Abstract
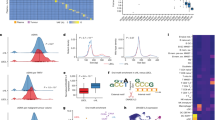
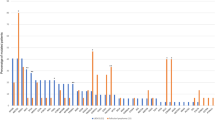
We used hybrid capture-targeted next-generation sequencing of circulating cell-free DNA (ccfDNA) of pediatric Hodgkin lymphoma (PHL) patients to determine pathogenic mechanisms and assess the clinical utility of this method. Hodgkin-Reed/Sternberg (HRS) cell-derived single nucleotide variants, insertions/deletions, translocations and VH-DH-JH rearrangements were detected in pretherapy ccfDNA of 72 of 96 patients. Number of variants per patient ranged from 1 to 21 with allele frequencies from 0.6 to 42%. Nine translocation breakpoints were detected. Genes involved in JAK/STAT, NFkB and PI3K signaling and antigen presentation were most frequently affected. SOCS1 variants, mainly deletions, were found in most circulating tumor (ct) DNAs, and seven of the nine translocation breakpoints involved SOCS1. Analysis of VH-DH-JH rearrangements revealed an origin of PHL HRS cells from partially selected germinal center B cells. Amounts of pretherapy ctDNA were correlated with metabolic tumor volumes. Furthermore, in all ccfDNA samples of 43 patients with early response assessment quantitative qPET < 3, indicative of a favorable clinical course, ctDNA was not detectable. In contrast, in five of six patients with qPET > 3, indicative of an unfavorable clinical course, ctDNA remained detectable. ccfDNA analysis of PHL is thus a suitable approach to determine pathogenic mechanisms and monitor therapy response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Mauz-Korholz C, Metzger ML, Kelly KM, Schwartz CL, Castellanos ME, Dieckmann K. et al. Pediatric Hodgkin lymphoma. J Clin Oncol. 2015;33:2975–85.
Castellino SM, Geiger AM, Mertens AC, Leisenring WM, Tooze JA, Goodman P, et al. Morbidity and mortality in long-term survivors of Hodgkin lymphoma: a report from the Childhood Cancer Survivor Study. Blood. 2011;117:1806–16.
Dorffel W, Riepenhausenl M, Luders H, Bramswig J, Schellong G. Secondary malignancies following treatment for Hodgkin’s lymphoma in childhood and adolescence. Dtsch Arztebl Int. 2015;112:320–7. i
Schaapveld M, Aleman BM, van Eggermond AM, Janus CP, Krol AD, van der Maazen RW, et al. Second cancer risk up to 40 years after treatment for Hodgkin’s lymphoma. N Engl J Med. 2015;373:2499–511.
Barros MH, Hassan R, Niedobitek G. Tumor-associated macrophages in pediatric classical Hodgkin lymphoma: association with Epstein−Barr virus, lymphocyte subsets, and prognostic impact. Clin Cancer Res. 2012;18:3762–71.
Barros MH, Vera-Lozada G, Soares FA, Niedobitek G, Hassan R. Tumor microenvironment composition in pediatric classical Hodgkin lymphoma is modulated by age and Epstein−Barr virus infection. Int J Cancer. 2012;131:1142–52.
Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017;17:223–38.
Kurtz DM, Green MR, Bratman SV, Scherer F, Liu CL, Kunder CA, et al. Noninvasive monitoring of diffuse large B-cell lymphoma by immunoglobulin high-throughput sequencing. Blood. 2015;125:3679–87.
Oberle A, Brandt A, Voigtlaender M, Thiele B, Radloff J, Schulenkorf A, et al. Monitoring multiple myeloma by next-generation sequencing of V(D)J rearrangements from circulating myeloma cells and cell-free myeloma DNA. Haematologica. 2017;102:1105–11.
Roschewski M, Dunleavy K, Pittaluga S, Moorhead M, Pepin F, Kong K, et al. Circulating tumour DNA and CT monitoring in patients with untreated diffuse large B-cell lymphoma: a correlative biomarker study. Lancet Oncol. 2015;16:541–9.
Rossi D, Diop F, Spaccarotella E, Monti S, Zanni M, Rasi S, et al. Diffuse large B-cell lymphoma genotyping on the liquid biopsy. Blood. 2017;129:1947–57.
