Abstract
Acute myeloid leukemia (AML) is a devastating disease, with the majority of patients dying within a year of diagnosis. For patients with relapsed/refractory AML, the prognosis is particularly poor with currently available treatments. Although genetically heterogeneous, AML subtypes share a common differentiation arrest at hematopoietic progenitor stages. Overcoming this differentiation arrest has the potential to improve the long-term survival of patients, as is the case in acute promyelocytic leukemia (APL), which is characterized by a chromosomal translocation involving the retinoic acid receptor alpha gene. Treatment of APL with all-trans retinoic acid (ATRA) induces terminal differentiation and apoptosis of leukemic promyelocytes, resulting in cure rates of over 80%. Unfortunately, similarly efficacious differentiation therapies have, to date, been lacking outside of APL. Inhibition of dihydroorotate dehydrogenase (DHODH), a key enzyme in the de novo pyrimidine synthesis pathway, was recently reported to induce differentiation of diverse AML subtypes. In this report we describe the discovery and characterization of BAY 2402234 - a novel, potent, selective and orally bioavailable DHODH inhibitor that shows monotherapy efficacy and differentiation induction across multiple AML subtypes. Herein, we present the preclinical data that led to initiation of a phase I evaluation of this inhibitor in myeloid malignancies.
Similar content being viewed by others
Introduction
AML is the second most common leukemia in adults, with the highest incidence in patients over 65 years of age. The 5 year overall survival of elderly patients with AML is <10%, indicating a need for more effective and well-tolerated therapies for AML [1].
During normal hematopoiesis, myeloid cells transition from hematopoietic progenitor cells that have a high proliferative capability to terminally differentiated cells with low proliferation rates and limited lifespans. During leukemogenesis, mutational events in hematopoietic stem cells disrupt normal differentiation and lead to a maturation arrest, which maintains leukemic cells in an undifferentiated state with the capacity to self-renew. As this differentiation block can occur at different progenitor stages in the setting of a variety of mutational events, AML is a highly heterogeneous disease [2]. This creates significant challenges to identifying targeted therapies that are effective against multiple AML subtypes.
Treatment of AML can be addressed from two different angles; by directly inducing apoptosis of blast cells with cytotoxic chemotherapy, or by inducing terminal differentiation of blast cells and thereby halting their capacity to self-renew. Induction of apoptosis with high dose cytarabine and daunorubicin has been the 1st line standard-of-care for many decades. However, many elderly patients are unable to receive high-dose chemotherapy due to its significant toxicity. More recently, therapies have been developed that target specific oncoproteins that mediate the differentiation arrest of AML blast cells, thereby inducing blast cell differentiation. These therapies, which include ATRA and arsenic trioxide (ASO) for APL and Enasidenib and Ivosidenib for IDH2- and IDH1-mutated AML, respectively, are well-tolerated even in elderly patients with multiple comorbidities [3,4,5]. Although data on the long-term outcomes of patients treated with Enasidenib or Ivosidenib are not yet known, the initial responses observed in patients with relapsed/refractory IDH mutant AML are encouraging. The overall response rate of patients treated with Enasidenib is 38.8%, with 19.6% of patients achieving complete remission [5]. The overall response rate for Ivosidenib-treated patients is 41.6%, with 21.6% of patients achieving complete remission [4]. In APL, the success of differentiation therapy is even more impressive. Treatment with ATRA, ASO and chemotherapy has a cure rate of over 80%, and many elderly patients treated with ATRA and ASO, or ASO alone, achieve durable remissions [6, 7].
Given the lack of effective differentiation therapy outside of APL (10–15% of AML) and, potentially, IDH mutant AML (18–22% of AML), we undertook a phenotypic screen to identify enzymes whose inhibition can overcome the differentiation blockade of AML cells transformed by constitutive expression of HoxA9. In this phenotypic screen, DHODH was the top hit [8]. DHODH is an enzyme that localizes to the inner mitochondrial membrane. It is the fourth and rate limiting step of the de novo pyrimidine synthesis pathway, converting dihydroorotate (DHO) to orotate. There are only two known mechanisms by which cells can acquire pyrimidines – de novo synthesis and recycling via the salvage pathway. Previous experiments revealed that inhibition of this critical enzyme causes a release of the differentiation blockade of blast cells across diverse AML subtypes [8, 9].
To target the differentiation arrest in AML in a therapeutically tractable manner, we developed BAY 2402234, a novel, highly potent and selective DHODH inhibitor. Herein, we present data on the preclinical activity of BAY 2402234, showing that BAY 2402234 has strong monotherapy efficacy and induces differentiation across multiple AML xenograft and PDX models in vitro and in vivo. These very promising preclinical observations are the rationale for a clinical evaluation of BAY 2402234 in myeloid malignancies that is currently ongoing (NCT 0340726).
Material and methods
Experimental procedures can be found in the Supplementary data.
Results
Properties of BAY 2402234
BAY 2402234 (Fig. 1a) is a novel potent and selective DHODH inhibitor that dose-dependently inhibits human full-length DHODH enzyme with an IC50 of 1.2 nM (Fig. 1b). BAY 2402234 is a neutral small molecule selective for DHODH with a LogD value of 2.7 and a molecular weight of 520.84 g/mol (Supplementary Table 1). BAY 2402234 has moderate plasma protein binding (fu = 9.9%), low blood clearance (CLb = 0.87 L/h/kg) and a high volume of distribution Vss = 3.0 L/kg. The oral bioavailability in mice is complete (F = 101%) and the half-life is 4.1 h.
