Hematol. 2017;92:515–9.
Hydbring P, Castell A, Larsson LG. MYC Modulation around the CDK2/p27/SKP2 Axis. Genes (Basel) 2017;8:174.
Bahram F, von der LN, Cetinkaya C, Larsson LG. c-Myc hot spot mutations in lymphomas result in inefficient ubiquitination and decreased proteasome-mediated turnover. Blood. 2000;95:2104–10.
Salghetti SE, Kim SY, Tansey WP. Destruction of Myc by ubiquitin-mediated proteolysis: cancer-associated and transforming mutations stabilize Myc. EMBO J. 1999;18:717–26.
Evrard SM, Pericart S, Grand D, Amara N, Escudie F, Gilhodes J, et al. Targeted next generation sequencing reveals high mutation frequency of CREBBP, BCL2 and KMT2D in high-grade B-cell lymphoma with MYC and BCL2 and/or BCL6 rearrangements. Haematologica. 2019;104:e154–e157.
Hemann MT, Bric A, Teruya-Feldstein J, Herbst A, Nilsson JA, Cordon-Cardo C, et al. Evasion of the p53 tumour surveillance network by tumour-derived MYC mutants. Nature. 2005;436:807–11.
Copie-Bergman C, Cuilliere-Dartigues P, Baia M, Briere J, Delarue R, Canioni D, et al. MYC-IG rearrangements are negative predictors of survival in DLBCL patients treated with immunochemotherapy: a GELA/LYSA study. Blood. 2015;126:2466–74.
Pedersen MO, Gang AO, Poulsen TS, Knudsen H, Lauritzen AF, Nielsen SL, et al. MYC translocation partner gene determines survival of patients with large B-cell lymphoma with MYC- or double-hit MYC/BCL2 translocations. Eur J Haematol. 2014;92:42–48.
Kluk MJ, Ho C, Yu H, Chen BJ, Neuberg DS, Dal Cin P, et al. MYC immunohistochemistry to identify MYC-driven B-cell lymphomas in clinical practice. Am J Clin Pathol. 2016;145:166–79.
Rosenwald A, Bens S, Advani R, Barrans S, Copie-Bergman C, Elsensohn MH et al. Prognostic significance of MYC rearrangement and translocation partner in diffuse large B-cell lymphoma: a study by the Lunenburg lymphoma biomarker consortium. J Clin Oncol. 2019; JCO1900743.
Chakraborty AA, Scuoppo C, Dey S, Thomas LR, Lorey SL, Lowe SW, et al. A common functional consequence of tumor-derived mutations within c-MYC. Oncogene. 2015;34:2406–9.
Kuttler F, Ame P, Clark H, Haughey C, Mougin C, Cahn JY, et al. c-myc box II mutations in Burkitt's lymphoma-derived alleles reduce cell-transformation activity and lower response to broad apoptotic stimuli. Oncogene. 2001;20:6084–94.
Xu-Monette ZY, Deng Q, Manyam GC, Tzankov A, Li L, Xia Y, et al. Clinical and biologic significance of MYC genetic mutations in de novo diffuse large B-cell lymphoma. Clin Cancer Res. 2016;22:3593–605.
Kluin PM, Harris NL, Stein H, Leoncini L, Campo E, Jaffe ES, et al. High-grade B-cell lymphoma. WHO Classification of tumours of haematpoietic and lymphoid tissues. Revised 4th edn. (2017) 335–41.
Leich E, Hoster E, Wartenberg M, Unterhalt M, Siebert R, Koch K, et al. Similar clinical features in follicular lymphomas with and without breaks in the BCL2 locus. Leukemia. 2016;30:854–60.
Bacon CM, Diss TC, Ye H, Liu H, Goatly A, Hamoudi R, et al. Follicular lymphoma of the thyroid gland. Am J Surg Pathol. 2009;33:22–34.
Pan Y, Meng B, Sun B, Guan B, Liang Y, Wang H, et al. Frequencies of BCL2 and BCL6 translocations in representative Chinese follicular lymphoma patients: morphologic, immunohistochemical, and FISH analyses. Diagn Mol Pathol. 2012;21:234–40.
Gu K, Fu K, Jain S, Liu Z, Iqbal J, Li M, et al. t(14;18)-negative follicular lymphomas are associated with a high frequency of BCL6 rearrangement at the alternative breakpoint region. Mod Pathol. 2009;22:1251–7.
Acknowledgements
The authors would like to thank Shubha Anand and Yuanxue Huang for their assistance with using TapeStation, Graeme Clark and Ezequiel Martin for their assistance with Illumina sequencing, and also Raiza Ulidan and Dan Jiang for their assistance with interphase FISH tests.
Funding
The research was supported by grants from Bloodwise (15002, 15019, 13006) UK. The REMoDL-B trial was endorsed by Cancer Research UK, reference number CRUKE/10/024, and funded by Janssen-Cilag. The Haematological Malignancy Research Network is supported by Bloodwise (15037). AS is partly funded by the National Institute for Health Research Oxford Biomedical Research Centre. DRW acknowledges UK Medical Research Council grant MR/L01629X/1 for infrastructure support. The views and opinions expressed are those of the authors and do not necessarily reflect those of the National Institute for Health Research, the UK National Health Service or the UK Department of Health.
Authors contributions
Targeted sequencing, mutation data collection and analyses: FC, AC, RD; Illumina sequencing analysis and variant calling: ZC, JST; Gene expression, FISH and IHC data: SB and AR; Gene expression data analysis and molecular subtyping: CS, MAC; The REMoDL-B trial was led by PWMJ, AJD, TC, JC; Case contribution: SB, CB, DP, AS, ER, HL; Survival analysis: FC, SC, CS; Study design and coordination: MQD, PWMJ, DRW, AJD, CB, RT, ER, JF, AS; Manuscript writing and preparation: MQD, FC with contributions from all authors. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cucco, F., Barrans, S., Sha, C. et al. Distinct genetic changes reveal evolutionary history and heterogeneous molecular grade of DLBCL with MYC/BCL2 double-hit. Leukemia 34, 1329–1341 (2020). https://doi.org/10.1038/s41375-019-0691-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0691-6