Scherer F, Kurtz DM, Newman AM, Stehr H, Craig AF, Esfahani MS. et al. Distinct biological subtypes and patterns of genome evolution in lymphoma revealed by circulating tumor DNA. Sci Transl Med. 2016;8:364ra155
Bessi L, Viailly PJ, Bohers E, Ruminy P, Maingonnat C, Bertrand P, et al. Somatic mutations of cell-free circulating DNA detected by targeted next-generation sequencing and digital droplet PCR in classical Hodgkin lymphoma. Leuk Lymphoma. 2019;60:498–502.
Camus V, Stamatoullas A, Mareschal S, Viailly PJ, Sarafan-Vasseur N, Bohers E, et al. Detection and prognostic value of recurrent exportin 1 mutations in tumor and cell-free circulating DNA of patients with classical Hodgkin lymphoma. Haematologica. 2016;101:1094–101.
Oki Y, Neelapu SS, Fanale M, Kwak LW, Fayad L, Rodriguez MA, et al. Detection of classical Hodgkin lymphoma specific sequence in peripheral blood using a next-generation sequencing approach. Br J Haematol. 2015;169:689–93.
Spina V, Bruscaggin A, Cuccaro A, Martini M, Di Trani M, Forestieri G, et al. Circulating tumor DNA reveals genetics, clonal evolution and residual disease in classical Hodgkin lymphoma. Blood. 2018;131:2413–25.
Vandenberghe P, Wlodarska I, Tousseyn T, Dehaspe L, Dierickx D, Verheecke M, et al. Non-invasive detection of genomic imbalances in Hodgkin/Reed-Sternberg cells in early and advanced stage Hodgkin’s lymphoma by sequencing of circulating cell-free DNA: a technical proof-of-principle study. Lancet Haematol. 2015;2:e55–65.
Hasenclever D, Kurch L, Mauz-Korholz C, Elsner A, Georgi T, Wallace H, et al. qPET—a quantitative extension of the Deauville scale to assess response in interim FDG-PET scans in lymphoma. Eur J Nucl Med Mol Imaging. 2014;41:1301–8.
Bea S, Valdes-Mas R, Navarro A, Salaverria I, Martin-Garcia D, Jares P, et al. Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. Proc Natl Acad Sci USA. 2013;110:18250–5.
Clipson A, Wang M, de Leval L, Ashton-Key M, Wotherspoon A, Vassiliou G, et al. KLF2 mutation is the most frequent somatic change in splenic marginal zone lymphoma and identifies a subset with distinct genotype. Leukemia. 2015;29:1177–85.
Green MR, Gentles AJ, Nair RV, Irish JM, Kihira S, Liu CL, et al. Hierarchy in somatic mutations arising during genomic evolution and progression of follicular lymphoma. Blood. 2013;121:1604–11.
Green MR, Kihira S, Liu CL, Nair RV, Salari R, Gentles AJ, et al. Mutations in early follicular lymphoma progenitors are associated with suppressed antigen presentation. Proc Natl Acad Sci USA. 2015;112:E1116–1125.
Guieze R, Robbe P, Clifford R, de Guibert S, Pereira B, Timbs A, et al. Presence of multiple recurrent mutations confers poor trial outcome of relapsed/refractory CLL. Blood. 2015;126:2110–7.
Kiel MJ, Velusamy T, Betz BL, Zhao L, Weigelin HG, Chiang MY, et al. Whole-genome sequencing identifies recurrent somatic NOTCH2 mutations in splenic marginal zone lymphoma. J Exp Med. 2012;209:1553–65.
Kridel R, Meissner B, Rogic S, Boyle M, Telenius A, Woolcock B, et al. Whole transcriptome sequencing reveals recurrent NOTCH1 mutations in mantle cell lymphoma. Blood. 2012;119:1963–71.
Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell. 2013;152:714–26.
Li H, Kaminski MS, Li Y, Yildiz M, Ouillette P, Jones S, et al. Mutations in linker histone genes HIST1H1 B, C, D, and E; OCT2 (POU2F2); IRF8; and ARID1A underlying the pathogenesis of follicular lymphoma. Blood. 2014;123:1487–98.
Lohr JG, Stojanov P, Carter SL, Cruz-Gordillo P, Lawrence MS, Auclair D, et al. Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell. 2014;25:91–101.