BAY 2402234 is a novel potent DHODH inhibitor that binds the ubiquinone pocket of DHODH. a Chemical structure of BAY 2402234. b Biochemical inhibition of full-length human DHODH by BAY 2402234. c Crystal structure of DHODH complexed with BAY 2402234. Strong electron density of BAY 2402234 (green) is visible between Helix A and B, contoured at 1.0σ (blue). The locations of the substrate (orotate; orange) and the prosthetic group (flavin mononucleotide/FMN; magenta) are indicated. d Protein-ligand interactions of BAY 2402234 (green) and DHODH. The alternative conformation of Arg136 is highlighted in cyan
Crystal structure of DHODH and BAY 2402234
To elucidate the structural details of inhibitor binding, recombinant human DHODH was complexed and co-crystallized with BAY 2402234. Refinement of the DHODH protein model against high-quality X-ray diffraction data revealed strong electron density fitting of the inhibitor, a bound flavin mononucleotide (FMN) and an orotate molecule (Fig. 1c, Supplementary Table 2). Overall, the protein exhibits the same fold and secondary structure topology as published X-ray structures of DHODH, with a central α/β-barrel and a small N-terminal domain crucial for inhibitor binding [10].
BAY 2402234 binds the ubiquinone binding site of DHODH between the N-terminal helices (Helix A & B), thereby demonstrating its molecular mode of action. The triazolone group of the inhibitor binds to the deep and more hydrophilic end of the cofactor pocket through direct hydrogen bonding with Tyr356 and Gln47 (Fig. 1d). The OH-group of BAY 2402234 interacts with Thr360 via a bound water molecule. At the entrance of the binding site the compound has extensive hydrophobic interactions with a large set of non-polar residues (Tyr38, Leu42, Met43, Leu46, Leu50, Leu58, Ala59, Phe62, Leu67, Leu68, Pro69, Met111, Leu359, and Pro364). The crystal structure also predicts a non-classical hydrogen bond between the hydroxyl group of Thr63 (H-bond donor) and the fluorine of the terminal phenyl group (H-bond acceptor). As previously observed, the non-coordinated Arg136 exhibits flexibility [11] and it is shown modeled in two different conformations (Fig. 1d).
BAY 2402234 induces differentiation and inhibits proliferation in AML cell lines across multiple AML subtypes
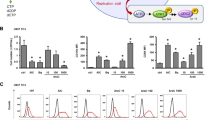
To investigate the effect of BAY 2402234 on AML cell differentiation, the ability of the drug to upregulate the differentiation marker CD11b was assessed in MOLM-13 and HEL cells. In both cell lines, dose-dependent upregulation of CD11b was observed after BAY 2402234 treatment (Fig. 2a, b, Supplementary Fig. 1a), with EC50s of 3.16 nM in MOLM-13 cells and 0.96 nM in HEL cells (Supplementary Table 3), and this effect was rescued by the addition of 100 µM exogenous uridine. Uridine bypasses the requirement for de novo pyrimidine synthesis by feeding into the salvage pathway, thereby negating the need for DHODH. The observation that addition of uridine inhibits induction of differentiation by BAY 2402234 indicates that the drug is acting ‘on-target’. BAY 2402234 similarly induces differentiation of a panel of additional AML cell lines at doses of 8.8 × 10−10 M to 9 × 10−9 M (Supplementary Table 3).
BAY 2402234 induces differentiation and inhibits proliferation of leukemia cells in vitro. a, b MOLM-13 (a) and HEL (b) cells were treated with increasing concentrations of BAY 2402234 for 4 days in the presence (blue) or absence (green) of 100 µM uridine and CD11b fluorescence intensity was then measured by FACS (RFU = relative fluorescence units). c TF-1 cells were treated for 3 or 6 days with the indicated concentrations of BAY 2402234 in the presence or absence of uridine and the percent CD11b-positive cells was determine by FACS. d THP-1 cells were treated with increasing concentrations of BAY 2402234 in the presence or absence of 100 µM uridine for 3 days and the number of surviving cells was then determined by Cell Titer Glow Assay (RLU = relative luminescence units). e–g TF-1 cells were treated with the indicated concentrations of BAY 2402234 in the presence or absence of uridine for a total of 9 days. The number of viable cells on days 3, 6 and 9 (e), the cell cycle state of the cells on day 3 (f) and the percent apoptotic cells on days 3 and 6 (g) were assessed. By day 9, no viable BAY 2402234-treated cells were detectable. Graphs f and g are representative results from three biological replicates
TF-1 cells were used to analyze the kinetics of CD11b upregulation by BAY 2402234. While an increase in CD11b was observed after 3 days of treatment, the increase after 6 days was more prominent (Fig. 2c, Supplementary Fig. 1b). Again, exogenous uridine was able to rescue the differentiation phenotype. Differentiation induction by BAY 2402234 treatment was further confirmed by Giemsa-Wright stainings for morphologic features of differentiation (Supplementary Fig. 2a) and by flow cytometry for CD14, another marker of myeloid differentiation (Supplementary Fig. 2b–d).
We also evaluated whether inhibition of DHODH has anti-proliferative effects in vitro. BAY 2402234 inhibited the proliferation of THP-1 cells with an IC50 of 2.6 nM (Fig. 2d) and inhibited the proliferation of nine other leukemia cell lines representing diverse AML subtypes, with a range of IC50s from 8.1 × 10−11 M to 8.2 × 10−9 M (Supplementary Table 4). In all cases, the addition of uridine completely abolished the anti-proliferative effect of BAY 2402234, confirming the on-target activity of BAY 2402234.
Finally, we asked if BAY 2402234 induces cell cycle arrest and apoptosis in AML cells. We treated TF-1 cells with BAY 2402234 and analyzed the proliferation and apoptosis of the cells at three different concentrations of the inhibitor and over multiple days. Treatment with the inhibitor dose-dependently slowed cell doubling (Fig. 2e), induced G2-M cell cycle arrest (Fig. 2f) and induced apoptosis (Fig. 2g) of the cells in a dose- and time-dependent manner. These effects were, once again, fully rescued by uridine supplementation.