Lohr JG, Stojanov P, Lawrence MS, Auclair D, Chapuy B, Sougnez C, et al. Discovery and prioritization of somatic mutations in diffuse large B-cell lymphoma (DLBCL) by whole-exome sequencing. Proc Natl Acad Sci USA. 2012;109:3879–84.
Love C, Sun Z, Jima D, Li G, Zhang J, Miles R, et al. The genetic landscape of mutations in Burkitt lymphoma. Nat Genet. 2012;44:1321–5.
Morin RD, Mungall K, Pleasance E, Mungall AJ, Goya R, Huff RD, et al. Mutational and structural analysis of diffuse large B-cell lymphoma using whole-genome sequencing. Blood. 2013;122:1256–65.
Okosun J, Bodor C, Wang J, Araf S, Yang CY, Pan C, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014;46:176–81.
Pasqualucci L, Trifonov V, Fabbri G, Ma J, Rossi D, Chiarenza A, et al. Analysis of the coding genome of diffuse large B-cell lymphoma. Nat Genet. 2011;43:830–7.
Quesada V, Ramsay AJ, Lopez-Otin C. Chronic lymphocytic leukemia with SF3B1 mutation. N Engl J Med. 2012;366:2530.
Richter J, Schlesner M, Hoffmann S, Kreuz M, Leich E, Burkhardt B, et al. Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nat Genet. 2012;44:1316–20.
Rossi D, Trifonov V, Fangazio M, Bruscaggin A, Rasi S, Spina V, et al. The coding genome of splenic marginal zone lymphoma: activation of NOTCH2 and other pathways regulating marginal zone development. J Exp Med. 2012;209:1537–51.
Schmitz R, Young RM, Ceribelli M, Jhavar S, Xiao W, Zhang M, et al. Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature. 2012;490:116–20.
Wang L, Lawrence MS, Wan Y, Stojanov P, Sougnez C, Stevenson K, et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N Engl J Med. 2011;365:2497–506.
Zhang J, Grubor V, Love CL, Banerjee A, Richards KL, Mieczkowski PA, et al. Genetic heterogeneity of diffuse large B-cell lymphoma. Proc Natl Acad Sci USA. 2013;110:1398–403.
Zhang J, Jima D, Moffitt AB, Liu Q, Czader M, Hsi ED, et al. The genomic landscape of mantle cell lymphoma is related to the epigenetically determined chromatin state of normal B cells. Blood. 2014;123:2988–96.
Reichel J, Chadburn A, Rubinstein PG, Giulino-Roth L, Tam W, Liu Y, et al. Flow sorting and exome sequencing reveal the oncogenome of primary Hodgkin and Reed-Sternberg cells. Blood. 2015;125:1061–72.
Tiacci E, Penson A, Schiavoni G, Ladewig E, Fortini E, Wang Y, et al. New recurrently mutated genes in classical Hodgkin lymphoma revealed by whole-exome sequencing os microdissected tumor cells. Blood. 2016;218:1088.
Raczy C, Petrovski R, Saunders CT, Chorny I, Kruglyak S, Margulies EH, et al. Isaac: ultra-fast whole-genome secondary analysis on Illumina sequencing platforms. Bioinformatics. 2013;29:2041–3.
Kuppers R, Dalla-Favera R. Mechanisms of chromosomal translocations in B cell lymphomas. Oncogene. 2001;20:5580–94.
Martin-Subero JI, Klapper W, Sotnikova A, Callet-Bauchu E, Harder L, Bastard C, et al. Chromosomal breakpoints affecting immunoglobulin loci are recurrent in Hodgkin and Reed-Sternberg cells of classical Hodgkin lymphoma. Cancer Res. 2006;66:10332–8.
Tiacci E, Ladewig E, Schiavoni G, Penson A, Fortini E, Pettirossi V, et al. Pervasive mutations of JAK-STAT pathway genes in classical Hodgkin lymphoma. Blood. 2018;131:2454–65.
Mansouri L, Noerenberg D, Young E, Mylonas E, Abdulla M, Frick M, et al. Frequent NFKBIE deletions are associated with poor outcome in primary mediastinal B-cell lymphoma. Blood. 2016;128:2666–70.
Pasqualucci L, Neumeister P, Goossens T, Nanjangud G, Chaganti RS, Kuppers R, et al. Hypermutation of multiple proto-oncogenes in B-cell diffuse large-cell lymphomas. Nature. 2001;412:341–6.