The human AML cell lines used in these studies harbor many genetic alterations that are highly recurrent in AML (Supplementary Table 5a and 5b). Six of the cell lines (NOMO-1, HEL, HL-60, SKM-1, THP-1, and TF-1) are TP53 mutant and three of the cell lines (EOL-1, MOLM-13, and MV4-11) are TP53 wild-type. Four of the cell lines (NOMO-1, MOLM-13, MV4-11, and THP-1) harbor MLL translocations; two (NOMO-1 and SKM-1) harbor KRAS mutations; three (HL-60, THP-1, and TF-1) harbor NRAS mutations. Oncogenic mutations in JAK2 (HEL), TET2 (HEL and SKM-1), ASXL1 (NOMO-1, SKM-1, and TF-1) and FLT3 (MOLM-13 and MV4-11) are also represented. Because upregulation of p53 has been proposed as a mechanism of anti-tumor activity by DHODH inhibition [12], we assessed whether TP53 mutation status affects the IC50 of BAY 2402234 in AML cell lines. We did not identify any correlation between sensitivity to BAY 2402234 and the TP53 mutation status of the AML cell lines.
BAY 2402234 shows strong monotherapy efficacy and induces differentiation in AML xenograft models in vivo
Dose-escalation studies were performed to determine the maximum tolerated dose (MTD) of BAY 2402234 in mice and daily dosing of up to 5 mg/kg for 10 days was found to be well tolerated (data not shown). We next assessed the anti-tumor activity of BAY 2402234 in three subcutaneous AML xenograft models. In MOLM-13 xenografts, a significant anti-tumor effect was observed at all four treatment doses (Fig. 3a). Unbound drug exposure after daily continuous treatment with BAY 2402234 was measured in murine plasma and was found to be dose- and time-dependent (Fig. 3b). Plasma levels of DHO were measured as a pharmacodynamic marker of target engagement. We observed a dose-dependent increase in DHO levels in BAY 2402234-treated mice (Fig. 3c), demonstrating DHODH inhibition in vivo. Likewise, in MV4-11 (Fig. 3d) and THP-1 (Supplementary Fig. 3a) xenografts, we saw significant anti-tumor efficacy in all dosing groups, confirming the strong anti-tumor effect of BAY 2402234 in AML mouse models.
BAY 2402234 shows monotherapy efficacy, target engagement, and differentiation induction in AML xenografts in vivo. a Serial measurements of tumor volumes in subcutaneous MOLM-13 AML xenografts treated daily with the indicated doses of BAY 2402234 starting after randomization on day 4 post transplantation (mean ± SEM, n = 10 mice/group; One way ANOVA on log transformed tumor volume). b, c Blood was drawn from mice treated daily for 16 days with the indicated doses of BAY 2402234 1, 3, 6, and 24 h after the final dose (mean, n = 2–3 mice/timepoint). Plasma levels of unbound BAY 2402234 (b) and plasma levels of DHO (c) were assessed to determine the pharmocokinetic and pharmacodynamic properties, respectively, of BAY 2402234. d Serial measurements of tumor volumes in subcutaneous MV4-11 AML xenografts treated daily with the indicated doses of BAY 2402234 starting after randomization on day 10 post transplantation (mean ± SEM, n = 12 mice/group; One way ANOVA on log transformed samples). e Measurement of CD11b fluorescence intensity by FACS of MV4-11 xenograft tumors after daily treatment with BAY 2402234 for 6 days (mean±SEM, n = 7; t-test) (RFU = relative fluorescence units). f Survival of disseminated HL-60 xenograft mice (n = 10/group; logrank Mantel-Cox test) treated daily with 5 mg/kg BAY 2402234 compared to standard AML dosing of cytarabine. In both cases, treatment was started 5 days post transplantation. g Survival of disseminated MV4-11 xenograft mice (n = 10/group; logrank Mantel-Cox test) treated daily with vehicle control or 4 mg/kg BAY 2402234 starting after randomization on day 3 post transplantation. The experiment was stopped on day 46 of treatment and peripheral blood and bone marrow from the surviving mice were harvested. h Determination of post-treatment tumor burden as a function of percent human HLA-ABC-positive cells in the peripheral blood and bone marrow of MV4-11 tumor bearing mice still alive on day 46 of treatment. *p < 0.05; **p < 0.01; ***p < 0.001
We also assessed the ability of BAY 2402234 to induce AML differentiation in vivo. MV4-11 tumors were harvested after daily dosing of 5 mg/kg BAY 2402234 for 6 days, and cell surface expression of CD11b was measured ex vivo. We detected an increase in CD11b expression in tumor cells from BAY 2402234-treated mice compared to vehicle-treated mice (Fig. 3e). Similar effects were seen in THP-1 xenografts treated with 5 mg/kg BAY 2402234 for 8 days (Supplementary Fig. 3b) and in MOLM-13 xenograft tumors harvested 24 h after a single 5 mg/kg dose of BAY 2402234 (Supplementary Fig. 3c).
The anti-tumor activity of BAY 2402234 was also evaluated in three systemic AML xenograft models transplanted by tail vein injection. In HL-60 xenografts, treatment with BAY 2402234 nearly doubled the median survival of mice from 13 to 25 days and showed improved efficacy compared to treatment with cytarabine (Fig. 3f). In MV4-11 xenografts, 70% of mice treated with BAY 2402234 were still alive at the study-endpoint (day 46 of treatment) whereas all of the vehicle-treated mice died by day 28 (Fig. 3g). The surviving MV4-11 xenografts were sacrificed on day 46 of treatment and were found to have <1% residual disease, as determined by flow cytometry for human HLA-ABC-positive cells in the peripheral blood and bone marrow of the mice (Fig. 3h). In a disseminated MOLM-13 xenograft model, tumor burden in the peripheral blood, bone marrow and spleens of control and BAY 2402234-treated mice was analyzed after 8 days of treatment. At that time point, over 40% of the peripheral blood cells in the control mice were human HLA-ABC-positive leukemia cells whereas human leukemia cells represented about 4% of circulating cells in the BAY 2402234 treated mice (Supplementary Fig. 4a). In the bone marrows of the control mice, 78% of the cells were human AML cells compared to 39% in the BAY 2402234 treated mice (Supplementary Fig. 4b); and in the spleens of the control mice, 44% of the cells were human AML cells compared to 6% in the treated mice (Supplementary Fig. 4c). Co-staining for human CD11b showed a dramatic increase in the percent CD11b-positive cells in the bone marrow of BAY 2402234-treated mice (Supplementary Fig. 4b), suggesting that BAY 2402234 induces AML cell differentiation in vivo.
BAY 2402234 prolongs survival and induces differentiation in AML PDX models
To test whether BAY 2402234 has therapeutic efficacy in PDX models, we tested five disseminated AML PDX models using endpoints of overall survival (AM5512, AM7577, and AM8096) and tumor burden at predefined time points (AML11655 and AML6252). These PDX models harbor mutations in a number of genes, including IDH1, IDH2, and NPM1, which are recurrently mutated in AML (Supplementary Table 6).
We observed increased survival of treated mice in all three survival endpoint studies: the AM5512 model (Fig. 4a), the AM7577 model (Fig. 4b), and the AM8096 model (Fig. 4c). In the AM8096 model, all of the vehicle-treated mice died by day 20 of treatment whereas 40% of the BAY 2402234-treated animals were still alive when the study was terminated after 47 days of treatment followed by a 4 week observation period.
BAY 2402234 shows monotherapy efficacy and differentiation induction in AML PDX models. a–c Survival of mice transplanted with AML PDX lines AM5512 (a), AM7577 (b), and AM8096 (c) treated daily with vehicle control or with 4 mg/kg BAY 2402234 (n = 5/group; logrank Mantel-Cox test). d, e Percent human CD45+ cells in the peripheral blood of AML PDX models AML11655 (d) and AML6252 (e) treated daily for 36 days (d) or 50 days (e) with vehicle control or with 4 mg/kg BAY 2402234 (mean ± SEM, n = 5; t-test). f, g Percent hCD11b/hCD45 double-positive AML cells in the peripheral blood of AML PDX models AML11655 (f) and AML6252 (g) treated daily with 4 mg/kg BAY 2402234 for 36 (f) and 50 (g) days (mean ± SEM, n = 5; t-test). In all cases, transplantation was by tail vein injection. Treatment was started for AM5512 and AM7577 models when 2–5% human CD45+ cells were detectable in the peripheral blood of the mice; treatment was started for AM8096, AML6252, and AML11655 models at predefined times post-transplantation (40, 3, and 4 days, respectively). *p < 0.05; **p < 0.01; ***p < 0.001
In the tumor burden endpoint studies, the percent circulating human CD45-positive AML cells was significantly decreased at study end in AML11655 (Fig. 4d) and AML6252 (Fig. 4e)-transplanted mice treated with BAY 2402234 compared to mice treated with vehicle control. Moreover, the spleen weights of BAY 2402234-treated mice were significantly reduced compared to vehicle-treated animals (Supplementary Fig. 5a, b). Together, these data indicate that BAY 2402234 significantly reduced tumor burden in AML PDX models. Co-staining of cells for human CD45 and human CD11b showed a significant increase in double-positive cells in BAY 2402234-treated mice compared to vehicle-treated mice (Fig. 4f, g), suggesting that BAY 2402234 induces AML cell differentiation in vivo in PDX models.
Single treatment with BAY 2402234 induces profound transcriptional changes
To elucidate the mode of action of BAY 2402234, MOLM-13 xenograft-bearing mice were treated with a single 5 mg/kg dose of BAY 2402234 and tumors were collected at five time points (3, 6, 24, 36, and 48 h) after treatment with BAY 2402234 or vehicle control and were subjected to RNA-sequencing. Gene expression analysis revealed that a single treatment with BAY 2402234 results in transcriptional changes (differential expression of 1.5-fold or more) as early as 3 and 6 h post treatment, and that the magnitude of transcriptional changes increases over time (Supplementary Table 7).
Gene set enrichment analysis (GSEA) was performed comparing the transcriptional profiles of tumors treated with BAY 2402234 or vehicle control for 24 h (Supplementary Table 8). This analysis found that BAY 2402234 treatment induces a similar transcriptional signature as that induced by knockdown of HOXA9 in MOLM-14 cells (Fig. 5a). Of note, HoxA9 knockdown has been shown to induce differentiation, cell cycle arrest and apoptosis of AML cells [13]. The transcriptional signature induced by BAY 2402234 also positively correlates with apoptosis-associated gene transcription (Fig. 5b) and negatively correlates with MYC target gene expression (Fig. 5c). Correlations with p53 pathway genes, E2F target genes and G2M checkpoint genes were also observed (Supplementary Fig. 6). Overall, this analysis further supports our finding that BAY 2402234 treatment induces the differentiation, cell cycle arrest and apoptosis of AML cells.
BAY 2402234 induces transcriptional changes linked to differentiation, apoptosis, and MYC repression. GSEA analysis of RNASeq read counts from MOLM-13 xenograft tumors harvested 24 h after a single treatment with 5 mg/kg BAY 2402234 or vehicle control. Enrichment plots are shown for signatures of (a) HOXA9 knockdown in MOLM-14 cells, (b) apoptosis-associated genes, and (c) MYC target genes
Examination of BAY 2402234-induced transcriptional changes over time revealed that a number of genes that play a role in the maturation block of AML cells are dysregulated by BAY 2402234 treatment in a time-dependent manner. These include HOXA9 [14], FLT3 [15], and c-MYC [16] (Supplementary Fig. 7a). Interestingly, we also detected rapid and significant downregulation of HMGA1 expression in BAY 2402234 treated cells (Supplementary Fig. 7b). HMGA1 has been reported to play an important role in the maintainence of the self-renewal in cancer stem cells [17]. Several other chromatin modifying enzymes, including KMT2A, KAT6A, and EZH2, were also dysregulated, albeit less dramatically, by BAY 2402234 treatment (Supplementary Fig. 7b). The effect of BAY 2402234 on EZH2 expression in vitro and in vivo was confirmed by real-time PCR (Supplementary Fig. 8a, b).