Lennerz JK, Hoffmann K, Bubolz AM, Lessel D, Welke C, Ruther N, et al. Suppressor of cytokine signaling 1 gene mutation status as a prognostic biomarker in classical Hodgkin lymphoma. Oncotarget. 2015;6:29097–110.
Mottok A, Renne C, Willenbrock K, Hansmann ML, Brauninger A. Somatic hypermutation of SOCS1 in lymphocyte-predominant Hodgkin lymphoma is accompanied by high JAK2 expression and activation of STAT6. Blood. 2007;110:3387–90.
Schif B, Lennerz JK, Kohler CW, Bentink S, Kreuz M, Melzner I, et al. SOCS1 mutation subtypes predict divergent outcomes in diffuse large B-Cell lymphoma (DLBCL) patients. Oncotarget. 2013;4:35–47.
Klein U, Goossens T, Fischer M, Kanzler H, Braeuninger A, Rajewsky K, et al. Somatic hypermutation in normal and transformed human B cells. Immunol Rev. 1998;162:261–80.
Sasanelli M, Meignan M, Haioun C, Berriolo-Riedinger A, Casasnovas RO, Biggi A, et al. Pretherapy metabolic tumour volume is an independent predictor of outcome in patients with diffuse large B-cell lymphoma. Eur J Nucl Med Mol Imaging. 2014;41:2017–22.
Tout M, Casasnovas O, Meignan M, Lamy T, Morschhauser F, Salles G, et al. Rituximab exposure is influenced by baseline metabolic tumor volume and predicts outcome of DLBCL patients: a Lymphoma Study Association report. Blood. 2017;129:2616–23.
Dukers DF, Meijer CJ, ten Berge RL, Vos W, Ossenkoppele GJ, Oudejans JJ. High numbers of active caspase 3-positive Reed-Sternberg cells in pretreatment biopsy specimens of patients with Hodgkin disease predict favorable clinical outcome. Blood. 2002;100:36–42.
Georgiadi EC, Sachinis N, Dimtsas G, Vassilakopoulos TP, Kittas C, Doussis-Anagnostopoulou IA. Evaluation of apoptosis in classical Hodgkin’s lymphoma comparing different methods. J BUON. 2012;17:746–52.
Lorenzen J, Thiele J, Fischer R. The mummified Hodgkin cell: cell death in Hodgkin’s disease. J Pathol. 1997;182:288–98.
Kuppers R, Engert A, Hansmann ML. Hodgkin lymphoma. J Clin Invest. 2012;122:3439–47.
Spina V, Bruscaggin A, Cuccaro A, Martini M, Di Trani M, Forestieri G, et al. Circulating tumor DNA reveals genetics, clonal evolution, and residual disease in classical Hodgkin lymphoma. Blood. 2018;131:2413–25.
Acknowledgements
We like to thank Annika Habbaba, Sebastian Schäfer, Cindy Arnold, Katharina Sack and Stefanie Rudloff for excellent technical assistance and Stefanie Avondstondt and Dieter Hoffmann from the Pediatric Hodgkin Lymphoma reference center for providing the clinical data documentation. The EuroNet-PHL-C2 trial is supported by a grant from the Deutsche Krebshilfe (Grant No.110674). This study was supported by the Deutsche Krebshilfe (Grant no. 111711), the Rhön Klinikum AG (RKA FL_21), the Elternverein für leukämie- und krebskranke Kinder Gießen and the Kinderkrebshilfe Oldtimer Markt Mainz.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Desch, AK., Hartung, K., Botzen, A. et al. Genotyping circulating tumor DNA of pediatric Hodgkin lymphoma. Leukemia 34, 151–166 (2020). https://doi.org/10.1038/s41375-019-0541-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0541-6
This article is cited by
-
Distinct Hodgkin lymphoma subtypes defined by noninvasive genomic profiling
Nature (2024)
-
Molecular characterization of an embryonal rhabdomyosarcoma occurring in a patient with Kabuki syndrome: report and literature review in the light of tumor predisposition syndromes
Familial Cancer (2023)
-
Circulating tumor DNA in B-cell lymphoma: technical advances, clinical applications, and perspectives for translational research
Leukemia (2022)
-
Circulating tumor DNA in Hodgkin lymphoma
Annals of Hematology (2022)
-
Molecular biology of Hodgkin lymphoma
Leukemia (2021)