Cell cycle exit is critical for terminal differentiation [18] and has been linked to DHODH inhibition in other cancer cell types in vitro [19, 20]. We examined the kinetics of dysregulation of several cell cycle regulatory genes by BAY 2402234. We found that P21/CDKN1A expression is induced within hours of BAY 2402234 treatment and then drops precipitously, whereas expression of CDK4 and CDK6 begins to decline immediately after BAY 2402234 treatment (Supplementary Fig. 7c). The effect of BAY 2402234 on P21/CDKN1A expression in vitro and in vivo was confirmed by real-time PCR (Supplementary Fig. 8a, b). These data are consistent with our observation that inhibition of DHODH inhibits the proliferation and induces the cell cycle arrest of AML cells in vitro and in vivo.
Finally, and in line with previous reports that DHODH inhibition reduces uridine diphosphate N-acetylglucosamine (UDP-GlcNAc) levels [8], we identified GNPNAT1 and UAP1 as being significantly downregulated by BAY 2402234 treatment (Supplementary Fig. 7d). These genes play a major role in UDP sugar metabolism and O-GlcNAc production.
Overall, the transcriptional changes induced by BAY 2402234 indicate a profound effect of inhibition of DHODH on multiple intracellular pathways, resulting in changes in cellular proliferation, differentiation and apoptosis.
Single treatment with BAY 2402234 induces profound proteomic changes
MOLM-13 AML xenograft tumor samples were used to investigate changes at the protein and phospho-protein levels in tumors after a single 5 mg/kg dose of BAY 2402234. Tumors were collected at 24 and 48 h after treatment of mice with BAY 2402234 or vehicle control and the protein lysates were then screened using an array of 622 antibodies (Supplementary Table 9). This analysis revealed similar effects of BAY 2402234 on mediators of proliferation and apoptosis as the transcriptional analysis (Fig. 6a, b). Several proteins linked to induction of apoptosis, including Caspase 3, Caspase 7, and cleaved PARP [21, 22] were among the most highly upregulated proteins at one or both of the sampled time points (Fig. 6a, Supplementary Table 10), and this effect was confirmed by western blot analysis of TF-1 cells treated with BAY 2402234 (Fig. 6c). Consistent with our transcriptomic data, P21/CDKN1A protein expression was also induced by BAY 2402234 treatment (Fig. 6d). Interestingly, BAY 2402234 treatment reduced levels of phospho-GSK3α/β (Fig. 6b, Supplementary Table 10). Inhibition of GSK3α has been found to promote AML cell differentiation and apoptosis and to prolong survival of AML mouse models [23]. BAY 2402234 treatment also reduced the phosphorylation of a number of other kinases that play a role in AML cell proliferation and self-renewal, including phospho-MEK1/2 and phospho-ERK1/2 [24], and phospho-Src and phospho-Lck [25,26,27] (Fig. 6b, Supplementary Table 10). These effects were confirmed by western blot analysis of MOLM-13 xenograft tumors after treatment with a single dose of BAY 2402234 (Fig. 6e). Levels of phospho-CREB Ser133, a downstream target of the MAPK pathway, were also reduced by BAY 2402234 treatment (Fig. 6b, Supplementary Table 10).
BAY 2402234 induces proteomic changes linked to cell cycle arrest, apoptosis, and kinase inactivation. a, b Results of an ex vivo protein screen of MOLM-13 xenograft tumors harvested 24 and 48 h after a single 5 mg/kg dose of BAY 2402234. Signal intensities of proteins involved in apoptosis (a) and kinases involved in proliferation and self-renewal signaling (b) are indicated (mean ± SEM, n = 5 tumors/timepoint; t-test) (RFU = relative fluorescence units). *p < 0.05; **p < 0.01; ***p < 0.001. c Western Blot analysis of cleaved PARP and cleaved Caspase 3 in TF-1 cells treated with the indicated concentrations of BAY 2402234 for the indicated times. TF-1 cells treated with doxorubicin serve as a positive control for apoptosis induction. d Western Blot analysis of p21 expression in MOLM-13 xenograft tumors harvested 24 h after treatment with a single 5 mg/kg dose of BAY 2402234 or with vehicle control. e Western Blot analysis of phospho-ERK1/2 Thr202/Tyr204, phospho-Lck Tyr505, and phospho-Src Tyr416 levels in individual MOLM-13 xenograft tumors harvested 24 or 48 h after treatment with a single 5 mg/kg dose of BAY 2402234 or with vehicle control
Discussion
Several inhibitors of DHODH have been described previously [28], with brequinar sodium and leflunomide/teriflunomide being the most well-known agents tested in humans. The anti-cancer activity of brequinar was tested in the 1990s in solid tumor patients [29,30,31], but brequinar was not given to patients with myeloid malignancies. Moreover, despite preclinical evidence supporting the need for continuous target engagement [32, 33], the clinical trials of brequinar in solid tumors used intermittent dosing schedules similar to those of classical chemotherapy. Leflunomide and its active metabolite teriflunomide are approved for the treatment of rheumatoid arthritis and multiple sclerosis, respectively. Although some encouraging preclinical data on their anti-tumor efficacy have been reported [34], leflunomide and teriflunomide are weak DHODH inhibitors, with IC50s in the low µM range. Additionally, several other targets of these drugs have been described, including receptor tyrosine kinases [35, 36], the aryl hydrocarbon receptor [37], monoamine oxidases [38], and cyclooxygenase 2 [39]. Given the low potency of leflunomide and teriflunomide against DHODH, it is likely that at least some of their described anti-tumor effects are not mediated by DHODH inhibition.
Given our recent findings regarding the effect of DHODH inhibition on AML cell differentiation [8], and given the high unmet clinical need in AML, our novel, potent and selective DHODH inhibitor BAY 2402234 may be of value in the treatment of myeloid malignancies. The molecule possesses many properties suitable for further clinical development. The drug metabolism and pharmacokinetics (DMPK) and physicochemical properties of BAY 2402234 allow for continuous oral dosing, while the crystal structure of the drug bound to DHODH, and the ability of uridine to rescue the effects of the drug, validate its specificity and selectivity for DHODH. Moreover, our in vitro and in vivo data show that BAY 2402234 treatment is both anti-proliferative and induces differentiation in multiple AML xenografts and PDX models that represent a diverse set of AML genotypes. This strongly suggests that BAY 2402234 will have broad therapeutic applicability in myeloid malignancies.
BAY 2402234 shows a dual mode of action of proliferation inhibition and differentiation induction. Interestingly, single-dose treatment with BAY 2402234 in an AML xenograft model induces transcriptomic, proteomic, and phospho-proteomic changes that can be linked to both modes of action, suggesting that both pathways are initiated early on by BAY 2402234 treatment. Whereas inhibition of proliferation can be explained by pyrimidine depletion, the underlying mechanism of induction of differentiation is still unclear. Unlike ATRA and mutant IDH inhibitors, which specifically target oncoproteins that are mechanistically involved in AML differentiation arrest, DHODH inhibition seems to have pleiotropic effects in cells. It is possible that the mechanism of anti-leukemic activity of DHODH inhibition more closely resembles that of other cancer therapies, like the BCL-2 inhibitor Venetoclax, that induce differentiation by targeting pathways critical for AML cell survival [40].
It is notable that BAY 2402234 induces acute downregulation of genes linked to stemness, which is the counterpoint to differentiation. One of the downregulated chromatin remodelers, HMGA1, has been strongly linked to stemness in pluripotent stem cells [41] as well as cancer stem cells [17], while the downregulated histone actelytransferase KAT6A has been reported to be essential for adult hematopoietic stem cell function [42]. Cell cycle exit also plays an important role in differentiation, and repression of one of the downregulated CDKs, CDK6, has been linked to leukemic differentiation [43, 44]. We also noted changes in phospho-Lck, which was previously reported to cooperate in cellular transformation with FLT3/ITD [27] and is linked to T and B cell leukemia [45, 46], as well as changes in the phosphorylation status of Src kinases, whose activities have been linked to cancer stemness in breast cancer [25, 26]. Finally, another notable effect of BAY 2402234 is the downregulation of key enzymes in UDP GlcNAc metabolism, which may explain our previous observation of global decreases in protein N-acetyl glycosylation after DHODH inhibition [8]. Given the role of O-GlcNAc in posttranslational modifications [47] and in chondrogenic, osteogenic and adipogenic differentiation [48,49,50,51], it is certainly plausible that O-GlcNAcylation also plays a role in myeloid differentiation. Interestingly, DHODH-induced reduction of O-GlcNAc was recently postulated to reduce c-Myc stability and thereby promote induction of differentiation [52].
Our findings demonstrate that the effects of DHODH inhibition extend far beyond pyrimidine depletion. Further investigations of the pleiotropic effects of DHODH inhibition will be required to fully understand how DHODH inhibition leads to induction of differentiation, cell cycle arrest and apoptosis in AML cells.
In summary, the novel DHODH inhibitor BAY 2402234 is active against diverse subtypes of AML in vitro and in vivo and shows a dual mode of action of proliferation inhibition and differentiation induction. Moreover, BAY 2402234 has favorable physicochemical and pharmacokinetic properties supportive of clinical development. On the basis of the work presented here, a clinical phase I trial of BAY 2402234 in myeloid malignancies (AML, MDS and CMML) has recently been initiated (NCT 03404726).
References
Ferrara F, Schiffer CA. Acute myeloid leukaemia in adults. Lancet. 2013;381:484–95.
Kavianpour M, Ahmadzadeh A, Shahrabi S, Saki N. Significance of oncogenes and tumor suppressor genes in AML prognosis. Tumour Biol. 2016;37:10041–52.
McCulloch D, Brown C, Iland H. Retinoic acid and arsenic trioxide in the treatment of acute promyelocytic leukemia: current perspectives. Onco Targets Ther. 2017;10:1585–601.
DiNardo CD, Stein EM, de Botton S, Roboz GJ, Altman JK, Mims AS, et al. Durable remissions with ivosidenib in IDH1-mutated relapsed or refractory AML. N Engl J Med. 2018;378:2386–98.
Stein EM, DiNardo CD, Pollyea DA, Fathi AT, Roboz GJ, Altman JK, et al. Enasidenib in mutant IDH2 relapsed or refractory acute myeloid leukemia. Blood. 2017;130:722–31.
Coombs CC, Tavakkoli M, Tallman MS. Acute promyelocytic leukemia: where did we start, where are we now, and the future. Blood Cancer J. 2015;5:e304.
Estey E, Garcia-Manero G, Ferrajoli A, Faderl S, Verstovsek S, Jones D, et al. Use of all-trans retinoic acid plus arsenic trioxide as an alternative to chemotherapy in untreated acute promyelocytic leukemia. Blood. 2006;107:3469–73.
Sykes DB, Kfoury YS, Mercier FE, Wawer MJ, Law JM, Haynes MK, et al. Inhibition of dihydroorotate dehydrogenase overcomes differentiation blockade in acute myeloid leukemia. Cell. 2016;167:171–86 e15.
Treindl F, Ruprecht B, Beiter Y, Schultz S, Dottinger A, Staebler A, et al. A bead-based western for high-throughput cellular signal transduction analyses. Nat Commun. 2016;7:12852.
Liu S, Neidhardt EA, Grossman TH, Ocain T, Clardy J. Structures of human dihydroorotate dehydrogenase in complex with antiproliferative agents. Structure. 2000;8:25–33.
Baumgartner R, Walloschek M, Kralik M, Gotschlich A, Tasler S, Mies J, et al. Dual binding mode of a novel series of DHODH inhibitors. J Med Chem. 2006;49:1239–47.
Ladds M, van Leeuwen IMM, Drummond CJ, Chu S, Healy AR, Popova G, et al. A DHODH inhibitor increases p53 synthesis and enhances tumor cell killing by p53 degradation blockage. Nat Commun. 2018;9:1107.
Faber J, Krivtsov AV, Stubbs MC, Wright R, Davis TN, van den Heuvel-Eibrink M, et al. HOXA9 is required for survival in human MLL-rearranged acute leukemias. Blood. 2009;113:2375–85.
Bach C, Buhl S, Mueller D, Garcia-Cuellar MP, Maethner E, Slany RK. Leukemogenic transformation by HOXA cluster genes. Blood. 2010;115:2910–8.
Sexauer A, Perl A, Yang X, Borowitz M, Gocke C, Rajkhowa T, et al. Terminal myeloid differentiation in vivo is induced by FLT3 inhibition in FLT3/ITD AML. Blood. 2012;120:4205–14.
Hoffman-Liebermann B, Liebermann DA. Interleukin-6- and leukemia inhibitory factor-induced terminal differentiation of myeloid leukemia cells is blocked at an intermediate stage by constitutive c-myc. Mol Cell Biol. 1991;11:2375–81.
Kim DK, Seo EJ, Choi EJ, Lee SI, Kwon YW, Jang IH, et al. Crucial role of HMGA1 in the self-renewal and drug resistance of ovarian cancer stem cells. Exp Mol Med. 2016;48:e255.
Ruijtenberg S, van den Heuvel S. Coordinating cell proliferation and differentiation: antagonism between cell cycle regulators and cell type-specific gene expression. Cell Cycle. 2016;15:196–212.
Dorasamy MS, Choudhary B, Nellore K, Subramanya H, Wong PF. Dihydroorotate dehydrogenase Inhibitors Target c-Myc and arrest melanoma, myeloma and lymphoma cells at S-phase. J Cancer. 2017;8:3086–98.
Mohamad Fairus AK, Choudhary B, Hosahalli S, Kavitha N, Shatrah O. Dihydroorotate dehydrogenase (DHODH) inhibitors affect ATP depletion, endogenous ROS and mediate S-phase arrest in breast cancer cells. Biochimie. 2017;135:154–63.
Ward TH, Cummings J, Dean E, Greystoke A, Hou JM, Backen A, et al. Biomarkers of apoptosis. Br J Cancer. 2008;99:841–6.
Mancinelli R, Carpino G, Petrungaro S, Mammola CL, Tomaipitinca L, Filippini A, et al. Multifaceted roles of GSK-3 in cancer and autophagy-related diseases. Oxid Med Cell Longev. 2017;2017:4629495.
Wagner FF, Benajiba L, Campbell AJ, Weiwer M, Sacher JR, Gale JP, et al. Exploiting an Asp-Glu “switch” in glycogen synthase kinase 3 to design paralog-selective inhibitors for use in acute myeloid leukemia. Sci Transl Med. 2018;10:eaam8460.
Naqvi S, Martin KJ, Arthur JS. CREB phosphorylation at Ser133 regulates transcription via distinct mechanisms downstream of cAMP and MAPK signalling. Biochem J. 2014;458:469–79.
Iliopoulos D, Hirsch HA, Struhl K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell. 2009;139:693–706.
Iliopoulos D, Hirsch HA, Wang G, Struhl K. Inducible formation of breast cancer stem cells and their dynamic equilibrium with non-stem cancer cells via IL6 secretion. Proc Natl Acad Sci USA. 2011;108:1397–402.
Marhall A, Kazi JU, Ronnstrand L. The Src family kinase LCK cooperates with oncogenic FLT3/ITD in cellular transformation. Sci Rep. 2017;7:13734.
Munier-Lehmann H, Vidalain PO, Tangy F, Janin YL. On dihydroorotate dehydrogenases and their inhibitors and uses. J Med Chem. 2013;56:3148–67.
Dodion PF, Wagener T, Stoter G, Drozd A, Lev LM, Skovsgaard T, et al. Phase II trial with Brequinar (DUP-785, NSC 368390) in patients with metastatic colorectal cancer: a study of the Early Clinical Trials Group of the EORTC. Ann Oncol. 1990;1:79–80.
Natale R, Wheeler R, Moore M, Dallaire B, Lynch W, Carlson R, et al. Multicenter phase II trial of brequinar sodium in patients with advanced melanoma. Ann Oncol. 1992;3:659–60.
Noe DA, Rowinsky EK, Shen HS, Clarke BV, Grochow LB, McGuire WB, et al. Phase I and pharmacokinetic study of brequinar sodium (NSC 368390). Cancer Res. 1990;50:4595–9.
Dexter DL, Hesson DP, Ardecky RJ, Rao GV, Tippett DL, Dusak BA, et al. Activity of a novel 4-quinolinecarboxylic acid, NSC 368390 [6-fluoro-2-(2’-fluoro-1,1’-biphenyl-4-yl)-3-methyl-4-quinolinecarb oxylic acid sodium salt], against experimental tumors. Cancer Res. 1985;45(11 Pt 1):5563–8.
Schwartsmann G, Peters GJ, Laurensse E, de Waal FC, Loonen AH, Leyva A, et al. DUP 785 (NSC 368390): schedule-dependency of growth-inhibitory and antipyrimidine effects. Biochem Pharmacol. 1988;37:3257–66.
Zhang C, Chu M. Leflunomide: a promising drug with good antitumor potential. Biochem Biophys Res Commun. 2018;496:726–30.
Mattar T, Kochhar K, Bartlett R, Bremer EG, Finnegan A. Inhibition of the epidermal growth factor receptor tyrosine kinase activity by leflunomide. FEBS Lett. 1993;334:161–4.
Shawver LK, Schwartz DP, Mann E, Chen H, Tsai J, Chu L, et al. Inhibition of platelet-derived growth factor-mediated signal transduction and tumor growth by N-[4-(trifluoromethyl)-phenyl]5-methylisoxazole-4-carboxamide. Clin Cancer Res. 1997;3:1167–77.
O’Donnell EF, Saili KS, Koch DC, Kopparapu PR, Farrer D, Bisson WH, et al. The anti-inflammatory drug leflunomide is an agonist of the aryl hydrocarbon receptor. PLoS ONE. 2010;5::e13128.
Petzer JP, Petzer A. Leflunomide, a reversible monoamine oxidase inhibitor. Cent Nerv Syst Agents Med Chem. 2016;16:112–9.
Hamilton LC, Vojnovic I, Warner TD. A771726, the active metabolite of leflunomide, directly inhibits the activity of cyclo-oxygenase-2 in vitro and in vivo in a substrate-sensitive manner. Br J Pharmacol. 1999;127:1589–96.
Konopleva M, Letai A. BCL-2 inhibition in AML: an unexpected bonus? Blood. 2018;132:1007–12.
Shah SN, Kerr C, Cope L, Zambidis E, Liu C, Hillion J, et al. HMGA1 reprograms somatic cells into pluripotent stem cells by inducing stem cell transcriptional networks. PLoS ONE. 2012;7:e48533.
Sheikh BN, Yang Y, Schreuder J, Nilsson SK, Bilardi R, Carotta S, et al. MOZ (KAT6A) is essential for the maintenance of classically defined adult hematopoietic stem cells. Blood. 2016;128:2307–18.
Matushansky I, Radparvar F, Skoultchi AI. Reprogramming leukemic cells to terminal differentiation by inhibiting specific cyclin-dependent kinases in G1. Proc Natl Acad Sci USA. 2000;97:14317–22.
Matushansky I, Radparvar F, Skoultchi AI. CDK6 blocks differentiation: coupling cell proliferation to the block to differentiation in leukemic cells. Oncogene. 2003;22:4143–9.
Koga Y, Kimura N, Minowada J, Mak TW. Expression of the human T-cell-specific tyrosine kinase YT16 (lck) message in leukemic T-cell lines. Cancer Res. 1988;48:856–9.
Von Knethen A, Abts H, Kube D, Diehl V, Tesch H. Expression of p56lck in B-cell neoplasias. Leuk Lymphoma. 1997;26:551–62.
Bond MR, Hanover JA. A little sugar goes a long way: the cell biology of O-GlcNAc. J Cell Biol. 2015;208:869–80.
Sun C, Shang J, Yao Y, Yin X, Liu M, Liu H, et al. O-GlcNAcylation: a bridge between glucose and cell differentiation. J Cell Mol Med. 2016;20:769–81.
Andres-Bergos J, Tardio L, Larranaga-Vera A, Gomez R, Herrero-Beaumont G, Largo R. The increase in O-linked N-acetylglucosamine protein modification stimulates chondrogenic differentiation both in vitro and in vivo. J Biol Chem. 2012;287:33615–28.
Ishihara K, Takahashi I, Tsuchiya Y, Hasegawa M, Kamemura K. Characteristic increase in nucleocytoplasmic protein glycosylation by O-GlcNAc in 3T3-L1 adipocyte differentiation. Biochem Biophys Res Commun. 2010;398:489–94.
Kim SH, Kim YH, Song M, An SH, Byun HY, Heo K, et al. O-GlcNAc modification modulates the expression of osteocalcin via OSE2 and Runx2. Biochem Biophys Res Commun. 2007;362:325–9.
Wu D, Wang W, Chen W, Lian F, Lang L, Huang Y, et al. Pharmacological inhibition of dihydroorotate dehydrogenase induces apoptosis and differentiation in acute myeloid leukemia cells. Haematologica. 2018;103:1472–83.
Acknowledgements
We thank the technical staff at Bayer. We thank Mike Jeffers, Martin Michels, Kristina Bundra, Jian Yu, Dennis Healy, and Charles Cai for proof reading the manuscript. We thank Helmholtz-Zentrum Berlin for the allocation of synchrotron radiation beamtime and the staff at beamline BL14.1 and moloX GmbH for data collection services. We thank the teams at CrownBiosciences, EPO GmbH, Charles River, and NMI TT.
Author information
Authors and Affiliations
Contributions
Manuscript drafting: AJ, JAL; manuscript revision: all authors; In vitro data: SC, LE, JAL, KB, and KZ; In vivo data: AJ, AE, PL, and AH; RNASeq analysis: HS, CM, RL, and AJ; DigiWest analysis: CM and AJ; Ex vivo analysis: SC, HM, CM, DS, and AJ; Chemical synthesis and analysis: SG, SF, and MB; Crystallography: AF; Result interpretation and study designs: AJ, CM, DBS, DTS, SC, JAL, and SG
Corresponding author
Ethics declarations
Conflict of interest
SC, CM, HS, KB, AF, AE, PL, KZ, HM, RL, DS, MB, AH, SG and AJ are employees of Bayer AG. SC, AH, RL, MB, SG and AJ are stockholders of Bayer AG. SC, CM, KZ, DBS, SF, SG and AJ are listed as inventors of Patent application WO2018/077923. DBS, DTS, DS, HM and AJ are listed as inventors of patent application WO2017/037022. DBS is a co-founder and owns equity in Clear Creek Bio. JAL, DBS, DTS, and LE received research funding from Bayer AG. DTS declares the following additional competing interests: Novartis: sponsored research; Agios: director; equity holder; Fate Therapeutics: consultant, equity holder; Magenta Therapeutics: director; equity holder; consultant; Clear Creek Bio: director; equity holder; consultant; Red Oak Medicines: director; equity holder, consultant; LifeVaultBio: director; equity holder; Fog Pharma: consultant
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Christian, S., Merz, C., Evans, L. et al. The novel dihydroorotate dehydrogenase (DHODH) inhibitor BAY 2402234 triggers differentiation and is effective in the treatment of myeloid malignancies. Leukemia 33, 2403–2415 (2019). https://doi.org/10.1038/s41375-019-0461-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-019-0461-5
This article is cited by
-
Targeted inhibition of DHODH is synergistic with BCL2 blockade in HGBCL with concurrent MYC and BCL2 rearrangement
BMC Cancer (2024)
-
Application of omics in the diagnosis, prognosis, and treatment of acute myeloid leukemia
Biomarker Research (2024)
-
Disulfiram treatment suppresses antibody-producing reactions by inhibiting macrophage activation and B cell pyrimidine metabolism
Communications Biology (2024)
-
DHODH inhibition represents a therapeutic strategy and improves abiraterone treatment in castration-resistant prostate cancer
Oncogene (2024)
-
Pharmacological induction of the hypoxia response pathway in Huh7 hepatoma cells limits proliferation but increases resilience under metabolic stress
Cellular and Molecular Life Sciences (2024)